Clear Sky Science · en
Contextual anatomy-guided deep learning for accurate fovea segmentation in diabetic retinopathy fundus images
Seeing the Danger in Diabetic Eyes
For millions of people with diabetes, a small spot at the very center of the eye’s retina—the fovea—can mean the difference between clear sight and permanent vision loss. Doctors already use color photographs of the back of the eye (fundus images) to screen for diabetic retinopathy, a leading cause of preventable blindness. But today’s automated tools mostly pinpoint the fovea as a single dot rather than outlining its true area, which is what doctors really need to decide who requires injections, who needs laser treatment, and who can safely wait. This study shows how a new data-driven artificial intelligence (AI) method can learn to see the fovea more like an eye specialist does, by paying attention to the entire anatomical landscape of the eye rather than just the fovea itself. 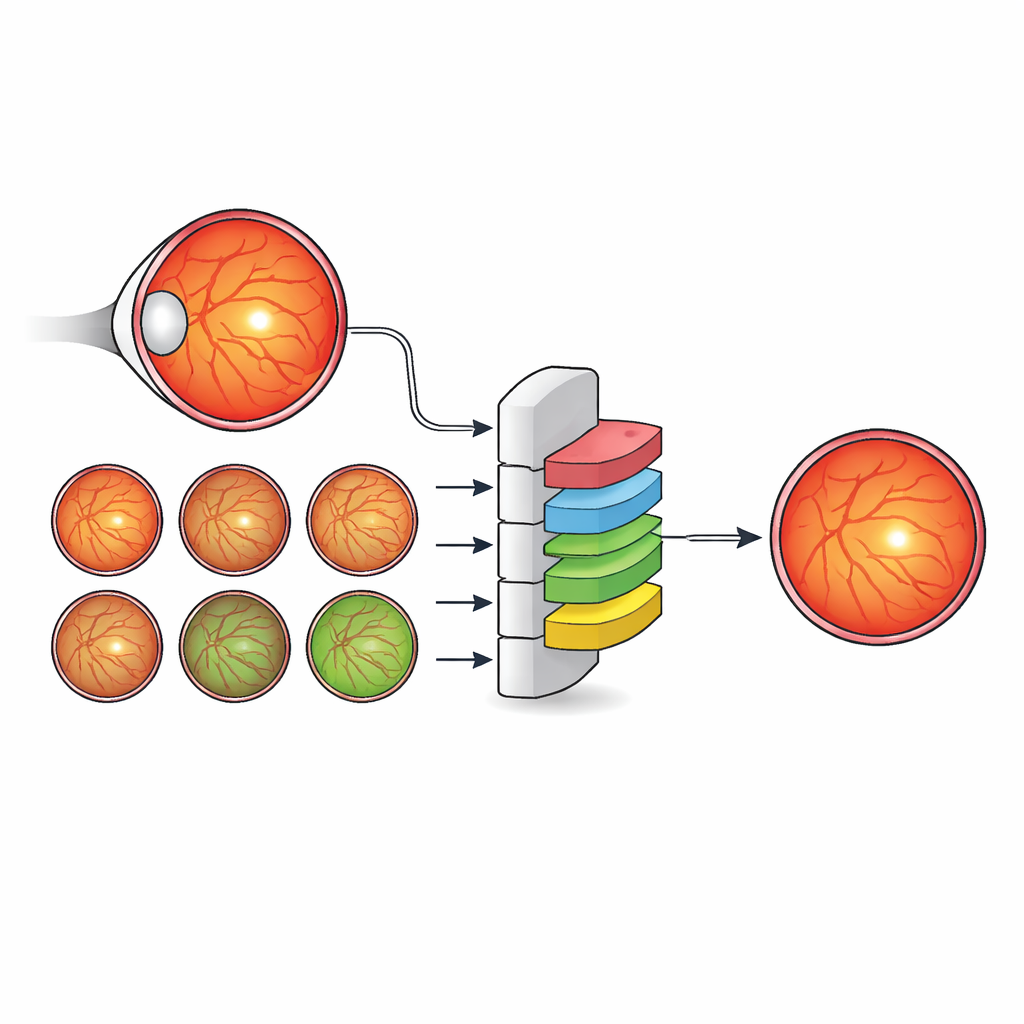
Why the Fovea Is Hard to Outline
The fovea is a tiny, shallow depression in the retina that gives us sharp central vision and rich color perception. On fundus images, its borders are fuzzy, its appearance varies from camera to camera, and disease can blur it even further. Traditional computer methods relied on simple brightness and geometric rules, which quickly break down in the presence of lesions or poor image quality. Newer deep learning systems have improved fovea detection, but they usually either treat each eye structure separately or rely on increasingly complex network designs. Both approaches often ignore a basic clinical reality: eye doctors do not look for the fovea in isolation—they use landmarks like the optic disc and the pattern of blood vessels to infer where the fovea must be, even when it is hard to see.
