Clear Sky Science · en
MSF-VMDNet for multi class segmentation of skin cancer whole slide images using a multi frequency dual encoder network
Why mapping skin cancer matters
When doctors diagnose skin cancer, they often look at ultra‑detailed microscope images of stained tissue slices. These pictures show not only the tumor, but also many kinds of normal skin structures tightly woven together. Drawing precise boundaries around each region by hand is slow and can vary from one specialist to another. This study presents a new computer method designed to automatically separate these tangled tissues with very high accuracy, potentially saving time and making diagnoses more consistent.
From simple spots to tangled tissue
Skin cancer is among the most common cancers worldwide, driven in part by increased exposure to ultraviolet light. Modern diagnosis usually begins with dermoscopy, a special kind of magnified skin photograph, and often continues with a biopsy: a tiny piece of skin is removed, stained, and examined under a microscope. In these whole‑slide images, tumors wind through layers such as the outer skin, deeper connective tissue, hair follicles, sweat glands, and fat. Many of these regions look alike in color and texture, and their borders can be fuzzy, making it difficult even for experts to clearly separate tumor from healthy tissue.
A new way to read complex pictures
Most earlier computer tools treated the problem like coloring a picture with only two crayons: “lesion” and “not lesion.” That may work for simpler surface photos, but it breaks down when a slide must be divided into many tissue types at once. The authors focus on a challenging dataset of non‑melanoma skin cancers in which each image must be split into up to ten distinct tissue classes. Their goal is to build a system that can understand the overall layout of the tissue while also catching tiny details along the borders—something traditional neural networks often struggle to balance.
Two brains are better than one
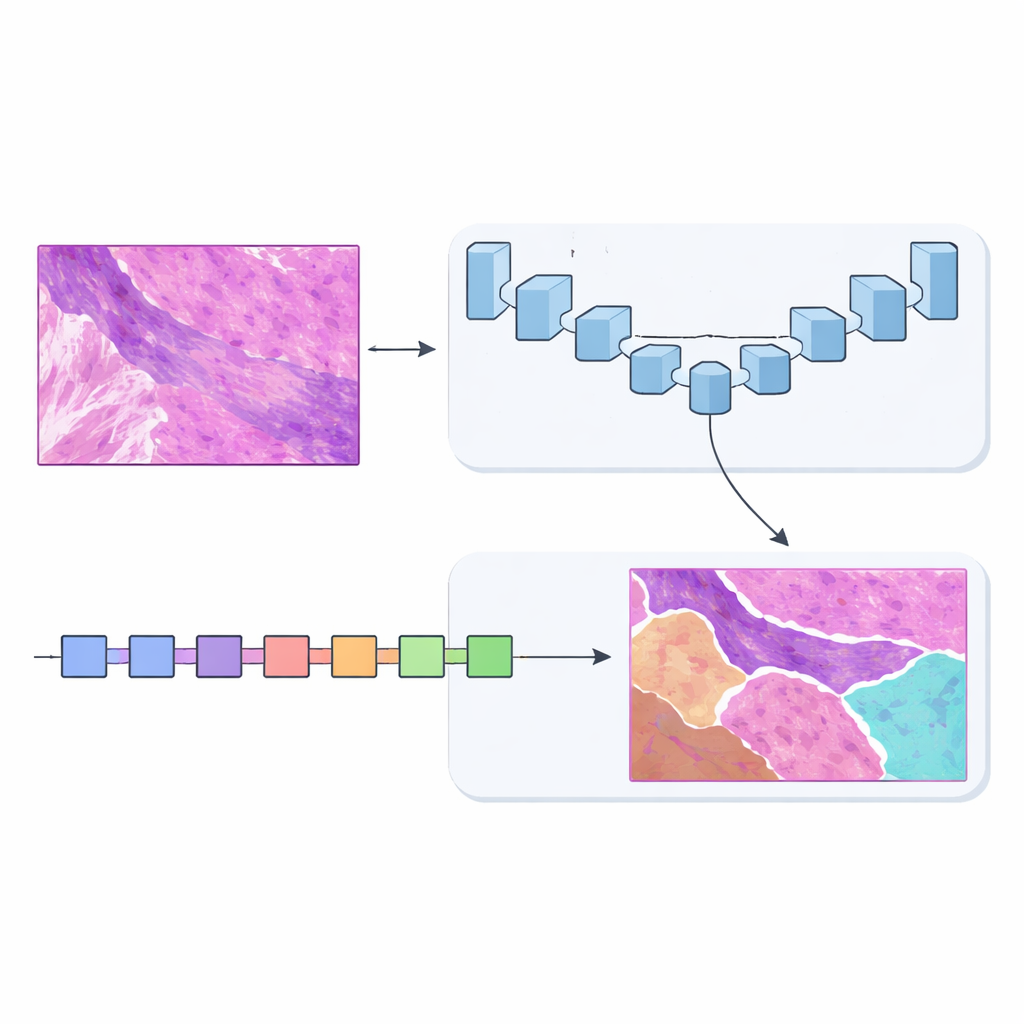
To solve this, the team designed MSF‑VMDNet, a dual‑encoder network that can be thought of as two complementary “brains” examining the same slide in parallel. One branch is based on U‑Net, a popular medical‑imaging model that excels at capturing fine local detail. The authors enhance it with a spectral module that converts the image into a collection of frequencies—some describing slow, smooth changes, others capturing sharp edges and textures—then recombines them to strengthen class boundaries. At the same time, a second branch built on a newer approach called Vision Mamba scans the slide in a more streamlined way, efficiently modeling long‑range relationships so that distant but related regions can inform one another.
