Clear Sky Science · en
Continental-scale genomic surveillance of Plasmodium falciparum malaria across sub-Saharan Africa with rapid nanopore sequencing
Why malaria genes matter to everyday life
Malaria still kills hundreds of thousands of people each year in sub-Saharan Africa, and the parasite is learning new tricks to escape our tests and medicines. This study shows how scientists and local health teams can rapidly read the parasite’s genetic code in ordinary African laboratories, so countries can see in near real time where malaria is becoming harder to diagnose and treat.
Growing trouble for tests and treatments
The deadliest malaria parasite, Plasmodium falciparum, is pushing back against two of our main tools. Some strains have lost genes called hrp2 and hrp3, which means standard rapid tests can miss infections. Others carry changes in a gene known as kelch13, which are linked to slower clearance of parasites after treatment with today’s frontline drugs. These worrying variants have been spreading across eastern, central, and southern Africa. Health agencies are considering switching types of tests or rotating drug combinations, but they need detailed and timely information on which parasite strains are circulating where.
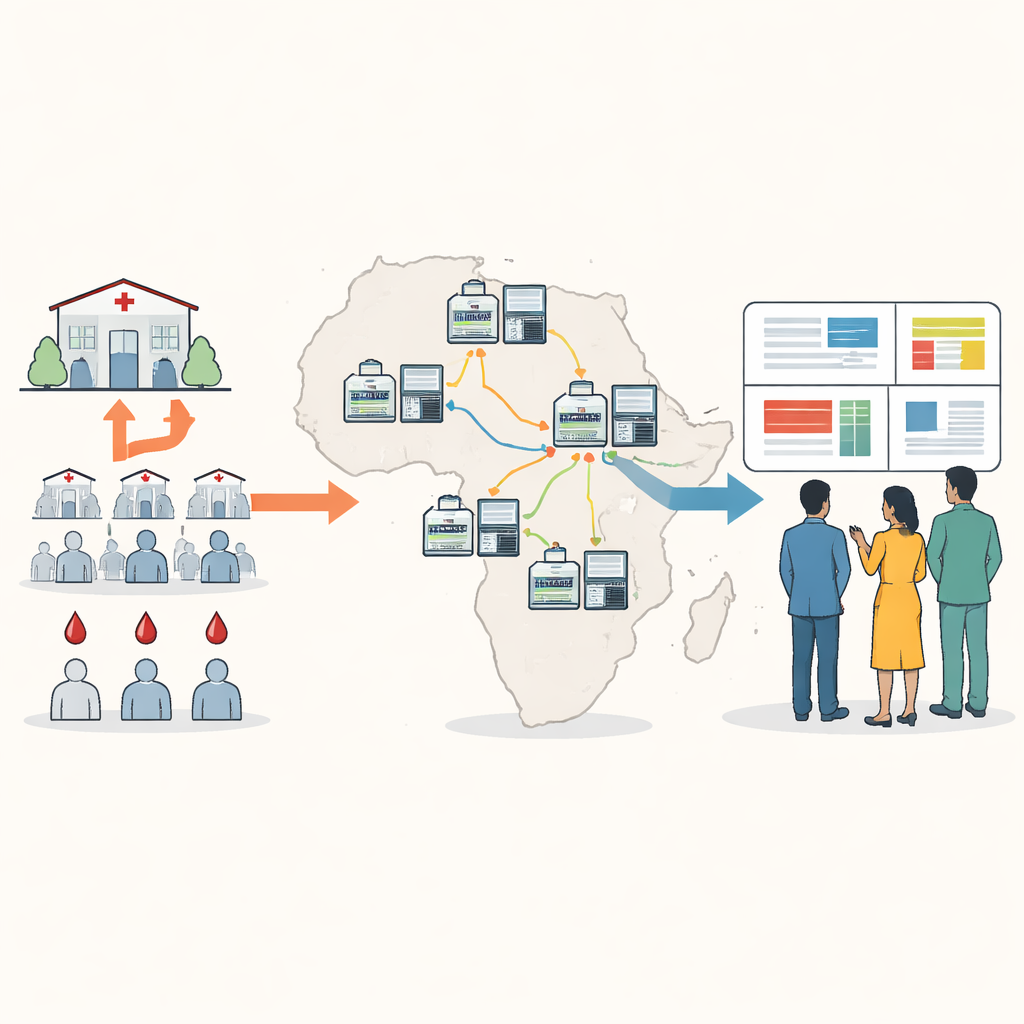
Bringing genetic tracking closer to the patients
Traditional DNA sequencing for malaria often requires shipping blood samples overseas or to a few big regional centers, which can be slow, expensive, and disconnected from local decision making. The authors designed a new approach built around a small portable device known as a nanopore sequencer. Using dried blood spots taken from routine malaria testing, local staff can amplify and sequence key parasite genes in about five hours, at a cost of under twenty-five dollars per sample. The new “minimal viable panel” focuses on ten genetic regions that reveal drug resistance, the presence or absence of hrp2 and hrp3, vaccine-relevant changes, and how many different parasite strains are mixed in a single infection.
