Clear Sky Science · en
Chromatin spatial analysis by METALoci unveils sex-determining 3D regulatory hubs
How the Genome’s Shape Helps Decide Sex
Every mammal begins life with tiny gonads that could become either testes or ovaries. We have long known many of the genes that push development toward male or female, but much less about how DNA’s three-dimensional folding inside the nucleus helps steer that choice. This study shows that the physical layout of our chromosomes creates powerful control centers—three-dimensional regulatory hubs—that help determine whether the embryonic gonad becomes testis or ovary, and reveals new players and hidden switches involved in that decision. 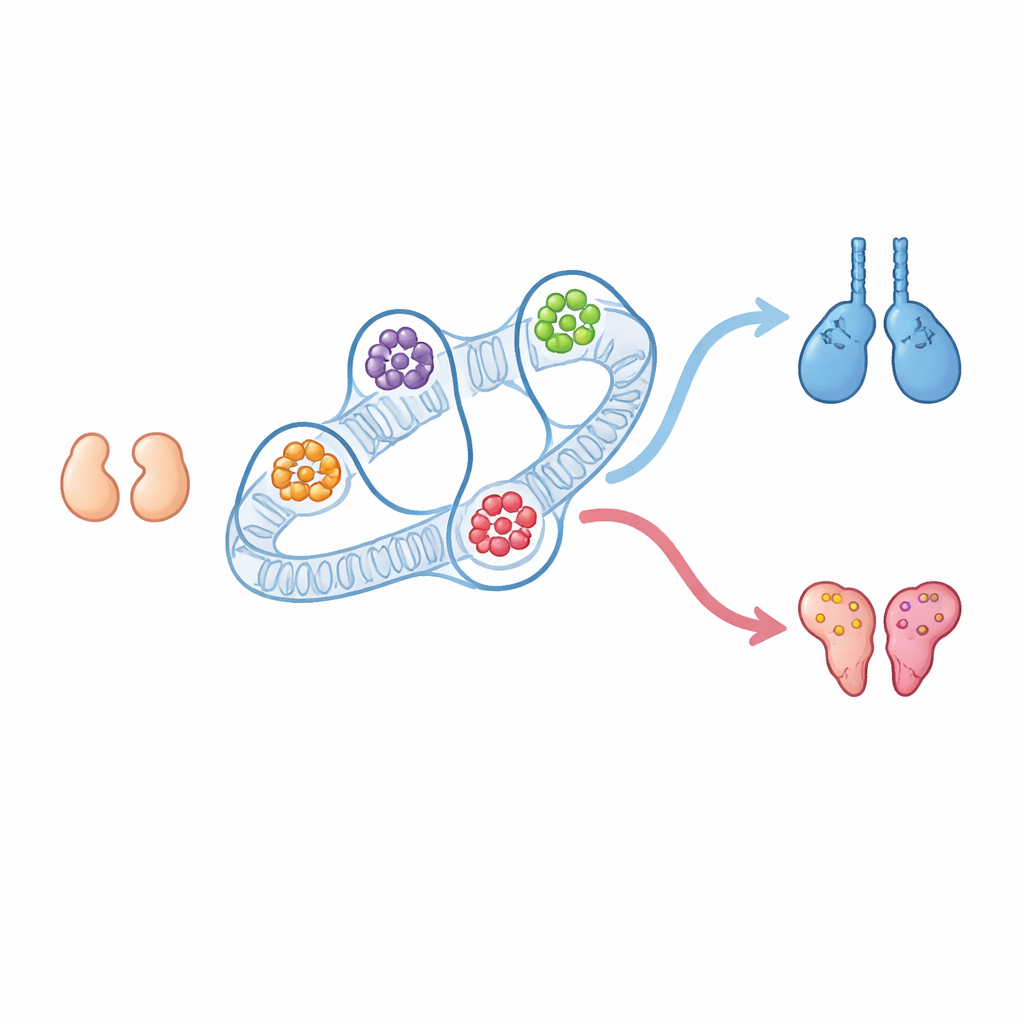
The Early Gonad at a Crossroads
In early mouse embryos, gonads in XX and XY individuals look and behave almost the same. They are “bipotential,” meaning they can still become either ovaries or testes. Later, a burst of activity from the Y-linked Sry gene in XY embryos triggers the testis pathway through the gene Sox9 and the signaling molecule Fgf9, while XX embryos, which lack Sry, turn on ovarian genes such as Wnt4 and Foxl2. The authors isolated specific supporting cells from these early gonads, both before and after sex is decided, and mapped how their DNA folds and which regions are active. Surprisingly, the broad chromosome neighborhoods that contact each other most often—known as compartments and domains—barely changed between the undetermined and sex-specific stages, even though gene activity shifts dramatically.
