Clear Sky Science · en
Enhanced drug disease association prediction through multimodal data integration and meta path guided global local feature fusion
Why finding new uses for old drugs matters
Developing a new medicine from scratch is slow, risky, and extremely expensive. Yet hidden in today’s pharmacies are many drugs that might also help treat other diseases, if only we could discover those extra uses. This study presents MedPathEx, a computer-based approach that sifts through large collections of biomedical data to predict which existing drugs are likely to work for which diseases, potentially speeding up the search for new treatments and making better use of medicines we already have.
Bringing many clues together in one big map
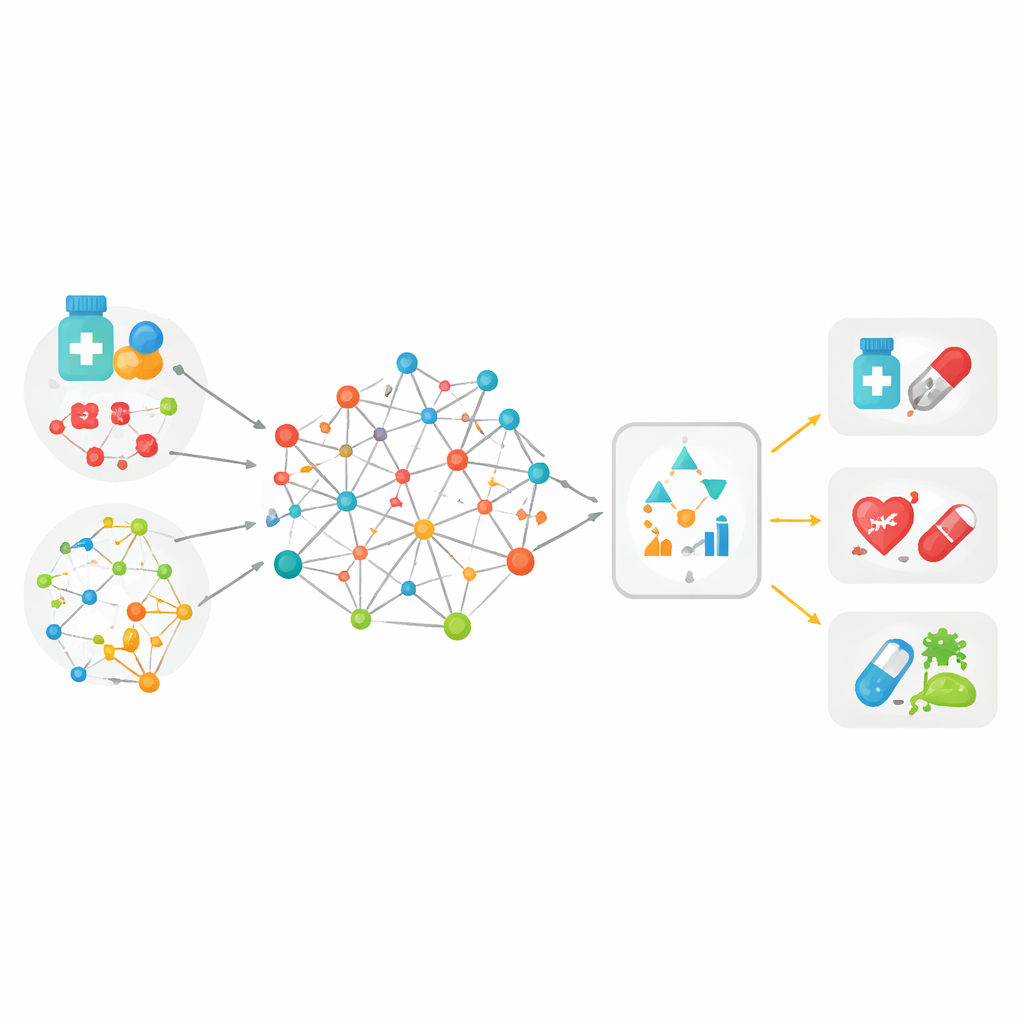
Most previous tools for matching drugs to diseases relied on just one type of information—for example, how similar two drug molecules look, or how closely two diseases are related in medical records. MedPathEx starts from the idea that no single clue is enough. The authors build a large “map” that links three kinds of players: drugs, diseases, and genes. On top of these links, they layer multiple kinds of clues: how drug molecules are built, how they are classified in therapy, what side effects they cause, how diseases appear in patients and in medical language, and what is known about gene function. By weaving all of this multimodal information into one heterogeneous network, each drug, disease, and gene is described in a richer, more realistic way than before.
