Clear Sky Science · en
Multi task learning based early prediction model for antibiotic resistance using multi institutional cohort data
Why this matters for everyday health care
Antibiotic resistance is quietly turning once-curable infections into life-threatening illnesses. Doctors must often choose antibiotics before lab tests reveal which drugs will work, a delay that can take several days. This study explores how computer models trained on hospital records can give doctors an early heads-up about which antibiotics are likely to fail, helping protect patients and slow the spread of resistant bacteria.
Antibiotics under pressure
Modern medicine leans heavily on antibiotics for everything from routine surgery to cancer care. Yet bacteria are evolving quickly, and new antibiotics are appearing more slowly than old ones are losing their power. In South Korea, where this study took place, up to one-third of antibiotic prescriptions in hospitals are judged inappropriate. Culture tests that reveal whether a germ is resistant usually take three to five days, forcing doctors to treat in the dark. The researchers set out to predict resistance to nine major groups of antibiotics, using only information available early in a patient’s hospital stay.
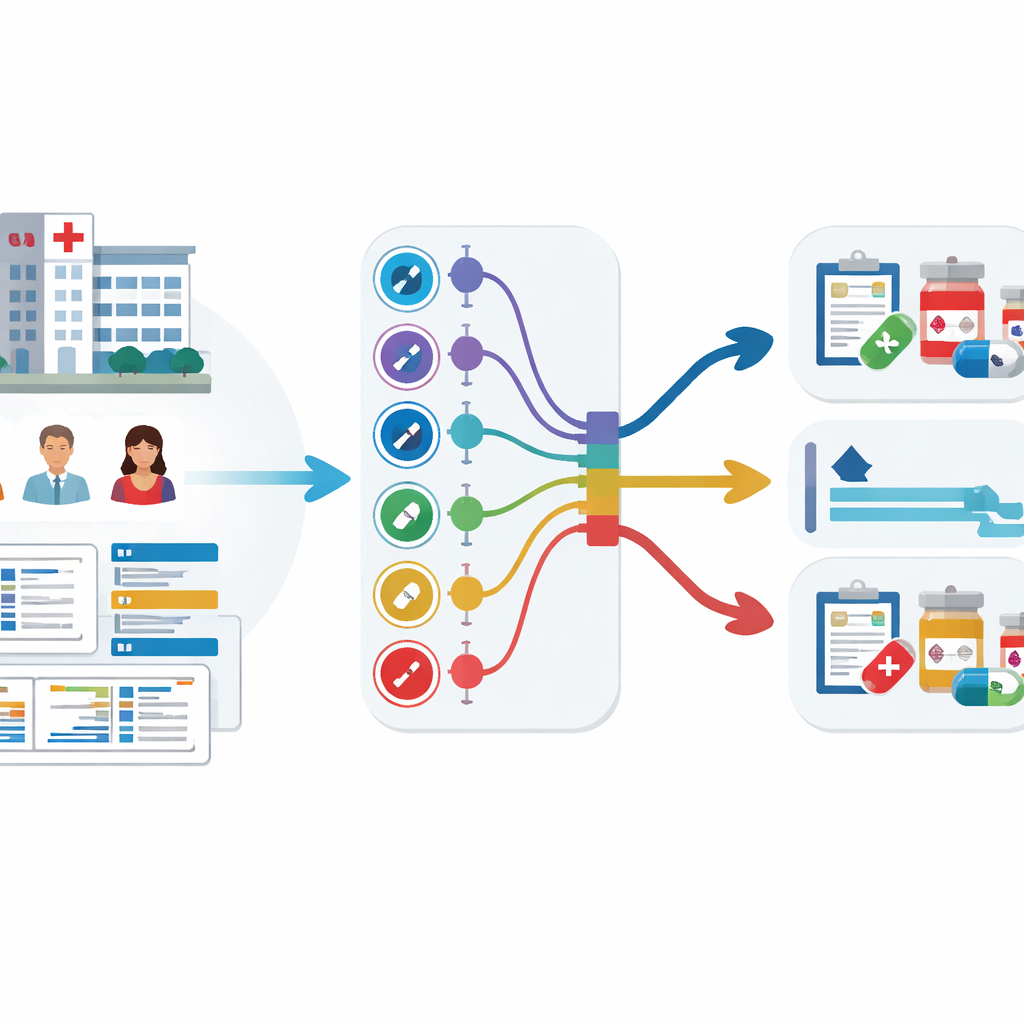
Turning hospital data into early warnings
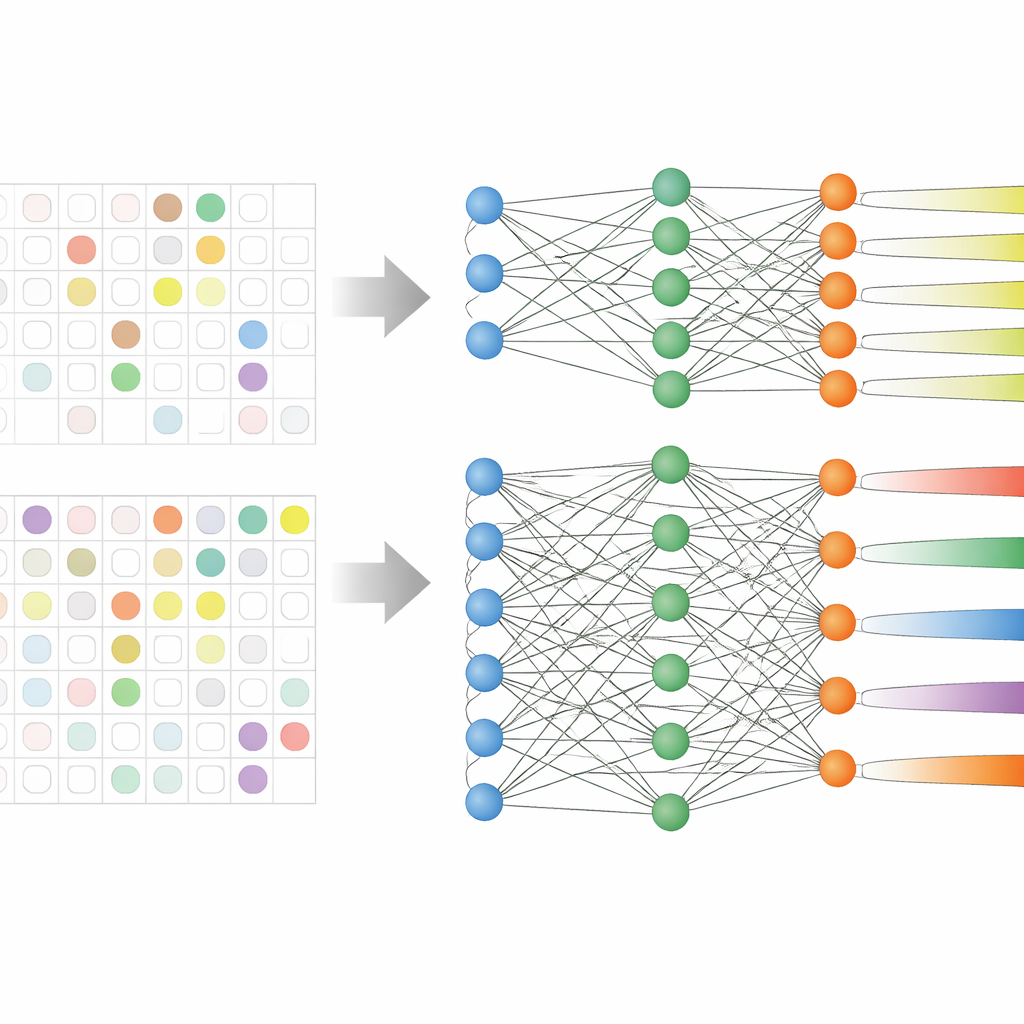
The team analyzed electronic medical records from 59,551 adult patients in three large Korean hospitals over more than a decade. These records included age, vital signs, length of stay, previous antibiotic use, and, crucially, past culture test results showing which antibiotics had already failed. Instead of training nine separate models—one for each antibiotic group—they used a strategy called multi-task learning, which allows a single model to learn several related prediction jobs at once. Two versions were tested: one with a shared core that branches into separate outputs for each antibiotic group (hard sharing), and one in which the tasks are linked more loosely (soft sharing).
Solving the problem of incomplete lab results