Teaching AI to Read the Eye’s Roadmap
The authors propose a different strategy: instead of inventing ever more intricate network architectures, they enrich the labels used to train a standard, efficient model. They created a new dataset, IDRiD-RETA-FV, in which eye specialists carefully drew pixel-level outlines not only for the fovea but also for the optic disc, the blood vessels, the overall retina, and the background. Two ophthalmologists followed a strict protocol and reached near-perfect agreement on these markings, providing a solid reference for the AI system. Into this dataset they plugged MNv4Fovea, a segmentation model based on a widely used architecture, but trained it to segment all anatomical structures at once. By progressively adding more context to the training labels—first only fovea versus background, then including the optic disc, then the retina, and finally the vessels—they could directly measure how much each piece of anatomy helps the model find the fovea.
How Context and Clever Augmentation Boost Accuracy
The results show that context matters enormously. When the model learned only from fovea and background labels, it correctly identified just over a quarter of foveal pixels. Simply adding the optic disc offered only modest gains. But once the retina outline was included, performance climbed sharply, and adding blood vessels pushed foveal recall above 90 percent. In other words, when the AI model was given the same roadmap eye doctors use—disc, vessels, and retinal boundary—it became far better at outlining the fovea. To help the system work across images from different cameras, the team also devised a novel way to vary image color and brightness. They modeled how other public eye datasets look using a statistical distribution called the Generalized Extreme Value (GEV) distribution and transformed training images to mimic those styles while preserving anatomy. This GEV-based augmentation nearly doubled the rate of successful fovea detections compared with standard geometric tweaks alone. 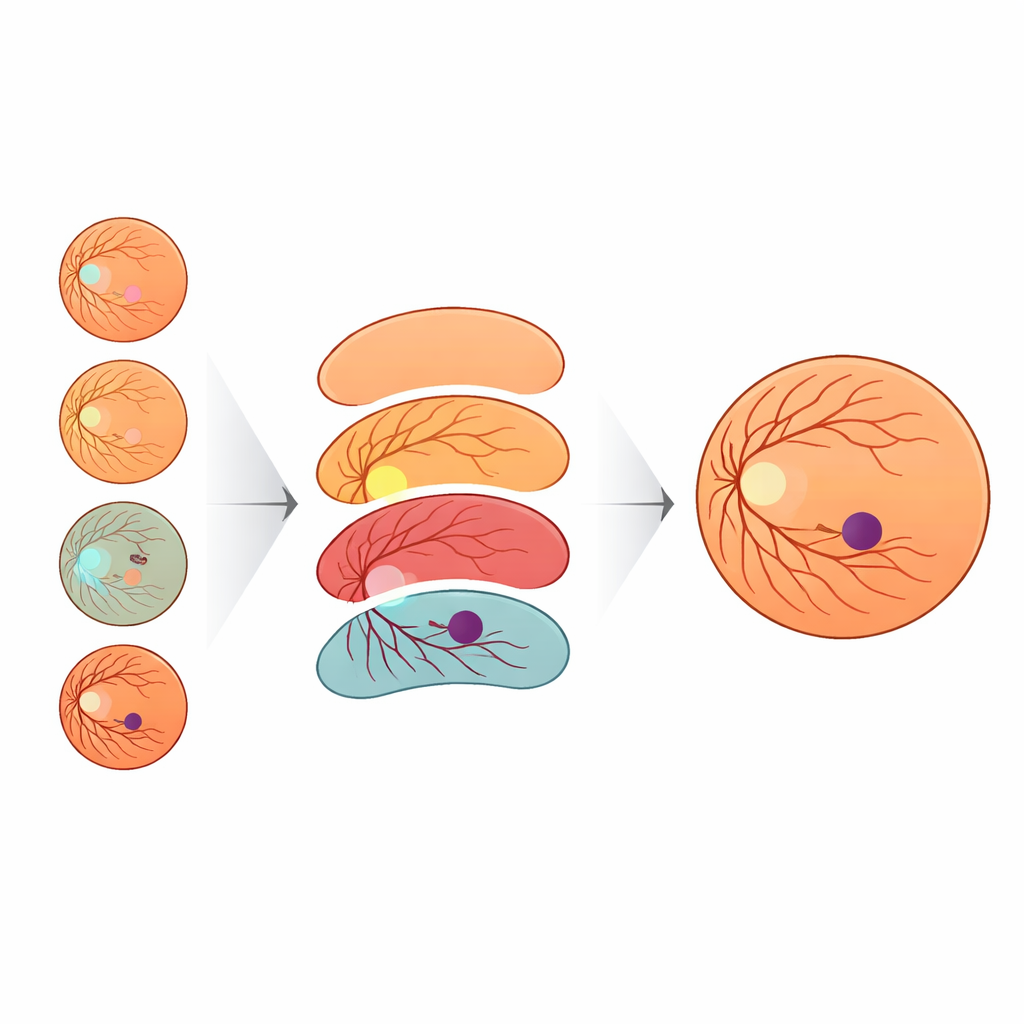