Discovering Hidden 3D Control Hubs
This mismatch between stable large-scale folding and highly dynamic gene activity suggested that the important action might be happening at finer scales. To uncover it, the team developed METALoci, a computational method that borrows tools from geography, where they are used to spot hotspots of, for example, pollution in a city. Instead of city blocks, METALoci treats small pieces of the genome as locations and uses Hi-C contact data to place them in a virtual map according to how often they touch. It then overlays chemical markers of activity, such as H3K27ac, to identify “hot” clusters where active enhancers and promoters gather in 3D space. These clusters, or metaloci, act as regulatory hubs in which groups of DNA switches and their target genes work together.
Rewiring During Sex Determination
Across the genome, METALoci revealed that these 3D hubs are extensively rewired as the gonads commit to ovary or testis. The number of strongly active hubs roughly doubled during differentiation, as did the number of strongly inactive zones, reflecting simultaneous turning on and off of entire gene programs. Known sex genes behaved in intuitive ways: for example, Sox9 gained a strong active hub in future Sertoli (testis) cells, while Bmp2 acquired an active environment in future granulosa (ovarian) cells. The authors tracked how each gene’s local environment moved between inactive and active states over time and between sexes, showing that male-specific genes typically experience particularly strong gains in regulatory activity during testis formation. 
A Noncoding Switch for a Key Sex Gene
One striking example came from Fgf9, a gene essential for promoting the testis pathway and blocking the ovarian one. People and mice with defects in FGF9 can undergo male-to-female sex reversal, yet the elements that control its activity in the gonad were unknown. Using METALoci, the researchers simulated deleting small pieces of DNA around Fgf9 in silico and asked which losses would disrupt its 3D regulatory hub. This pointed to a broad, gene-free stretch about a quarter of a million bases downstream. When the team removed a 306-kilobase chunk spanning most of this region in mice, Fgf9 expression in fetal testes dropped roughly by half. XY embryos developed gonads ranging from mixed ovotestes to ovary-like organs, closely mirroring complete Fgf9 knockouts—but without the lethal lung defects those knockouts usually cause. Smaller deletions showed that much of the control is concentrated in a central 93-kilobase subregion, yet regulatory power is shared among several enhancers, providing redundancy.
Shared Regulators of Male and Female Identity
To understand how these hubs connect into wider gene circuits, the authors combined their 3D maps with single-cell RNA data and reconstructed regulatory networks. They found known sex-determining factors at key positions, but also highlighted transcription factors not previously linked to sex determination. Among them were Meis1 and Meis2, which emerged as strong “negative regulators” of both male and female differentiation programs. Functional tests in genetically engineered mice showed that removing Meis1 alone, or three of the four Meis1/Meis2 gene copies, causes patches of cells in XY gonads to adopt ovarian-like identity and, conversely, some XX cells to adopt testis-like identity. This indicates that Meis genes act redundantly as guardians of proper sexual identity in both directions.
Why This Work Matters
For a non-specialist, the key message is that sex determination is not controlled only by which genes you have, but also by how your DNA is folded in 3D and how clusters of distant regulatory elements communicate with their target genes. METALoci reveals these hidden hubs and shows that, even when the overall genome scaffold looks stable, the internal wiring within domains can be dramatically rewired to shift cell fate. By pinpointing a powerful noncoding region that governs Fgf9 and uncovering new factors like the Meis genes, this work offers fresh leads for understanding disorders of sex development and demonstrates how 3D genome mapping can uncover crucial regulatory switches that traditional gene-focused approaches miss.
Citation: Mota-Gómez, I., Rodríguez, J.A., Dupont, S. et al. Chromatin spatial analysis by METALoci unveils sex-determining 3D regulatory hubs. Nat Struct Mol Biol 33, 577–589 (2026). https://doi.org/10.1038/s41594-026-01749-z
Keywords: sex determination, 3D genome, chromatin architecture, gene regulation, noncoding DNA