Listening to many frequencies at once
Frequency information plays a central role in this architecture. By moving between the ordinary image space and a frequency‑based view, the network can treat broad shapes and tiny structures differently. Low‑frequency components help it understand the overall layout of the tumor and the surrounding skin layers, while high‑frequency components sharpen the edges where one tissue type ends and another begins. Carefully designed modules merge these views back into a spatial map, and an additional decoding block (called SCConv) filters out redundant signals while boosting truly informative patterns. The result is a cleaner, more confident map of each tissue region.
How well does it work in practice?
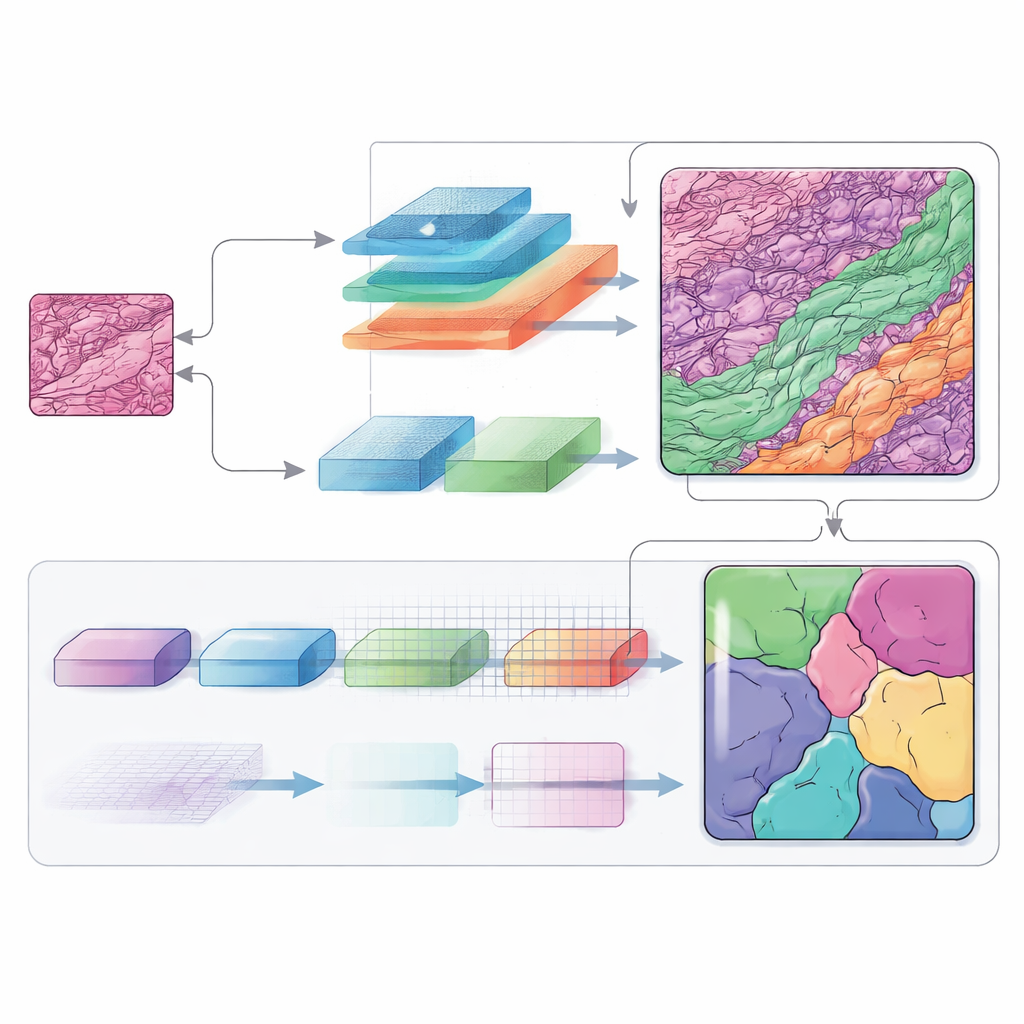
The researchers tested MSF‑VMDNet on the non‑melanoma skin cancer dataset and compared it with a wide range of leading segmentation models, including classic U‑Net, Transformer‑based methods, and other Vision Mamba hybrids. Their system produced clearer tumor outlines and fewer mistakes in tricky areas where tumor, hair follicles, and inflamed tissue intertwine. On standard overlap measures, it surpassed all competitors, reaching around 95 percent agreement with expert‑drawn masks. To check whether the method generalizes beyond one lab and one cancer type, the team also evaluated it on three additional collections: dermoscopic photos of skin lesions, microscope images of different kinds of cell nuclei, and CT scans of abdominal organs. In all cases, the method remained highly accurate and statistically outperformed strong baselines.
What this means for patients and doctors
In plain terms, MSF‑VMDNet is an automated map‑maker for medical images that can separate tumors from the crowded landscape of normal tissue with unusual precision. Although it does not replace the judgment of pathologists, it can provide a fast, detailed starting point, reducing the manual effort required and helping ensure that subtle tumor regions are not overlooked. Because the same approach works well on very different kinds of medical images, it could become a versatile tool for many diagnostic tasks. With further development and integration of clinical information, systems like this may support more reliable skin‑cancer grading and, ultimately, better‑informed treatment choices.
Citation: Zhang, J., Pu, Q., Tian, J. et al. MSF-VMDNet for multi class segmentation of skin cancer whole slide images using a multi frequency dual encoder network. Sci Rep 16, 11722 (2026). https://doi.org/10.1038/s41598-026-40044-1
Keywords: skin cancer imaging, medical image segmentation, deep learning in pathology, histopathology analysis, AI diagnostics