Clear Sky Science · en
Chemical capture of diazo metabolites reveals biosynthetic hydrazone oxidation
Why unstable molecules matter to us
Chemists have long known that some of the most powerful medicines and industrial reagents are also the most fragile. They fall apart so quickly that finding them in nature can be like trying to catch soap bubbles in a storm. This study tackles one such family of fleeting molecules, called diazo compounds, which are made by microbes and can both damage cells and drive useful chemical reactions. By inventing a way to "catch" these unstable molecules as soon as they appear, the researchers uncover new chemistry in a human lung pathogen and reveal an enzyme that could become a green tool for manufacturing valuable reagents.
Hidden weapons from tiny microbes
Microorganisms are miniature chemical factories. Among the many compounds they make are diazo-containing natural products, which carry a pair of nitrogen atoms ready to burst apart and release energy. That reactivity lets them attack biological targets, giving rise to antibiotic and anticancer activity, and it also makes them prized reagents in synthetic chemistry labs. Yet only a few dozen natural diazo compounds are known. The team’s survey of genetic data uncovered hundreds of microbial gene clusters that look capable of making diazo molecules, scattered across soil bacteria, marine microbes and pathogens. The mismatch between the genetic potential and the small number of known compounds suggested that many diazo molecules remain unseen, likely because they are too unstable for standard discovery methods that rely on isolating stable, abundant products.
Catching fleeting molecules with a chemical hook
To solve this problem, the researchers flipped the usual search strategy. Instead of asking whether a microbe’s extracts kill cancer cells or bacteria, they asked whether its metabolites share a specific kind of chemical reactivity. They designed a small probe molecule based on a strained ring-shaped alkyne that reacts selectively and rapidly with diazo groups, forming a more stable ring called a pyrazole. When mixed with microbial broth, this probe acts like a chemical hook: if any diazo compound is present, the probe latches on and converts it into a heavier, more easily detected product. Using sensitive mass spectrometry and comparative metabolomics, the team compared samples taken immediately after adding the probe with samples incubated overnight. Signals that grew strongly over time and showed the characteristic doubled peak pattern of the probe products flagged hidden diazo metabolites.
