Clear Sky Science · en
CREsted: modeling genomic and synthetic cell-type-specific enhancers across tissues and species
Why tiny switches in DNA matter
Every cell in your body carries the same DNA, yet brain cells, blood cells and muscle cells behave very differently. A big reason is a hidden layer of control made of short DNA switches called enhancers, which decide when and where genes turn on. This article presents CREsted, a software toolkit that uses modern artificial intelligence to read these switches directly from DNA and even design new ones. The work shows how we can move from simply listing genetic parts to actively understanding and engineering them across tissues and species.
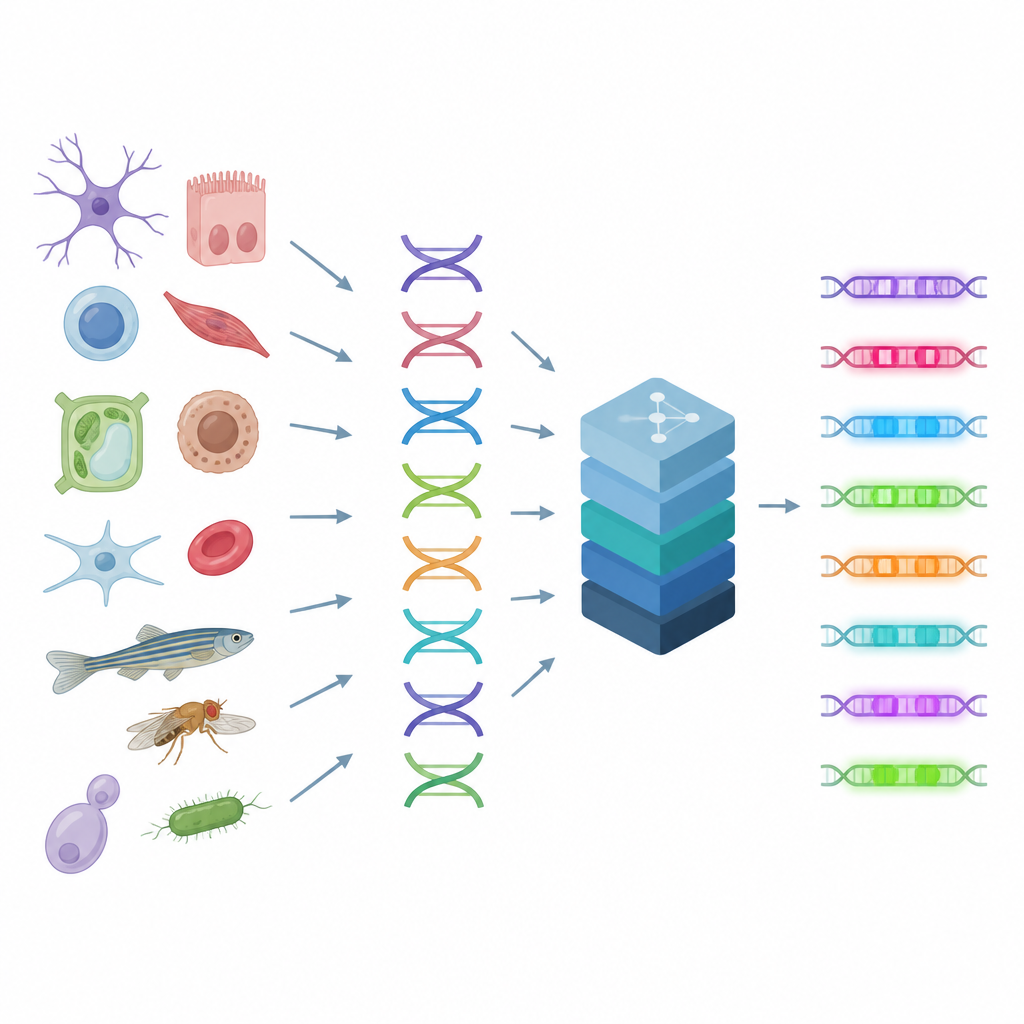
Reading the cell’s control switches
Enhancers act like knobs on a soundboard, combining signals from many proteins to fine tune gene activity in each cell type. Because multiple DNA patterns can produce similar outcomes, the rules behind enhancers are complex and hard to guess by eye. The authors build on a technique that measures how open or closed each stretch of DNA is in thousands of individual cells, a clue that reveals where active enhancers sit in the genome. CREsted takes these measurements, links them to the underlying DNA sequences and trains deep learning models to predict how accessible each region will be in many cell types at once. This turns raw sequence into a map of regulatory activity.
A toolkit from data to insight
CREsted is more than a single model: it is an end to end pipeline. It first cleans and reshapes single cell data into a form that reduces technical bias between cell types. Then it trains flexible neural networks that can either classify active regions or predict graded accessibility values. Importantly, CREsted does not stop at prediction. It can zoom in to identify which individual letters of DNA matter most for a given cell type, cluster recurring patterns and match these patterns to likely regulatory proteins using existing databases and gene expression data. Finally, it includes design tools that iteratively “evolve” synthetic DNA sequences so that the model predicts strong activity in one chosen cell type and little activity elsewhere.
Testing the toolkit in brain, blood, cancer and fish
The authors showcase CREsted on several rich datasets. In the mouse motor cortex, their models predict which regions of DNA are open in different neuron and support cell types with high accuracy and outperform a leading general purpose framework. By highlighting key sequence patterns, CREsted recovers known regulator proteins for specific neuron classes and can even explain how a single letter change in a motif may switch activity between neuron subtypes. In human blood cells, a related model rediscovers many previously tested binding sites in classic immune enhancers and aligns well with independent protein binding experiments, supporting that the learned sequence patterns are biologically meaningful.
