Clear Sky Science · en
Trimbody with rigid AI-designed scaffolds enables atomic-resolution cryo-EM structure determination of small proteins
Seeing the Smallest Building Blocks
Proteins are the tiny machines that keep our cells alive, but many of them are too small to be seen clearly with today’s most powerful imaging tools. This paper introduces “Trimbody,” a clever protein-based helper that makes these tiny targets look bigger and more rigid under a cryo-electron microscope, allowing scientists to see them at near-atomic detail. The method promises to speed up basic biology and the development of nanobody-based medicines, especially for drug targets that were previously too small or too flexible to image well.
Why Small Proteins Are Hard to See
Cryo-electron microscopy (cryo-EM) has revolutionized structural biology by letting researchers visualize large biological molecules in a near-native frozen state. However, when proteins get smaller than about 50–100 kilodaltons (a common size unit for proteins), they become difficult to distinguish from noise in the images. Their outlines are faint, and they lack striking features that computer programs need to assemble sharp three-dimensional models. Existing strategies try to “bulk up” these small proteins by attaching them to larger partners, but many of these systems are complicated to build, somewhat floppy, or work only for a narrow set of targets. Scientists have needed a simple, robust way to make small proteins look big, bright, and rock-solid for cryo-EM.
A Two-Part Helper That Acts Like a Rigid Handle
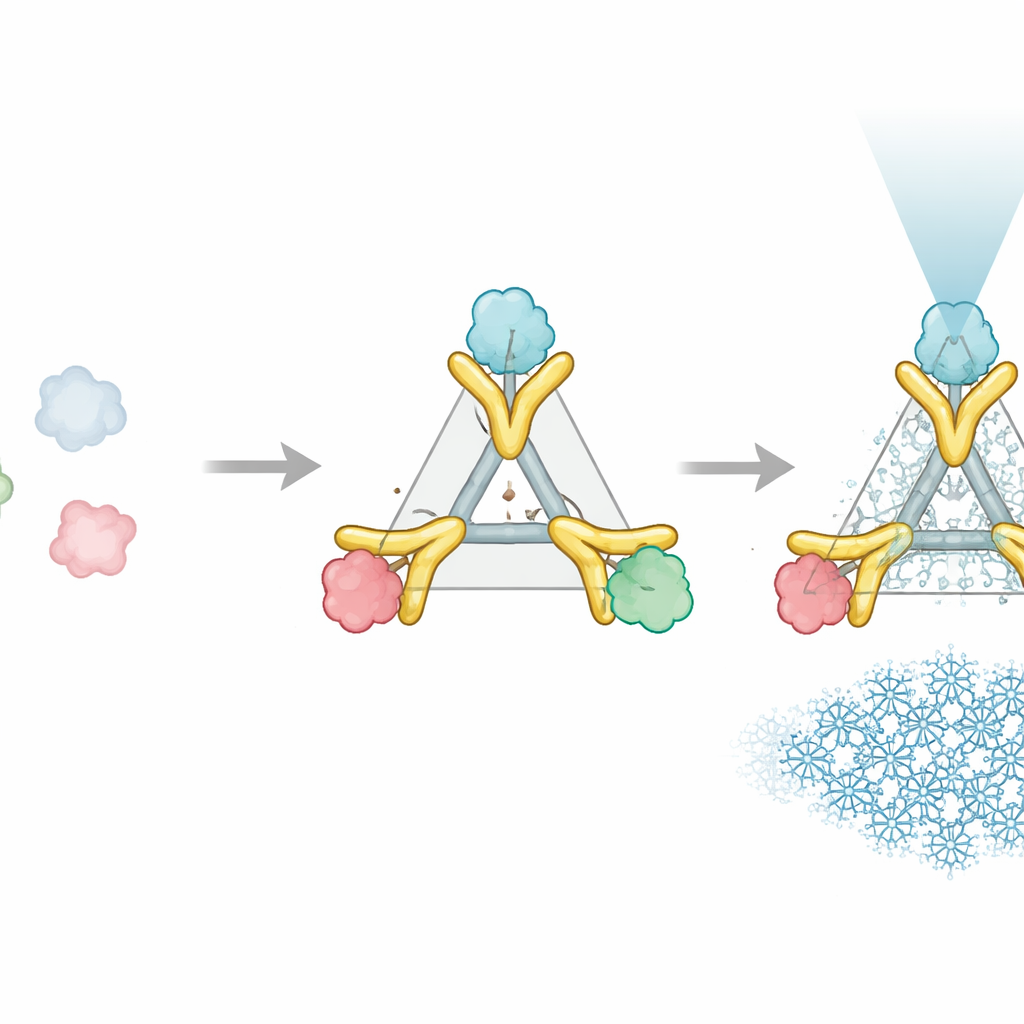
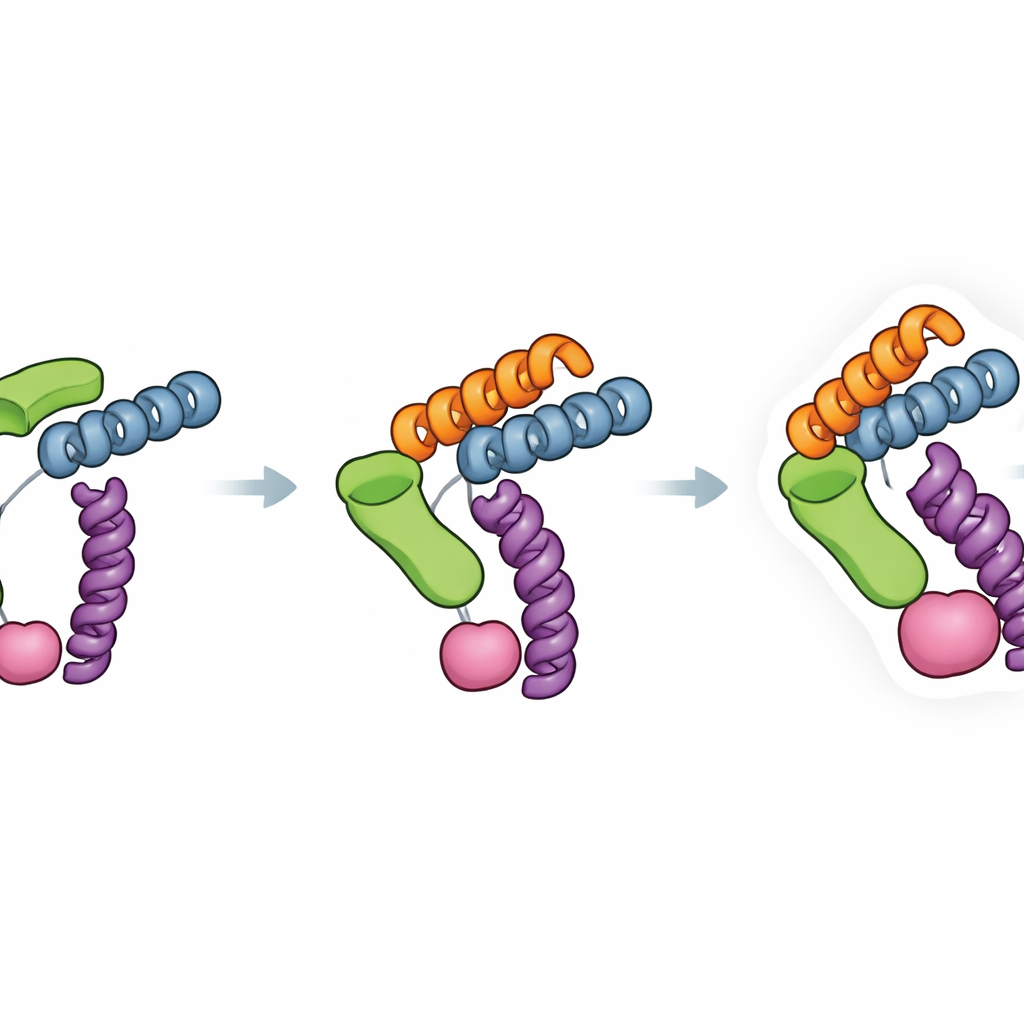
The Trimbody system solves this problem with just two engineered parts that snap together around the protein of interest. The first part is a trimeric scaffold—a three-armed protein assembly that forms a stable, rigid core. The second part is a customized nanobody, a tiny antibody fragment that binds tightly to a chosen protein target. This nanobody is extended with a specially designed “TAIL” domain that grips the scaffold, while an AI-designed “H3” helical bundle stabilizes the scaffold itself from the inside. Together, these additions act like braces and crossbars, locking the nanobody and scaffold into a single stiff unit. When three nanobody–target pairs attach to the three arms of the scaffold, the formerly tiny protein now behaves as part of a large, symmetric, and easily recognized object in the microscope.
Putting Trimbody to the Test