Simple software on an ordinary laptop
To make the method usable in places with limited internet and computing power, the team created a software package called Nomadic. It runs on the same laptop that controls the sequencer and works entirely offline. As sequencing proceeds, the program maps each read to the parasite genome, checks quality, and flags important mutations, all while displaying intuitive charts in a local dashboard. The software also produces compact summary files that are easy to share by email or slow connections, allowing partners to combine results from many sites without moving large raw data files.
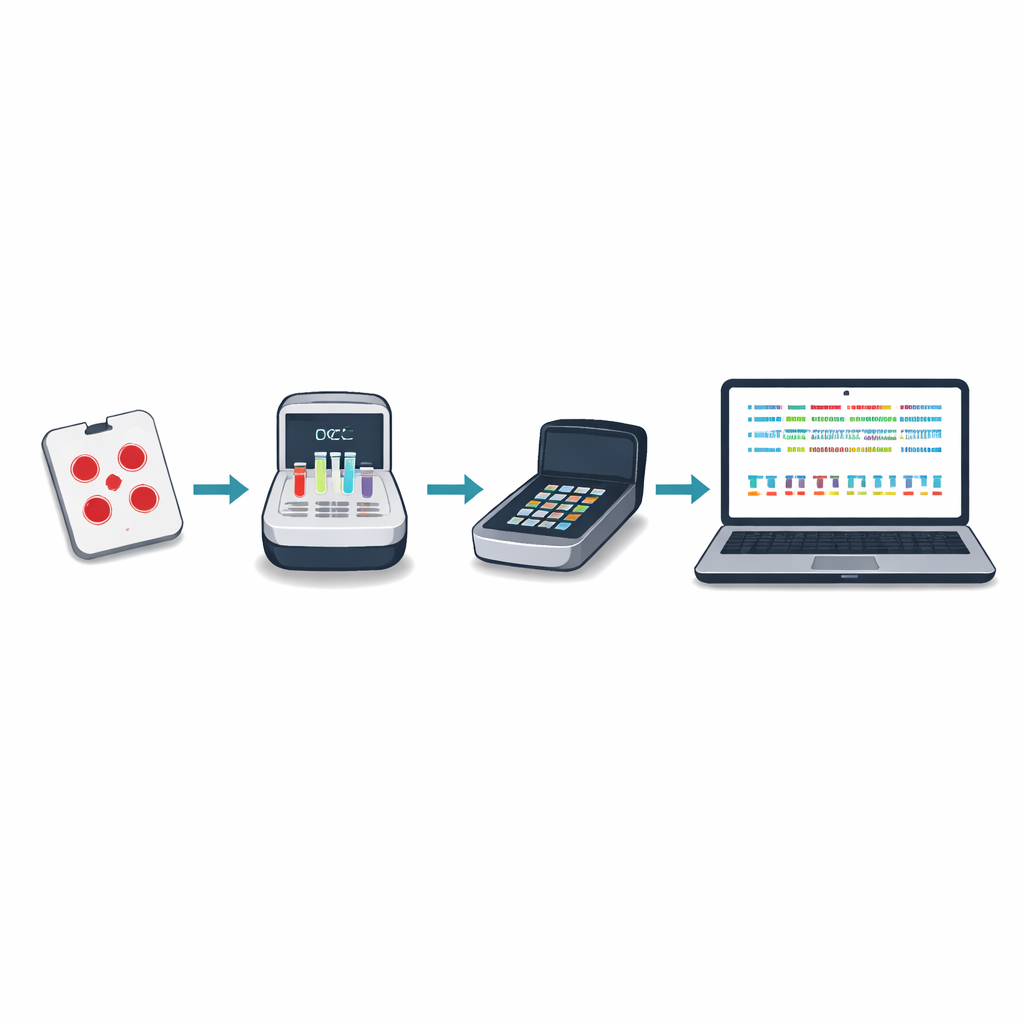
Testing the system across Africa
Between 2024 and 2025, laboratories in six African countries used the protocol to sequence and analyze over one thousand malaria samples, most of them processed locally by 13 scientists with varied levels of prior experience. The method produced strong and consistent coverage of the target genes across a wide range of parasite levels in blood. A new variant-calling tool, named Delve, could reliably detect genetic changes carried by minor parasite strains that made up as little as five to ten percent of an infection, a level standard tools often miss. The same sequencing runs also correctly identified almost all deletions in the hrp2 and hrp3 genes when compared with conventional laboratory methods, and in some mixed infections it detected deletions that standard tests overlooked.
What this means for the fight against malaria
This work shows that high-quality genetic surveillance of malaria does not have to be confined to distant high-tech centers. By pairing an affordable, streamlined laboratory protocol with laptop-based analysis, ordinary public health laboratories can track drug resistance markers and stealthy diagnostic-evading parasites in near real time. While other methods remain better suited for some specialist tasks, this approach is well matched to the urgent need for fast, practical information to guide test choices and treatment policies across sub-Saharan Africa.
Citation: Mwenda, M., Mosler, K., Bohmeier, B. et al. Continental-scale genomic surveillance of Plasmodium falciparum malaria across sub-Saharan Africa with rapid nanopore sequencing. Nat Commun 17, 4218 (2026). https://doi.org/10.1038/s41467-026-72358-z
Keywords: malaria, genomic surveillance, drug resistance, nanopore sequencing, sub-Saharan Africa