Clear Sky Science · en
Mitochondrial apoptosis gene-based pathomics for ovarian cancer prognosis
A New Way to Read Cancer Clues
Ovarian cancer is often called a “silent killer” because it usually stays unnoticed until it has already spread, making it hard to treat and predict who will do well. This study offers a fresh approach: it teaches computers to read hidden patterns in microscope images of tumors and link them to how tumor cells decide to live or die. By doing so, the researchers aim to sort patients into higher- and lower-risk groups more accurately than current methods, and to uncover weak points that new treatments might target.
Looking Deeper into Tumor Pictures
Pathologists traditionally judge cancer by looking at stained tissue slices under a microscope and describing what they see. While this method is powerful, it is also subjective and struggles to capture subtle details. The team behind this study used digital “whole-slide” images of ovarian tumors and cut each large image into many smaller tiles. A deep learning system called ResNet50 scanned these tiles to pull out thousands of mathematical features that describe patterns in cell shape, texture, and how cells and supporting tissues are arranged. In parallel, specialized software quantified more classic image traits, such as nucleus size and tissue structure, to build a rich picture of each tumor’s appearance. 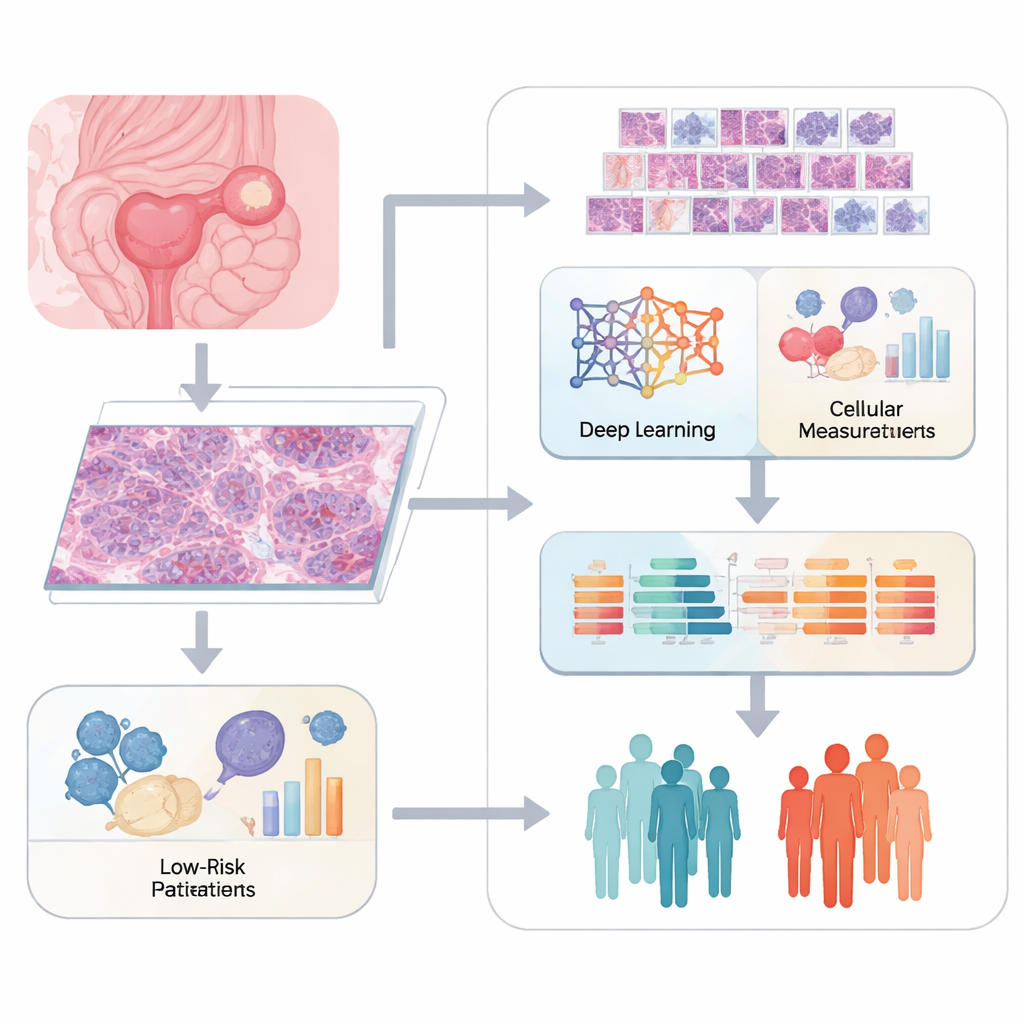
Connecting Images to Cell Death Signals
Instead of stopping at what the tumor looks like, the researchers tied these image features to a vital life–death switch inside cells known as mitochondrial apoptosis. This process is governed by a family of proteins that can either push a damaged cell to self-destruct or help it resist death, a key reason some tumors shrug off chemotherapy. Using gene activity data from hundreds of ovarian cancer samples, the team scored how strongly the mitochondrial death pathway was turned on in each patient. They then kept only those image features that were clearly linked to this pathway, building a “pathomics” bridge between tumor appearance and internal wiring.
Building a Risk Score for Patients
From thousands of initial measurements, the researchers whittled the set down through several layers of statistical filtering and machine learning, arriving at 12 core image-based features that best predicted how long patients survived. These features were combined into a single number, a risk score they call the MAR-PTL model. Patients were split into high- and low-risk groups based on this score. The model correctly distinguished outcomes in both a training group and a separate test group, outperforming standard clinical measures like disease stage and age. In other words, the hidden visual fingerprints of a tumor, once decoded, carried powerful information about a patient’s future.
A Key Survival Protein and Its Neighborhood
Digging deeper, the study highlighted one gene, BCL2L2, as a central player. This gene helps block mitochondrial cell death, and higher levels of it were linked to worse survival. Intriguingly, specific deep learning features from the tissue images rose or fell in step with BCL2L2, suggesting that the computer could “see” how this molecular brake on cell death reshapes tumor architecture. Single-cell analysis, which looks at thousands of individual cells one by one, showed that BCL2L2-rich cancer cells had a distinctive energy‑related metabolism and communicated intensely with nearby support cells such as fibroblasts and cells lining blood vessels. These interactions appeared to form a nurturing neighborhood that helps tumors resist stress and spread. 
What This Could Mean for Patients
Taken together, this work shows that microscope images hold far more than meets the eye. By fusing digital pathology with knowledge of how tumor cells choose life or death, the MAR-PTL model offers a sharper way to flag ovarian cancer patients at highest risk. The discovery of BCL2L2 as a key hub, and its tight links to both tumor metabolism and its surrounding support cells, points to new treatment ideas—for example, drugs that disarm this survival switch or disrupt its communication with nearby tissues. While these findings still need to be tested in prospective studies and laboratory experiments, they pave the way toward more personalized care and, ultimately, better chances for women facing this challenging disease.
Citation: Qin, Lh., Huang, X., Yang, C. et al. Mitochondrial apoptosis gene-based pathomics for ovarian cancer prognosis. Sci Rep 16, 13231 (2026). https://doi.org/10.1038/s41598-026-40121-5
Keywords: ovarian cancer, digital pathology, deep learning, mitochondrial apoptosis, tumor microenvironment