Clear Sky Science · en
Dbf4-dependent kinase finetunes Ino80 function at chromosome replication origins
Why this matters for our DNA
Every time a cell divides, it must copy its entire DNA without major mistakes. Failures in this copying step can destabilize the genome and are closely linked to cancer. This study uncovers how a key cell-cycle enzyme not only switches on the DNA copying machinery, but also tidies up the local packaging of DNA so that replication can start smoothly and restart under stress. 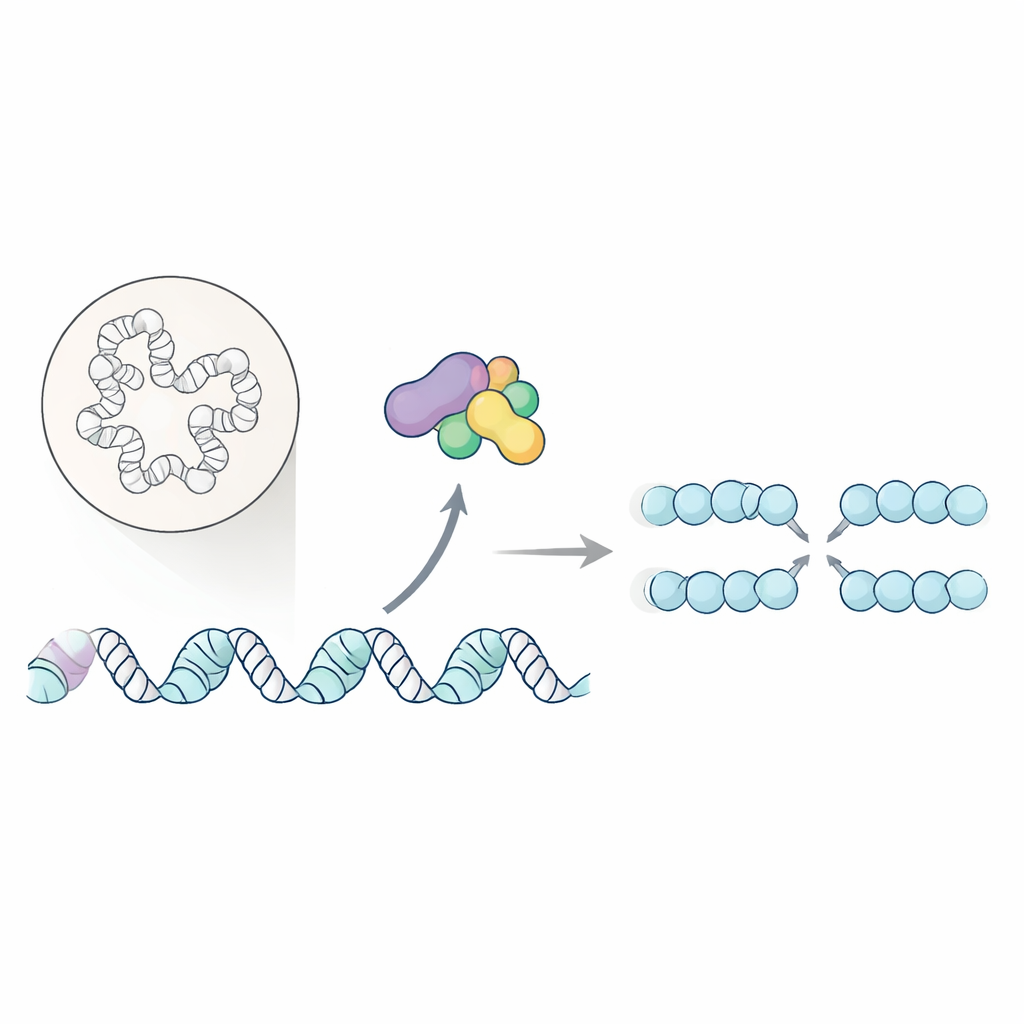
A traffic light for starting DNA copying
Cells rely on specialized "origins" along their chromosomes where DNA replication begins. A well-known enzyme pair called Dbf4-dependent kinase (DDK) helps start this process by modifying the core DNA-unwinding machine, the helicase. Until now, only a few direct targets of DDK were known, leaving much of its role mysterious. The authors set out to chart, on a global scale, which nuclear proteins depend on DDK for phosphorylation—a chemical tag that often serves as an on/off switch for protein function.
Scanning the nucleus for DDK’s hidden partners
To do this, the researchers used budding yeast as a model and combined two complementary strategies to turn down DDK activity in living cells: a temperature-sensitive mutant of its catalytic subunit and a drug-induced replication stress that indirectly blocks DDK’s action. They then isolated nuclei and used mass spectrometry to catalog thousands of phosphorylated protein fragments, comparing samples with active versus inhibited DDK. By focusing on sites that decreased in both inhibition setups, they built a high-confidence list of around 400 DDK-dependent targets. Many of these proteins are bound to chromatin, the DNA–protein material that organizes the genome, hinting that DDK has a broader role in shaping the physical landscape where replication occurs.
A remodeling machine tuned by a small protein tail
Among the strongest hits was Arp8, part of a large chromatin remodeling machine called the INO80 complex. INO80 is known to reposition nucleosomes—bead-like structures of DNA wrapped around histone proteins—and to arrange them into regular arrays around replication origins. The team showed biochemically that DDK directly phosphorylates two specific serines in an unstructured tail of Arp8. When they replaced these serines with alanines, which cannot be phosphorylated, the overall composition of INO80 stayed intact, but its internal architecture changed: cross-linking experiments revealed weakened contacts between key modules of the complex. Functionally, the mutant INO80 burned ATP much more slowly and was markedly worse at sliding nucleosomes into new positions, even though it could still bind DNA and nucleosomes, sometimes even more tightly than normal.
