Clear Sky Science · en
CSWin-MDKDNet: cross-shaped window network with multi-dimensional fusion and knowledge distillation for medical image segmentation
Sharper Views Inside the Body
Modern medicine relies heavily on images—CT scans, MRIs and skin photos—to spot organs, tumors and other structures. But before doctors or computers can measure or track disease, they often need to "color in" each organ or lesion precisely, a task called segmentation. This paper introduces a new artificial intelligence system, CSWin-MDKDNet, that makes this outlining step more accurate and efficient across several types of medical images, potentially improving diagnosis, treatment planning and follow-up care for many patients.
Why Drawing Boundaries Matters
When radiologists plan surgery, measure a heart’s pumping strength or estimate the size of a skin lesion, they depend on clear boundaries in their images. Traditionally, human experts draw these outlines by hand, which is slow, tiring and can vary from one person to another. Earlier computer methods based on convolutional neural networks learned to recognize local patterns like edges and textures, and they transformed medical image analysis. Yet these systems still struggled to see the "big picture"—how distant parts of an image relate to one another—while also keeping fine details along organ edges intact. This trade-off between global context and local precision has limited the reliability of automated tools in demanding clinical settings.
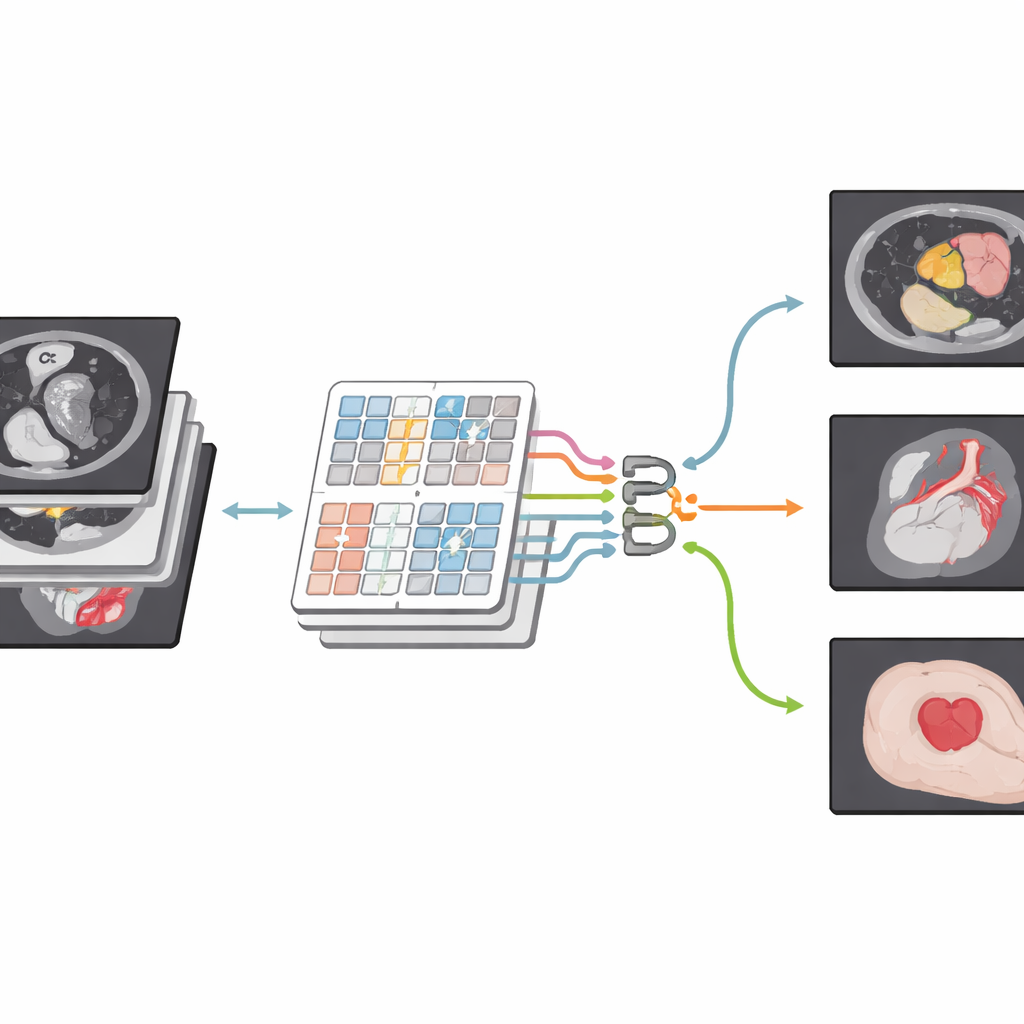
A New Way of Looking at Medical Images
The authors build on a newer family of models known as Transformers, originally developed for language but now widely used in computer vision. Their network, CSWin-MDKDNet, starts by breaking a medical image into patches and passing them through a Transformer module that views the picture in cross-shaped stripes, horizontally and vertically. This design allows the system to connect far-apart regions—such as the top and bottom of the liver—without an explosion in computation. Around this core, the model adopts a U-shaped encoder–decoder layout that has become standard in medical imaging: one path gradually shrinks the image to capture high-level structure, while another path expands it back to full size, producing a detailed segmentation map that lines up with the original scan.
Blending Details from Many Directions
Simply stacking more layers and attention blocks can make a model powerful but also bloated and unfocused. To address this, the authors introduce a Multi-dimensional Selective Fusion module that acts like a smart mixer for image features. It looks at information across three aspects at once: the different "channels" that encode various visual cues, the spatial layout that captures where edges and textures occur, and several scales that range from fine details to broad context. By using targeted weighting rather than treating all features equally, this module boosts information that truly helps distinguish one organ from another—such as the subtle, irregular outline of the pancreas—while muting distractions from noise and background tissue.