Seeing both the neighborhood and the whole city
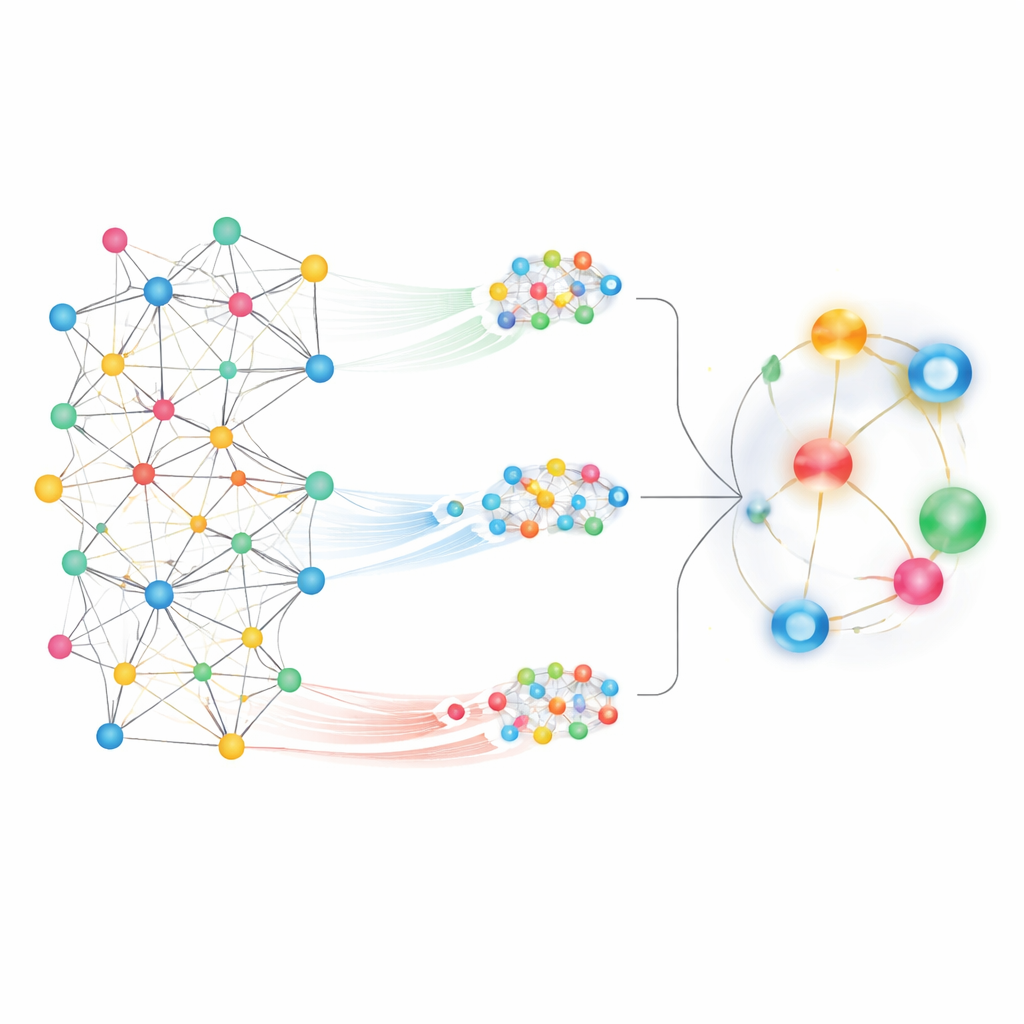
Looking at such a huge map, one challenge is deciding which patterns matter. MedPathEx tackles this by combining two complementary views of the network. First, it zooms in on local neighborhoods, tracing short, meaningful paths—for example, drug → gene → disease—to capture how specific genes might connect a medication to an illness. These patterned walks, called meta-paths, highlight fine-grained relationships. Second, the method zooms out to consider the global picture, allowing each node to “pay attention” to many others across the entire map. This global attention view picks up broader trends and long-range connections that would be missed by only following local paths.
Blending different views into a single signal
To turn these network views into predictions, MedPathEx uses modern neural-network techniques to convert the complex map into compact numerical fingerprints for every drug, disease, and gene. One part of the model learns from similarity graphs built within each type (drug–drug, disease–disease, gene–gene). Another part focuses on the local meta-path neighborhoods, while a third captures the global structure of the whole network. The model then learns how much weight to give each of these sources and fuses them into a single, combined representation for each node. When the system compares the combined fingerprints of a given drug and disease, it outputs a score that reflects how likely the pair is to be truly connected in the real world.
Testing performance and probing what matters
The researchers put MedPathEx to the test using public databases that catalog hundreds of thousands of known links among thousands of drugs, diseases, and genes. In rigorous five-fold cross-validation, MedPathEx outperformed a variety of competing approaches, including classic machine-learning methods and several advanced graph-based models. Measures of accuracy such as AUC, average precision, and F1 score were all higher, showing that the method more reliably separates true drug–disease pairs from false ones. When components were removed one by one, performance dropped, especially when either the local meta-path view or the global attention view was taken away, underscoring that both perspectives are essential. The analysis also showed that the local meta-path information contributes the most, with global and similarity features providing important refinements.
Real-world examples in heart disease and high blood pressure
Beyond numbers, the authors checked whether MedPathEx’s top-ranked suggestions for two common conditions—coronary artery disease and hypertension—matched what is known from the medical literature. For each disease, the system proposed several drugs, some of which are already used clinically for related heart problems, lending credibility to the approach. Others are less obvious candidates that nevertheless have biological links, such as affecting inflammation, blood vessel function, or plaque buildup in arteries. Network diagrams of drugs, genes, and diseases illustrate how these medicines might influence disease-related pathways, pointing to plausible mechanisms that could be explored in laboratory or clinical studies.
What this means for future medicines
In simple terms, MedPathEx shows that by combining many sources of biomedical data and looking at both local and global patterns in how drugs, genes, and diseases connect, computers can do a better job of guessing which medicines might help which illnesses. While this tool does not replace clinical trials or real-world testing, it can narrow down the vast search space and highlight promising drug candidates—especially for complex diseases like heart disease and high blood pressure. As more detailed biological data become available and as such models are further validated, approaches like MedPathEx could become powerful partners in the quest to repurpose existing drugs and design more efficient treatment strategies.
Citation: Wu, S., Wang, W., Jiao, H. et al. Enhanced drug disease association prediction through multimodal data integration and meta path guided global local feature fusion. Sci Rep 16, 11038 (2026). https://doi.org/10.1038/s41598-026-36223-9
Keywords: drug repurposing, drug–disease associations, heterogeneous networks, graph neural networks, computational drug discovery