Clear Sky Science · en
Delineation of the heterogeneity underlying genomic instability in hereditary breast cancers reveals four disease subtypes
Why this research matters for families
Many women who develop breast cancer have a strong family history of the disease but no mutation in the well‑known BRCA1 or BRCA2 genes. Doctors know these tumors often behave differently and can respond in special ways to treatment, yet the reasons have been hazy. This study uses whole‑genome analysis to show that hereditary breast cancers without BRCA1/2 mutations are not one single entity but fall into four clear genetic groups, each with its own pattern of DNA damage and potential treatment vulnerabilities. Understanding these hidden differences could help tailor therapies far more precisely for patients and their relatives.
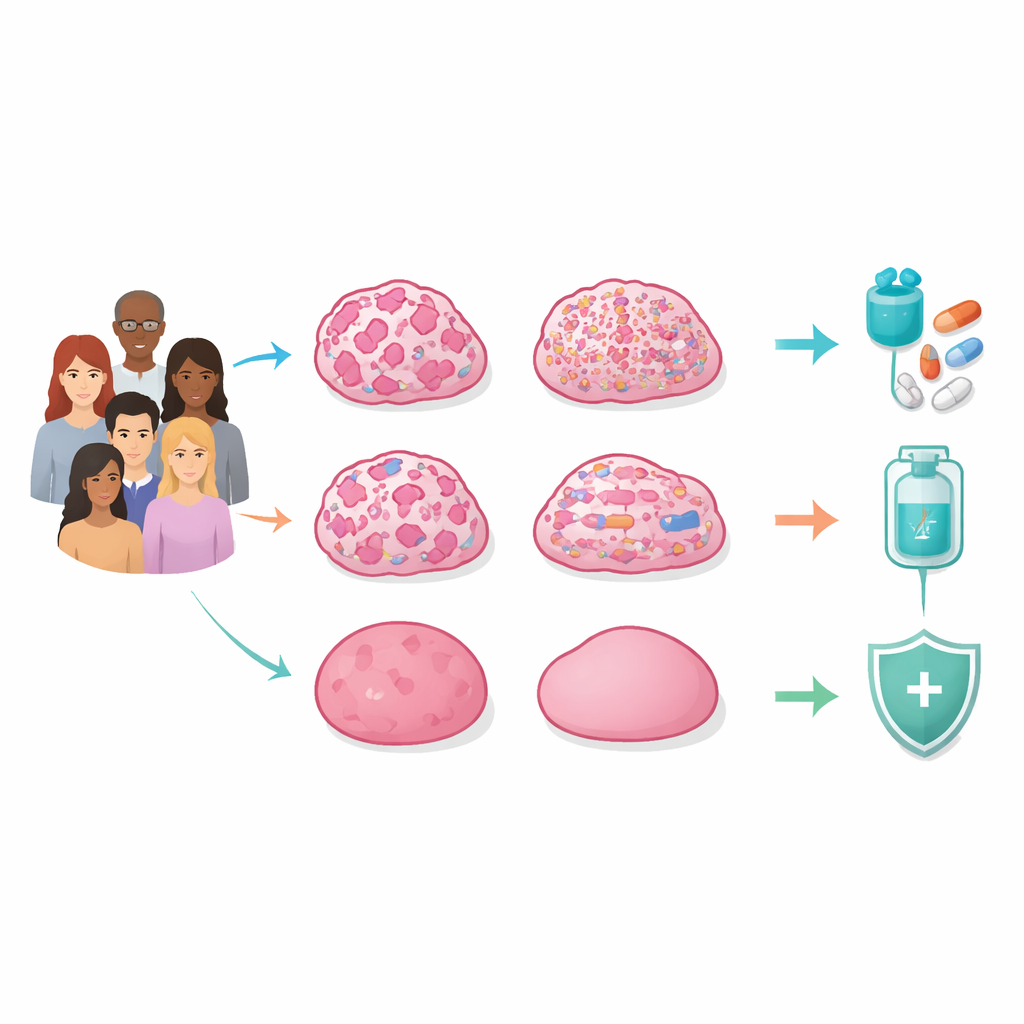
Four hidden patterns inside inherited tumors
The researchers sequenced the entire genomes of tumors and normal tissue from 129 Korean women with suspected hereditary breast cancer who tested negative for harmful BRCA1/2 variants. They looked not just at individual gene mutations, but also at large and small breaks, losses, gains, and rearrangements across the chromosomes, and at chemical tags on DNA and gene activity. By integrating all these layers, they found that the central feature distinguishing tumors was not a single gene, but the overall pattern of genomic instability—how damaged and disordered the DNA had become.
DNA repair failure, mutation storms, and quiet genomes
One major group, called the homologous recombination–deficient (HRD) subtype, showed widespread “chromosomal scars” caused by failure of a key DNA repair system. Even without inherited BRCA1/2 mutations, many of these tumors had lost BRCA function through loss of the remaining copy or by silencing the gene’s promoter with extra methylation. Their chromosomes were heavily rearranged, often with catastrophic shattering events, and frequently carried TP53 mutations. These tumors tended to be triple‑negative breast cancers and to have poorer survival, but they also appeared especially vulnerable to drugs that exploit DNA repair weakness, such as PARP inhibitors.