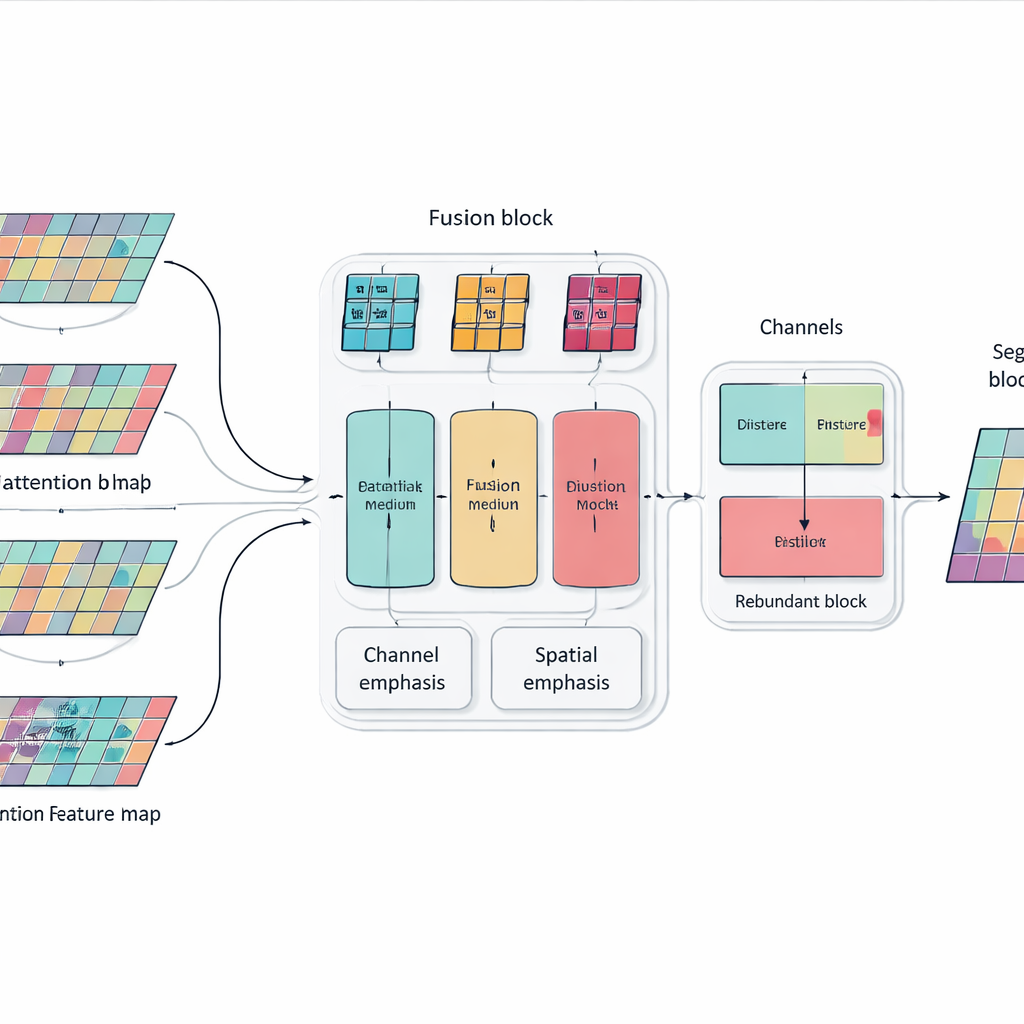
Teaching the Network Not to Repeat Itself
Another problem with very deep networks is redundancy: later layers can end up repeating patterns already learned earlier, wasting capacity and sometimes confusing the decision process. Instead of adding extra pruning modules, the researchers introduce a simple training rule inspired by knowledge distillation. Within each block of the network, they encourage deeper channels to absorb the most useful information from shallower ones while avoiding unnecessary duplication. This internal "teacher–student" relationship nudges the model toward compact, consistent representations, which helps it generalize better to new patients and different scanners without increasing the cost of running the system.
Proven Gains Across Organs and Modalities
To test their approach, the team evaluated CSWin-MDKDNet on three demanding benchmarks. On multi-organ abdominal CT scans, the system achieved the highest average overlap between its predictions and expert labels, particularly improving on hard-to-outline organs like the pancreas. On cardiac MRI, it offered more precise contours of the heart’s chambers and muscle, which are critical for measuring heart function. On a large collection of skin lesion photos, it produced cleaner boundaries than several strong competing models. Notably, these gains came with fewer parameters and lower computation than classic Transformer-based designs, meaning the method is better suited for practical deployment in clinics and hospitals.
Clearer Outlines for Better Care
In everyday terms, this work shows how smarter software can trace the shapes of organs and lesions in medical images more accurately, while using computer resources more efficiently. By combining a broad view of the image with carefully tuned attention to important details, and by discouraging wasteful repetition inside the network, CSWin-MDKDNet delivers more reliable digital outlines that doctors can trust. Such improvements may not be visible to patients directly, but they can support more precise surgery planning, more consistent tracking of disease over time and ultimately more confident decisions at the bedside.
Citation: Cui, G., Lin, H., Sun, L. et al. CSWin-MDKDNet: cross-shaped window network with multi-dimensional fusion and knowledge distillation for medical image segmentation. Sci Rep 16, 11532 (2026). https://doi.org/10.1038/s41598-026-40690-5
Keywords: medical image segmentation, deep learning, transformer networks, organ and lesion analysis, computer-aided diagnosis