Real hospital data are messy: not every patient is tested against every antibiotic. Earlier studies often threw out these partially labeled cases, shrinking their datasets and ignoring patterns that span different drugs. Here, the researchers rewrote the model’s learning rule so it simply skips missing lab results when calculating its training error. This allowed them to keep nearly all patients in the analysis, even when some antibiotic results were absent. The hard-sharing model especially benefited from this setup, because it could learn common signals of resistance while still fine-tuning its predictions for each drug group.
How well did the models perform?
When tested on data from hospitals they had not seen before, the multi-task models generally outperformed more traditional methods like logistic regression and standard boosting algorithms. On average, the hard-sharing model showed the best balance of accuracy and stability across the nine antibiotic classes, and it was particularly strong for drug groups with the scarcest data, such as aminoglycosides. A separate analysis that explains which features drive the predictions found that a patient’s previous resistance to an antibiotic was the single most important factor, followed by resistance or use of related drugs, as well as age and length of stay. Subgroup analyses showed that hard sharing worked best when prior culture results for a given antibiotic were available, while soft sharing was better when such history was missing.
What this means for patients and clinicians
The study suggests that smart use of hospital records can provide early, reasonably reliable forecasts of antibiotic resistance across several major drug families. Even modest gains in prediction accuracy, applied to large numbers of patients, could help doctors choose narrower, more targeted antibiotics sooner and step down from broad drugs more confidently. That, in turn, may reduce complications, shorten hospital stays, and slow the broader march of resistance. The authors caution that their model still needs real-world testing and refinement, especially when past culture data are unavailable, but they argue that multi-task learning offers a promising way to make better use of imperfect medical data and support more careful antibiotic use.
Citation: Kim, Y., Jeong, I., Park, JH. et al. Multi task learning based early prediction model for antibiotic resistance using multi institutional cohort data. Sci Rep 16, 11891 (2026). https://doi.org/10.1038/s41598-026-41185-z
Keywords: antibiotic resistance, clinical decision support, electronic health records, machine learning, multi-task learning