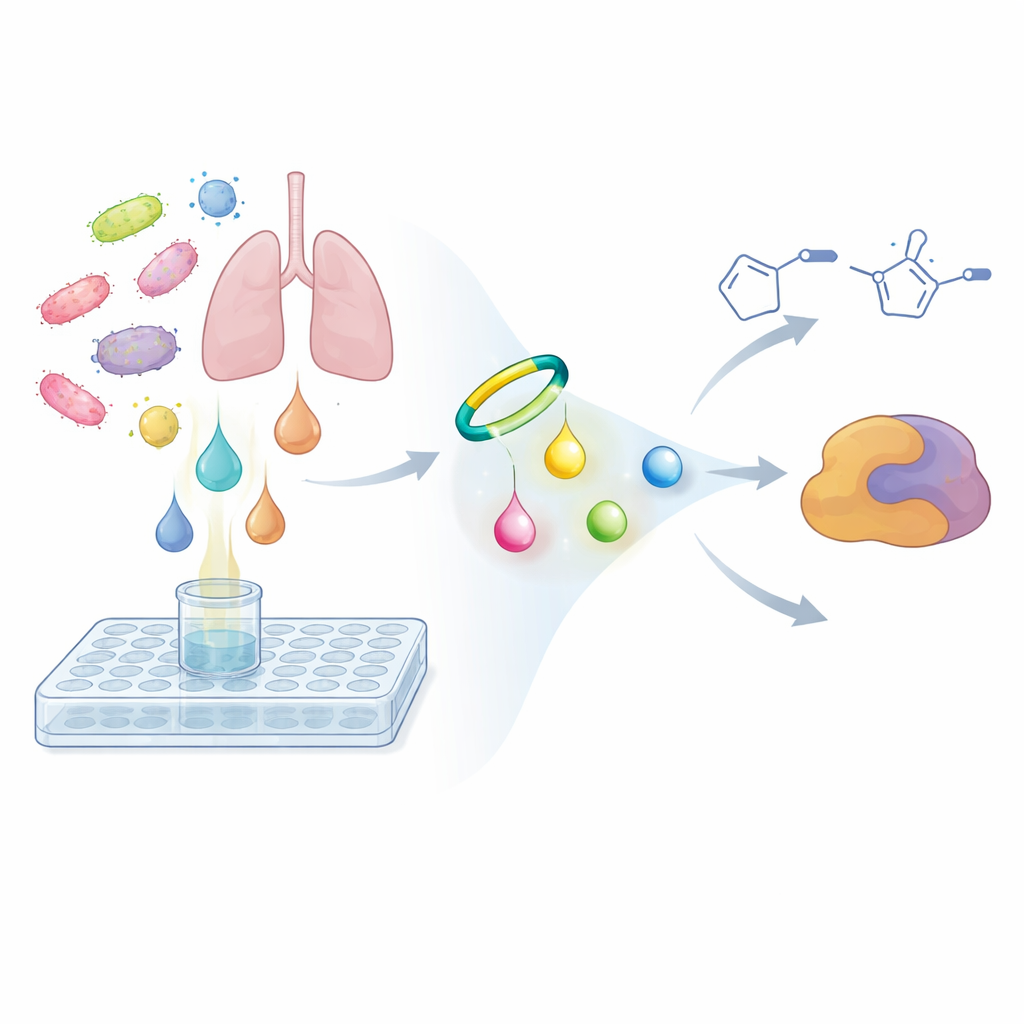
New reactive molecules from a lung pathogen
Screening genetically selected bacteria with this probe-based workflow led the team to the human lung pathogen Nocardia ninae. In its culture broth, they observed two pairs of probe-derived products that appeared only after incubation, had the expected nitrogen-rich composition, and matched the behavior expected for captured diazo compounds. By subtracting the known formula of the probe and reasoning through the remaining atoms, they deduced that the original natural products were two very simple yet previously unseen molecules: 4-diazo-3-oxobutanoic acid and diazoacetone. These are tiny building blocks compared with many natural products, but they are highly reactive. Control experiments showed that the unmodified molecules were essentially invisible to routine analysis, while the probe-labeled versions could be detected at concentrations as low as a few hundred parts per billion, underscoring how easily such metabolites could have been missed.
Uncovering an unusual enzyme pathway
Finding the molecules was only half the story; the researchers then traced how the bacterium makes them. By examining its genome, they pinpointed a cluster of genes, dubbed dob, that resembled known pathways for building nitrogen–rich units called hydrazones, but with extra enzymes hinting at new steps. By moving this gene cluster into a different bacterium and showing that the host began producing the same probe-detected products, they confirmed its role. Detailed biochemical experiments revealed a stepwise assembly line: common amino acids are reshaped into a hydrazone-bearing intermediate, which is extended by a polyketide synthase enzyme to form a small hydrazone acid. A second enzyme, a metalloenzyme named Dob3, then performs a rare chemical feat: it oxidizes the hydrazone into a diazo group. This reaction, called hydrazone oxidation, had been proposed but not directly demonstrated in nature.