Proving It Works Across Many Datasets
MNv4Fovea was tested both on the new expert-labeled dataset and on several well-known public collections of retinal images. On the held-out test split of IDRiD-RETA-FV, it achieved high overlap between predicted and true foveal regions and located the foveal center within only a few pixels—essentially always detecting the fovea when it was present. On external datasets such as MESSIDOR, REFUGE, ARIA, and MAPLES-DR, the model matched or exceeded state-of-the-art methods in how close its foveal predictions were to the reference points. Interestingly, in some cases the numerical scores appeared modest because the external datasets defined “fovea” differently—for example, marking a broad macular zone instead of the tiny clinical fovea—yet expert review showed that the model’s predictions were anatomically sensible and often closer to specialist opinion than to the original labels.
What This Means for Patients and Future Tools
For people living with diabetes, the study highlights a path toward more reliable, widely deployable screening tools that do more than place a dot on the retina. By learning from the full anatomical context—disc, vessels, and retinal outline—rather than relying on architectural tricks alone, MNv4Fovea can precisely map the foveal region, a key step in deciding whether swelling or lesions are close enough to threaten central vision. The authors argue that this data-centric, anatomy-guided approach can complement future advances in network design, and that similar strategies may benefit many other areas of medical imaging where subtle structures must be found in complex, diseased landscapes.
Citation: Chankhachon, S., Kansomkeat, S., Bhurayanontachai, P. et al. Contextual anatomy-guided deep learning for accurate fovea segmentation in diabetic retinopathy fundus images. Sci Rep 16, 10388 (2026). https://doi.org/10.1038/s41598-026-40287-y
Keywords: diabetic retinopathy, retinal imaging, fovea segmentation, deep learning, medical image analysis