Clear Sky Science · en
Non-destructive transcriptomics via vesicular export
Watching Living Cells Without Breaking Them
Our cells are constantly turning genes on and off, and scientists track this activity by reading out RNA, the messenger molecules that carry genetic instructions. But there is a catch: nearly all current methods require breaking cells open, so each measurement destroys the very cells being studied. This paper introduces a way to eavesdrop on a cell’s RNA conversations over many days without harming the cells, opening the door to following how the same cells respond to drugs or mature into tissues like heart muscle or neurons.
Turning Cells Into Gentle Messengers
The researchers developed a platform they call non-destructive transcriptomics by vesicular export, or NTVE. Instead of cracking cells open, NTVE persuades them to regularly send out tiny bubbles, or vesicles, that safely carry samples of their RNA into the surrounding fluid. These vesicles are built using a stripped-down form of an HIV protein shell that naturally buds off from the cell surface. By adding molecular “handles” inside this shell, the team makes sure that RNA molecules are pulled into the budding vesicles, and by adding distinct markers on the vesicle surface, they can later separate vesicles from different cell types that are grown together. 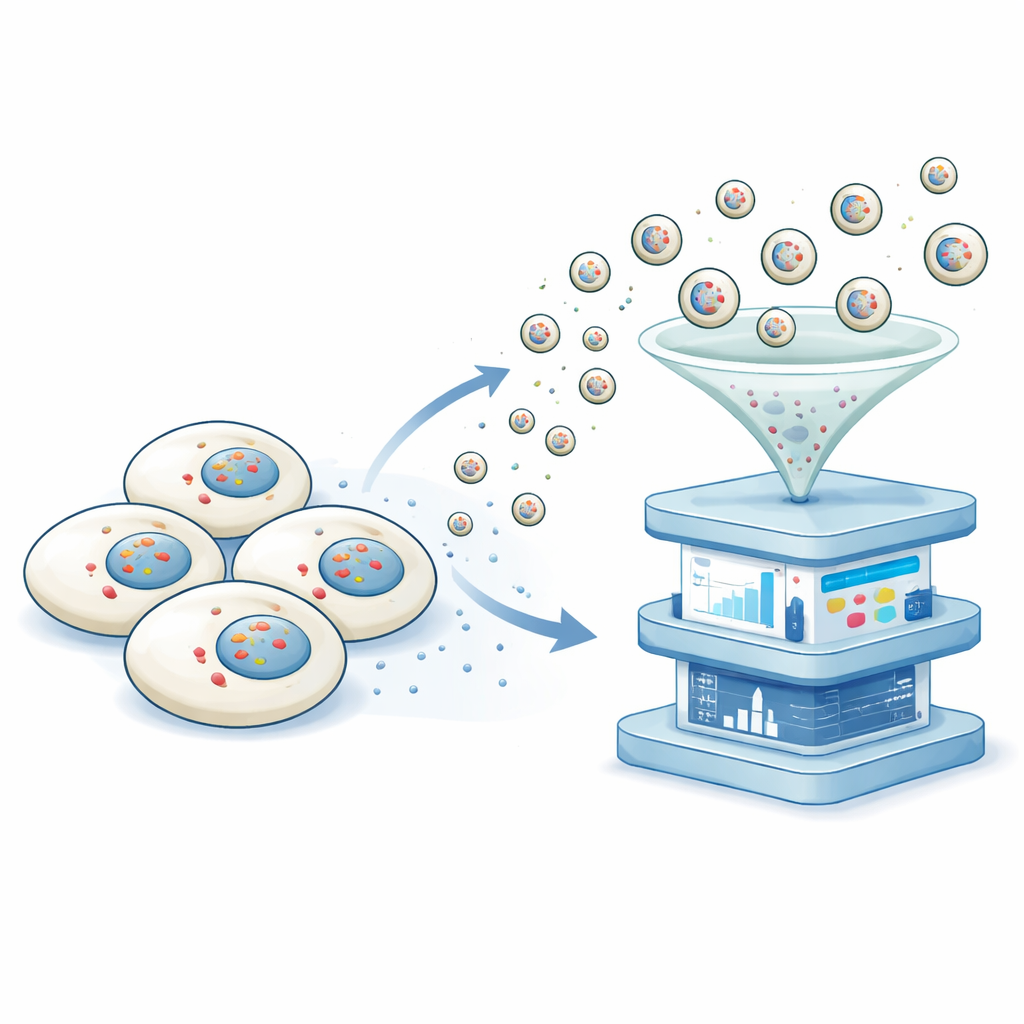
Catching the Full RNA Picture
A key challenge is making sure these exported vesicles truly reflect what is going on inside the cell, rather than just grabbing a biased subset of RNAs or leaking damaged material. To solve this, the authors fused part of a normal cellular protein that binds to the tail of most messenger RNAs onto the inside of the vesicle shell. This design draws in a broad and balanced collection of RNA messages from near the cell surface. By comparing RNA from vesicles with RNA from traditional destructive sampling of the same cells, they showed very strong agreement across more than 14,000 genes. Importantly, mitochondrial RNAs, which might indicate membrane damage and leakage, were strongly depleted, suggesting that the cells remain intact and healthy while exporting RNA.
Following Cells Through Change
With NTVE in place, the team tested whether it could track how cells respond over time to specific triggers. In human cells, NTVE correctly reported the activation of a single “switched on” gene driven by a CRISPR-based activator, and it closely matched conventional methods when cells were exposed to the immune signal interferon-gamma, capturing the same pathways and response genes. The researchers then moved to more delicate systems: primary mouse neurons and human stem cells. By packaging the NTVE machinery in viral vectors or transposons, they created neuron and stem cell lines that could be induced with a simple drug to start vesicle production. In neurons, NTVE captured the wave of gene activity that follows stimulation with forskolin, a compound known to activate memory-related signaling.