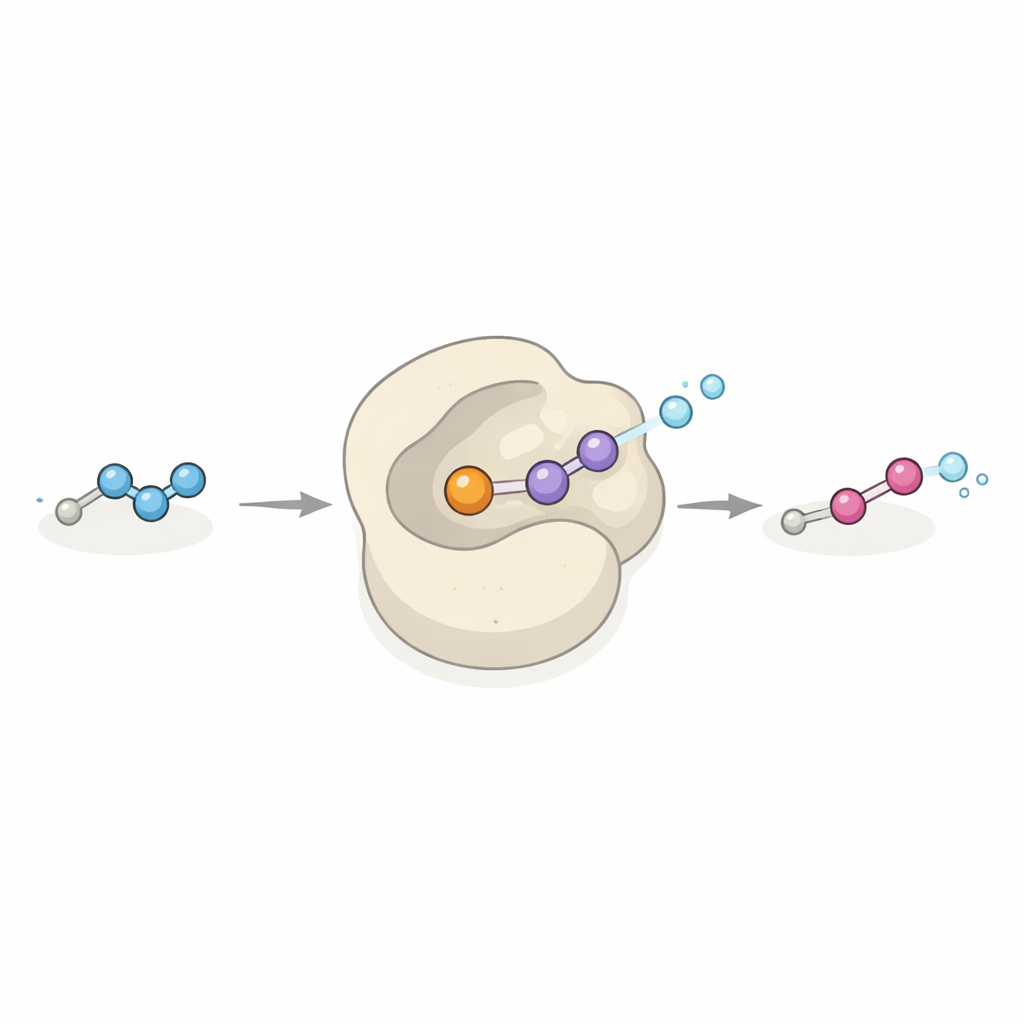
A new kind of catalyst with industrial promise
Dob3 belongs to a family of proteins that usually add oxygen atoms to carbon–hydrogen or nitrogen–hydrogen bonds using a pair of iron atoms and oxygen from the air. Here, Dob3 extends that playbook by turning a relatively stable hydrazone into a highly energized diazo group in a concerted two-electron oxidation step. The enzyme accepts a range of different hydrazone substrates in test-tube experiments, converting them into their corresponding diazo compounds. Because many diazo reagents are cornerstones of modern synthetic chemistry, especially for forging carbon–carbon bonds through so-called carbene transfer reactions, Dob3 could become a versatile “green” catalyst for making these reagents from simple precursors under mild, environmentally friendly conditions.
What this work means going forward
For non-specialists, the key message is that chemistry happening inside microbes is both richer and more delicate than we have been able to see. By using reactivity itself as a beacon—catching unstable molecules the moment they are made—this study reveals new diazo compounds inside a human pathogen and firmly establishes hydrazone oxidation as a natural route to these reactive groups. The discovery of Dob3, a diiron enzyme that performs this challenging transformation, opens doors to engineering microbes to make industrial diazo reagents sustainably. More broadly, the work suggests that many reactive natural products remain part of a microbial “dark matter” of chemistry, waiting to be uncovered with similarly clever, reactivity-guided search strategies.
Citation: Pfeifer, K., Van Cura, D., Wu, K.J.Y. et al. Chemical capture of diazo metabolites reveals biosynthetic hydrazone oxidation. Nature 652, 517–525 (2026). https://doi.org/10.1038/s41586-025-10079-x
Keywords: diazo natural products, reactivity-based screening, microbial metabolites, biocatalysis, enzyme discovery