CREsted also probes more applied questions. In cancer, it compares a “mesenchymal like” cell state that appears in both melanoma and glioblastoma, using models trained on cell lines and on patient tumor samples. The enhancer patterns show shared themes but also important differences, such as specific motifs present only in tumors. In another test, the authors ask whether specialized “foundation” models trained on vast genomic datasets truly outperform smaller, task focused models. After careful fine tuning, these large models still struggle to match the cell type specific resolution of CREsted’s own architecture, suggesting that dedicated training on high quality single cell data remains crucial.
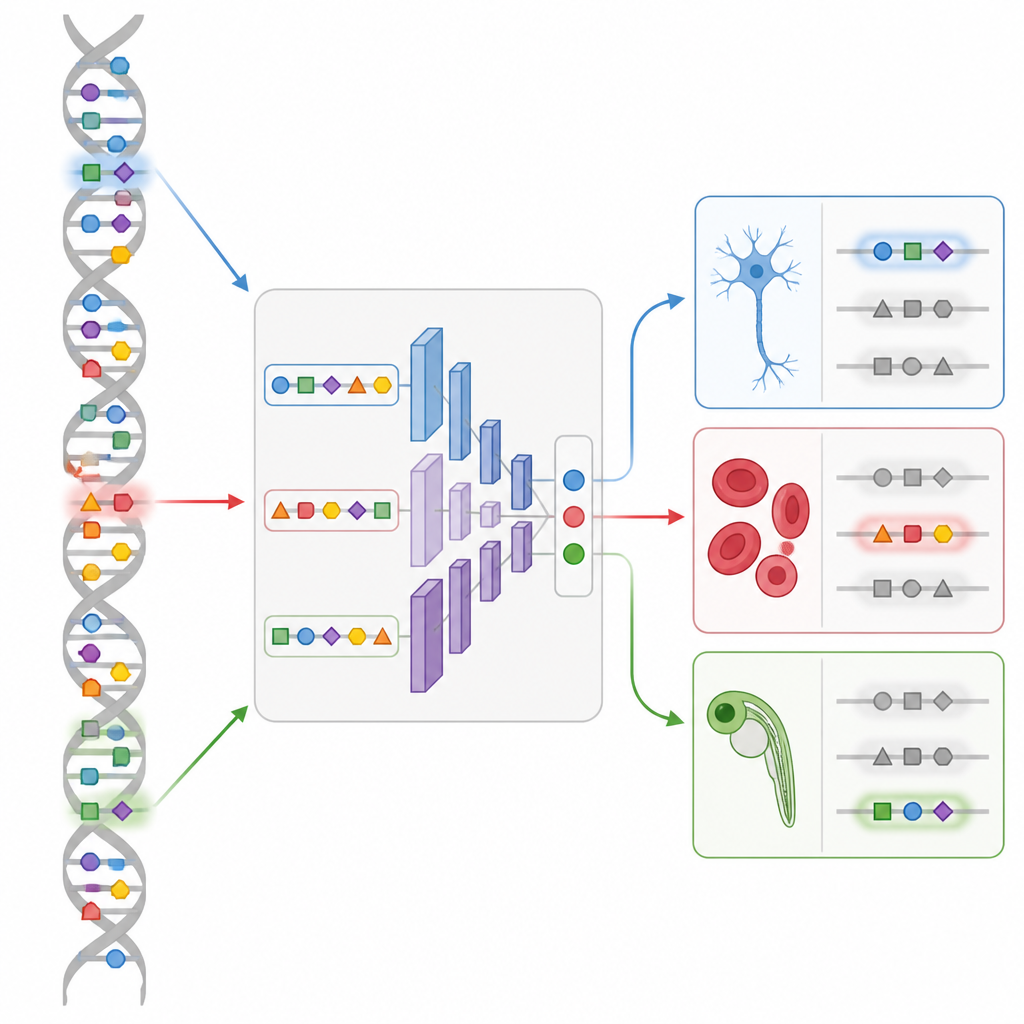
Designing new switches in a living embryo
The most striking demonstration comes from zebrafish development. Using a single cell map of DNA accessibility across many embryo stages, the team trains a CREsted model called DeepZebrafish. They then let the design module generate completely synthetic enhancers predicted to turn on only in heart muscle, only in body muscle, only in blood vessel lining, or in controlled combinations of heart and muscle. When these artificial sequences are placed in front of a fluorescent reporter and injected into fish eggs, many light up exactly in the intended tissues. Analysis of their sequences reveals recognizable patterns for major developmental regulators, confirming that the model has captured meaningful “grammar” rather than random quirks.
What this means for understanding and engineering genomes
To a non specialist, CREsted can be viewed as a microscope for the regulatory layer of the genome and a drafting tool for new genetic parts. It translates stretches of DNA into predictions about which cell types will use them, points to the key letters and motifs that drive that behavior and can suggest new sequences that should behave in a desired way. By working across brain, blood, cancer and whole developing animals, the study shows that a unified approach can reveal both shared and cell type specific rules of gene control. In the long run, such tools may help researchers build more precise genetic markers, improve disease models and better understand how small changes in DNA can ripple through cells and tissues.
Citation: Kempynck, N., De Winter, S., Blaauw, C.H. et al. CREsted: modeling genomic and synthetic cell-type-specific enhancers across tissues and species. Nat Methods 23, 946–959 (2026). https://doi.org/10.1038/s41592-026-03057-2
Keywords: enhancer modeling, deep learning genomics, single cell chromatin, cis regulatory elements, synthetic enhancers