Mutation‑heavy and copy‑number‑heavy cancers
A second group, the mutation‑dominant (MUT) subtype, had comparatively low HRD scores but carried unusually high numbers of point mutations. Many of these changes bore the hallmark of APOBEC enzymes, which can trigger local “storms” of mutations known as kataegis. MUT tumors showed strong signals of immune activation, including genes used by killer T cells and markers of pro‑inflammatory macrophages, suggesting that they may be particularly good candidates for immunotherapy, even though many would be classified as hormone‑receptor‑positive by standard tests. A third group, the copy‑number–dominant (CN) subtype, instead accumulated many small DNA losses and gains while keeping an overall near‑normal chromosome count. These focal changes often removed classic tumor‑suppressor genes and were accompanied by an activated, fibroblast‑rich tumor stroma, hinting at sensitivity to platinum or taxane chemotherapy, potentially in combination with drugs that target the supporting tissue.
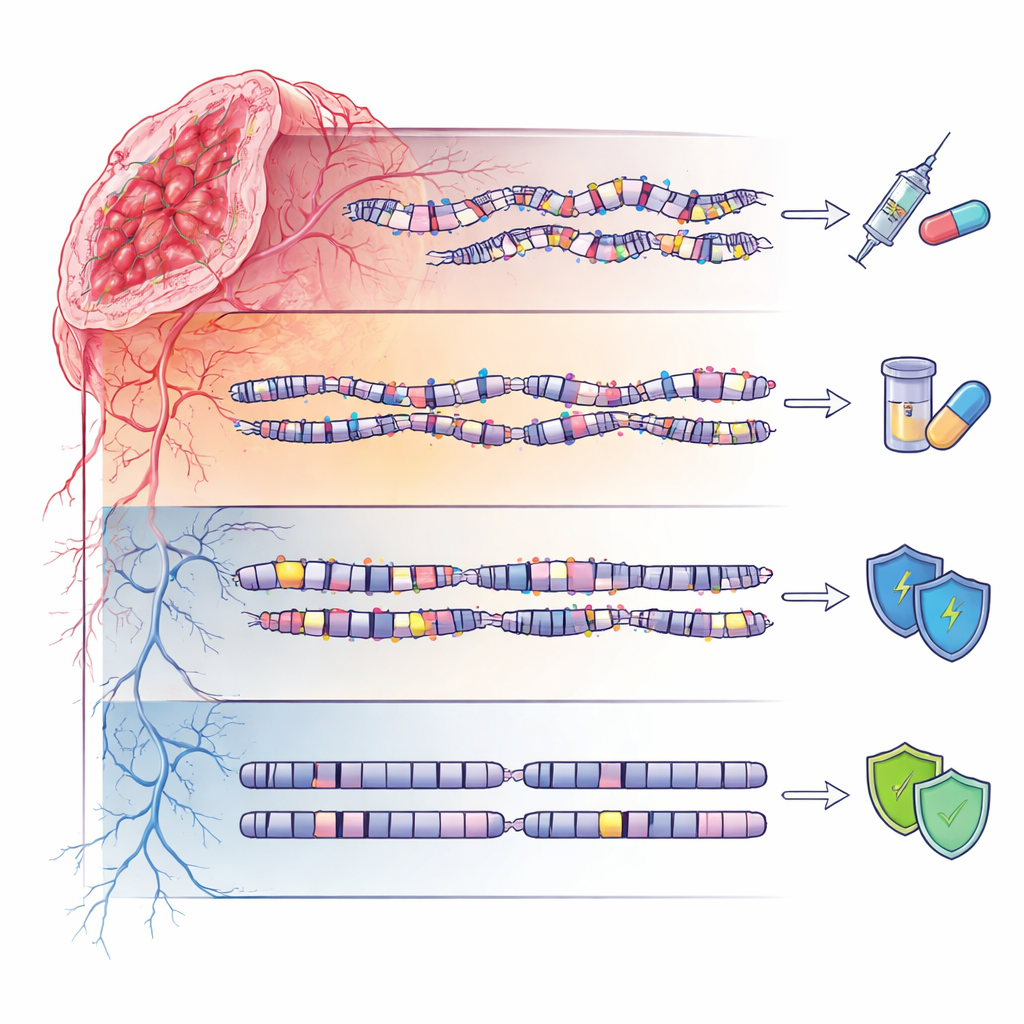
Quiet genomes and inherited risk
The fourth group, termed genome‑stable (GS), showed relatively calm DNA landscapes with few mutations or large‑scale disruptions. Yet these patients were not free of inherited influences: they were enriched for rare germline variants in several DNA repair–related genes, many of which are still classified as “variants of uncertain significance.” This suggests that in some families, subtle inherited changes may predispose to cancer without producing dramatic genomic chaos, underscoring the need for careful, ancestry‑aware interpretation of genetic test results, especially in non‑European populations.
What this means for future care
By moving beyond single‑gene tests and traditional expression‑based subtypes, this work proposes an “integrative genomic instability index” that combines repair defects, mutation burden, copy‑number changes, and structural signatures to place hereditary breast cancers into four mechanistic groups. In practical terms, it supports matching HRD tumors to PARP inhibitors, CN‑driven tumors to specific chemotherapies and stromal‑targeting strategies, and mutation‑heavy tumors to immunotherapy, while highlighting the need for ongoing reevaluation of germline variants in genome‑stable cases. For patients and clinicians, the study points toward a future in which inherited breast cancer care is guided not just by BRCA status, but by a richer picture of how each tumor’s DNA has gone wrong.
Citation: Kim, S., Lee, S., Kim, H. et al. Delineation of the heterogeneity underlying genomic instability in hereditary breast cancers reveals four disease subtypes. Exp Mol Med 58, 1254–1268 (2026). https://doi.org/10.1038/s12276-026-01693-4
Keywords: hereditary breast cancer, genomic instability, BRCA-negative, tumor subtypes, personalized therapy