From local spacing to global replication success
To connect these molecular defects to chromosome behavior, the authors rebuilt chromatin in vitro on a library of yeast DNA fragments containing hundreds of replication origins. With normal INO80 and the origin recognition complex present, they observed neatly phased nucleosome arrays flanking a nucleosome-free gap at each origin. In contrast, the Arp8 mutant produced less precise arrays with altered linker distances between nucleosomes. Using a fully reconstituted replication system, they found that chromatin assembled with mutant INO80 supported weaker initiation at origins, even though the growth of individual replication tracts that did start looked similar. In live yeast carrying the same Arp8 mutations, entry into DNA synthesis after cell-cycle arrest was delayed, spontaneous recombination events increased, and cells were especially sensitive to hydroxyurea, a drug that depletes DNA building blocks and stalls replication. Yet global gene expression, including that of cell-cycle and replication genes, was largely unchanged, arguing that the main problem lies in chromatin architecture at origins rather than in how many replication factors are made. 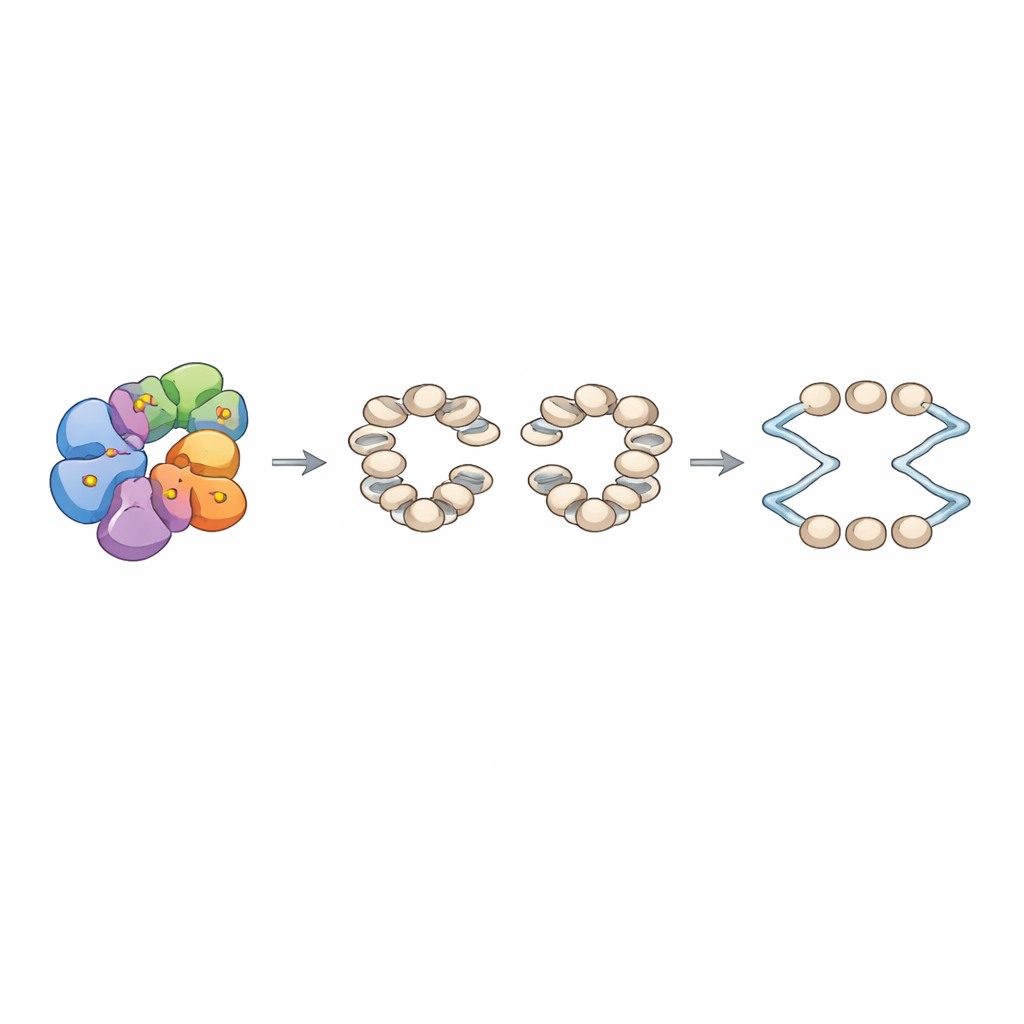
How careful DNA packing safeguards our genomes
Taken together, the work reveals that DDK does more than flick the switch on the core replication machinery. By phosphorylating Arp8, it finetunes the INO80 remodeler so that nucleosomes around replication origins are spaced just right. This orderly packing appears to be a gatekeeper for efficient origin firing and for safely restarting stalled replication forks under stress. In practical terms, the study shows that the way DNA is wrapped and arranged is not just an obstacle that replication must overcome—it is an active, regulated feature that helps determine when and where replication begins, with direct implications for maintaining genome stability.
Citation: Bansal, P., Lahiri, S., Kumar, C.N. et al. Dbf4-dependent kinase finetunes Ino80 function at chromosome replication origins. Nat Commun 17, 3029 (2026). https://doi.org/10.1038/s41467-026-70698-4
Keywords: DNA replication, chromatin remodeling, cell cycle kinase, genome stability, yeast model