To see whether Trimbody works in practice, the authors applied it to several very different small proteins. They imaged a human immune-related protein (Galectin-10), a green fluorescent protein often used as a laboratory marker, the front-end domain of the cancer-linked molecule Nectin-4, and a bacterial membrane transporter that moves lactose across cell membranes. In each case, the Trimbody complex formed clean, well-behaved particles on cryo-EM grids and produced highly detailed three-dimensional reconstructions with resolutions around 2.3–2.6 angstroms—fine enough to distinguish most individual side chains of amino acids. Focused computational refinement on just the nanobody–target region yielded even clearer views, revealing subtle features like flexible loops, buried chemical groups, and binding surfaces that were previously hard to capture. Importantly, for proteins with existing crystal structures, the Trimbody-based cryo-EM models matched them closely, demonstrating that the helper scaffold does not distort the natural shape of the target.
Designed Rigidity and Broad Compatibility
A key strength of Trimbody lies in how deliberately its rigidity was engineered. The H3 domain forms a three-way helical brace that locks the projecting binding module to the trimeric base, while the TAIL domain and its four-helix linker create a second anchor between the nanobody and scaffold. Measurements of binding strength showed that this dual anchoring improved the nanobody–scaffold attachment by about four orders of magnitude, greatly reducing unwanted motion that can blur cryo-EM images. The authors also analyzed many existing nanobody–protein structures to check how often Trimbody would run into steric clashes—situations where the scaffold would bump into the protein itself. Their survey suggests that, for most nanobodies and targets, the system can be made compatible by adjusting a small set of framework residues and testing complex formation with standard biochemical assays.
Opening a Clearer Window on Tiny Targets
In everyday terms, Trimbody acts like a precision-crafted handle that lets microscopes “grab” and align very small proteins without shaking. Because it relies on just two fusion proteins that are straightforward to produce in bacteria, the approach is both cost-effective and accessible to many laboratories. By turning a wide variety of nanobody–target pairs into stiff, cryo-EM–friendly complexes, Trimbody offers a general route to atomic-level views of proteins that were previously too small or too elusive to study. This capability can accelerate the understanding of disease-related proteins and support faster, structure-guided development of nanobody-based drugs and diagnostics.
Citation: Song, J., Qi, L., Li, Y. et al. Trimbody with rigid AI-designed scaffolds enables atomic-resolution cryo-EM structure determination of small proteins. Nat Commun 17, 3135 (2026). https://doi.org/10.1038/s41467-026-69941-9
Keywords: cryo-electron microscopy, nanobody scaffolds, protein structure, AI-designed proteins, small membrane proteins