Watching Stem Cells Become Tissues
Because NTVE does not kill cells, it is especially powerful for watching slow, multi-day processes like stem cell differentiation. The authors engineered human induced pluripotent stem cells to express NTVE and then pushed them along three developmental paths: toward nerve-like, muscle-like, and gut-like lineages. By sampling vesicle RNA every day, they reconstructed the branching trajectories of these cell populations and evaluated which marker genes best distinguished each lineage. In a separate experiment, they tracked stem cells maturing into beating heart cells over nine days. Daily NTVE measurements captured the rise and fall of key cardiac genes and showed that the surge in heart-specific markers coincided with the first visible contractions. 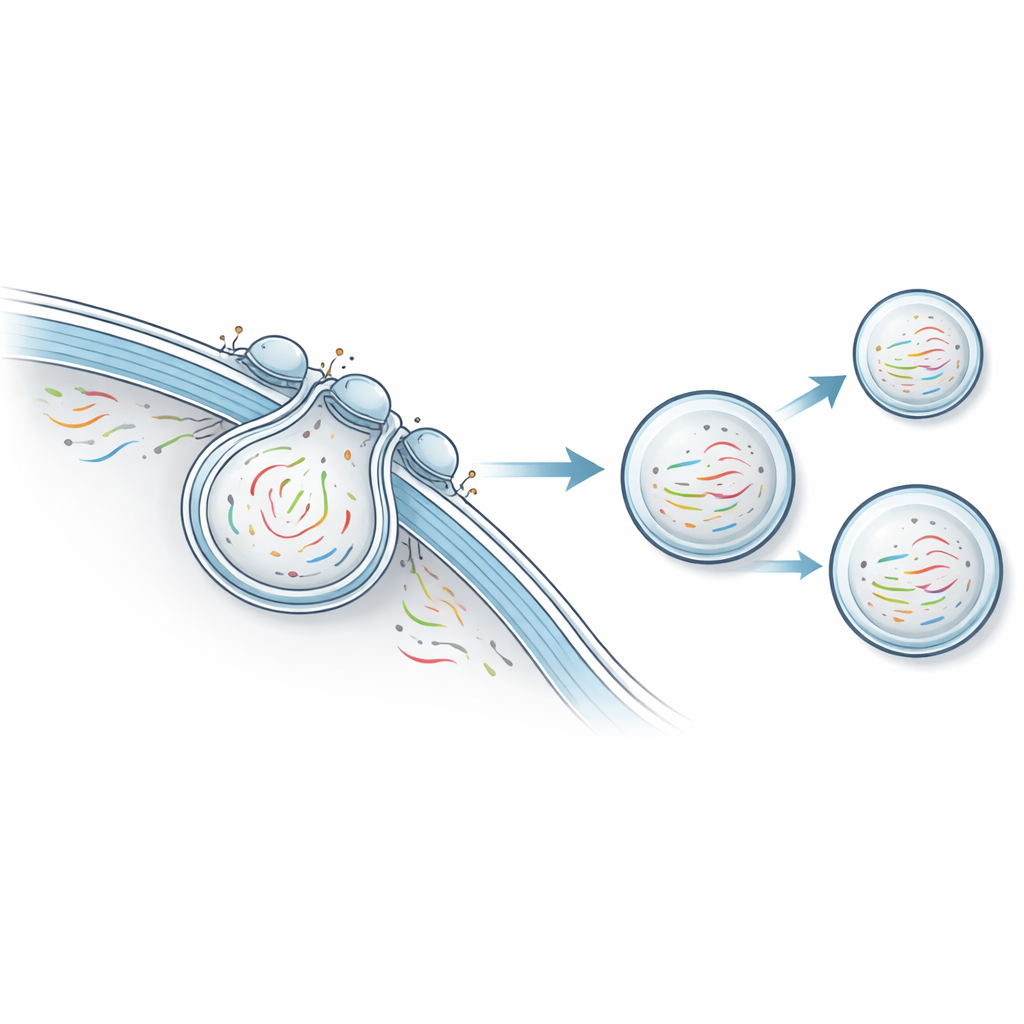
Sending Messages Between Cells
Beyond passive monitoring, NTVE vesicles can also be turned into delivery vehicles. The team demonstrated that vesicles could be loaded with messenger RNAs or gene-editing protein complexes and directed to specific “receiver” cells using viral coat proteins as targeting devices. In co-cultured cells, this allowed one population to send vesicles that flipped genetic switches or rewrote DNA in another population, creating a programmable channel for cell-to-cell communication that rides on the same vesicle platform used for RNA readout.
Why This Matters for Future Research
For non-specialists, the key idea is that NTVE lets scientists repeatedly take “liquid biopsies” of living cells in a dish, reading their gene activity day after day without sacrificing the cells. This overcomes a major limitation of standard RNA profiling, where each time point requires a new batch of cells and obscures how individual cells evolve. With NTVE, each culture—or even each organoid—can serve as its own baseline, improving the power to detect subtle changes and making long-term studies of development, disease models, and drug responses more precise. Although the method currently focuses on cytoplasmic messenger RNAs and cannot yet assign vesicle RNAs to single cells, it already offers an accessible and versatile way to watch the molecular life of cells unfold in real time.
Citation: Armbrust, N., Grosshauser, M., Geilenkeuser, J. et al. Non-destructive transcriptomics via vesicular export. Nat Commun 17, 3812 (2026). https://doi.org/10.1038/s41467-026-72072-w
Keywords: RNA export, live-cell transcriptomics, extracellular vesicles, stem cell differentiation, gene expression dynamics