Clear Sky Science · en
Joint modeling of cellular heterogeneity and condition effects with scPCA in single-cell RNA-seq
Why this matters for understanding cells
Modern biology can now read out which genes are active in thousands of individual cells at once. But when scientists compare cells across different treatments, ages, or genetic backgrounds, the sheer volume of data becomes overwhelming, and technical quirks can easily hide real biological changes. This paper introduces a new analytical tool, scPCA, that helps researchers disentangle what truly changes in cells under different conditions from the natural diversity that exists between cell types.
From noisy cell data to clear patterns
Single-cell RNA sequencing measures the activity of thousands of genes in each cell, producing extremely high-dimensional data. To make sense of this, researchers usually compress the data with methods like principal component analysis (PCA), which finds a small set of “axes” capturing the main patterns of variation. However, traditional approaches blend together two very different sources of variation: inherent differences between cell types and changes caused by an experiment, such as a drug treatment. The authors argue that this mixing can mislead downstream analyses like clustering cells into types or searching for treatment effects.
A new way to share structure across conditions
scPCA tackles this problem by explicitly telling the factorization model which condition each cell came from, and then learning a separate—yet linked—set of gene-expression patterns for every condition. Instead of forcing all samples to share exactly the same underlying structure, scPCA allows each condition to have its own version of each component, gently shifted from a designated reference condition. This preserves a common coordinate system for comparing cells across conditions, while still capturing systematic expression shifts driven by treatments, aging, or genetic changes.
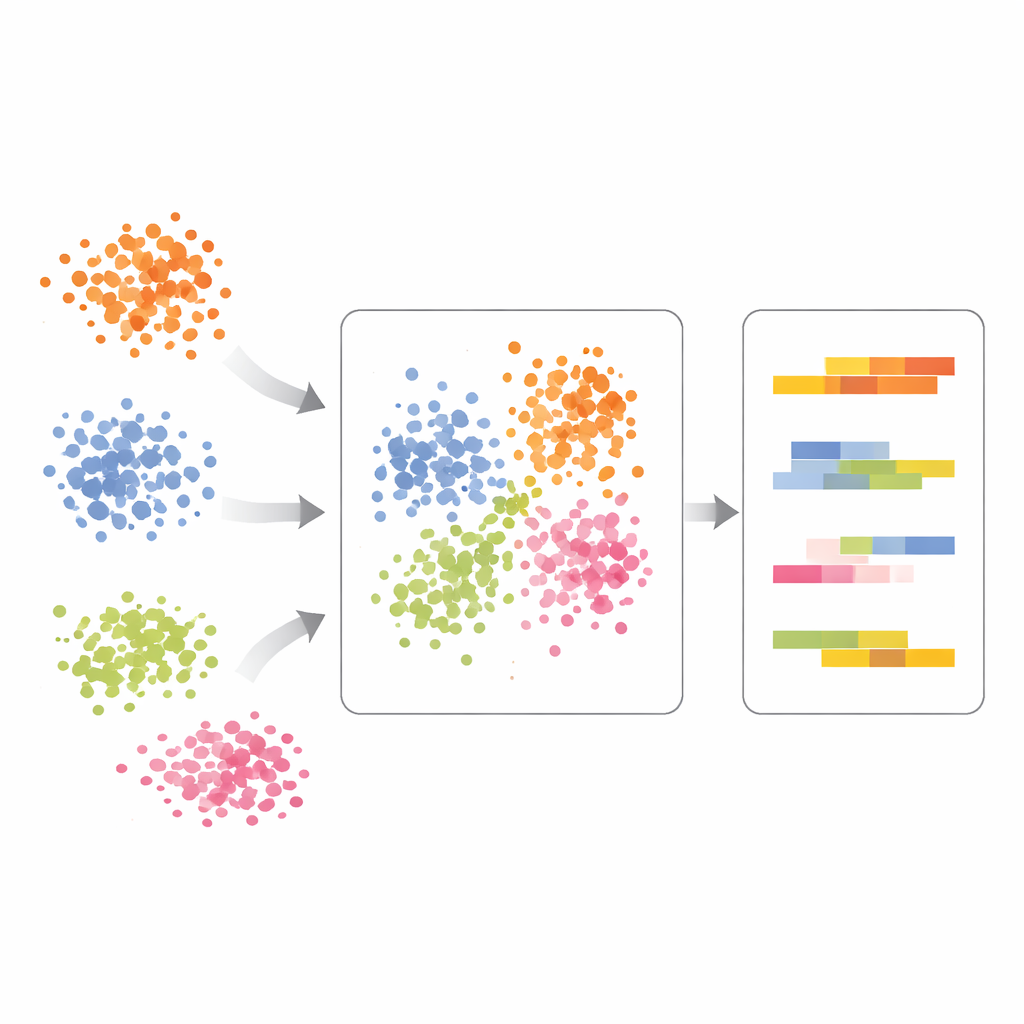
Seeing true treatment effects in immune cells
The authors demonstrate scPCA on immune cells from lupus patients, some left untreated and others stimulated with interferon-beta, a strong immune signal. Standard analysis caused cells to cluster by both cell type and treatment, making the results hard to interpret. With scPCA, cells of the same type from different conditions aligned much better, revealing that the main axis of variation still reflected immune cell lineages rather than treatment alone. Only after accounting for cell type did scPCA highlight treatment-driven shifts in specific genes in myeloid cells, including those tied to interferon signaling and altered protein handling inside the cell. This showed that the method can cleanly separate who the cells are from how they respond.
Untangling technical artifacts and aging effects
Experiments often suffer from batch effects: subtle differences caused by sample processing rather than biology. Using a mixture of two human cell lines measured in separate batches, the authors show that standard PCA preserves these technical differences, while scPCA can largely remove them by treating batch as a conditioning variable. The method then reveals which genes were responsible for the apparent batch separation, including markers specific to each cell line. In a more complex example, scPCA is applied to lung cells from young and old mice. It finds components that align with major cell types—such as pneumocytes, ciliated cells, macrophages, and T cells—and then pinpoints age-related gene changes within each, including stress and immune-response genes that fit with the idea of “inflammaging.”
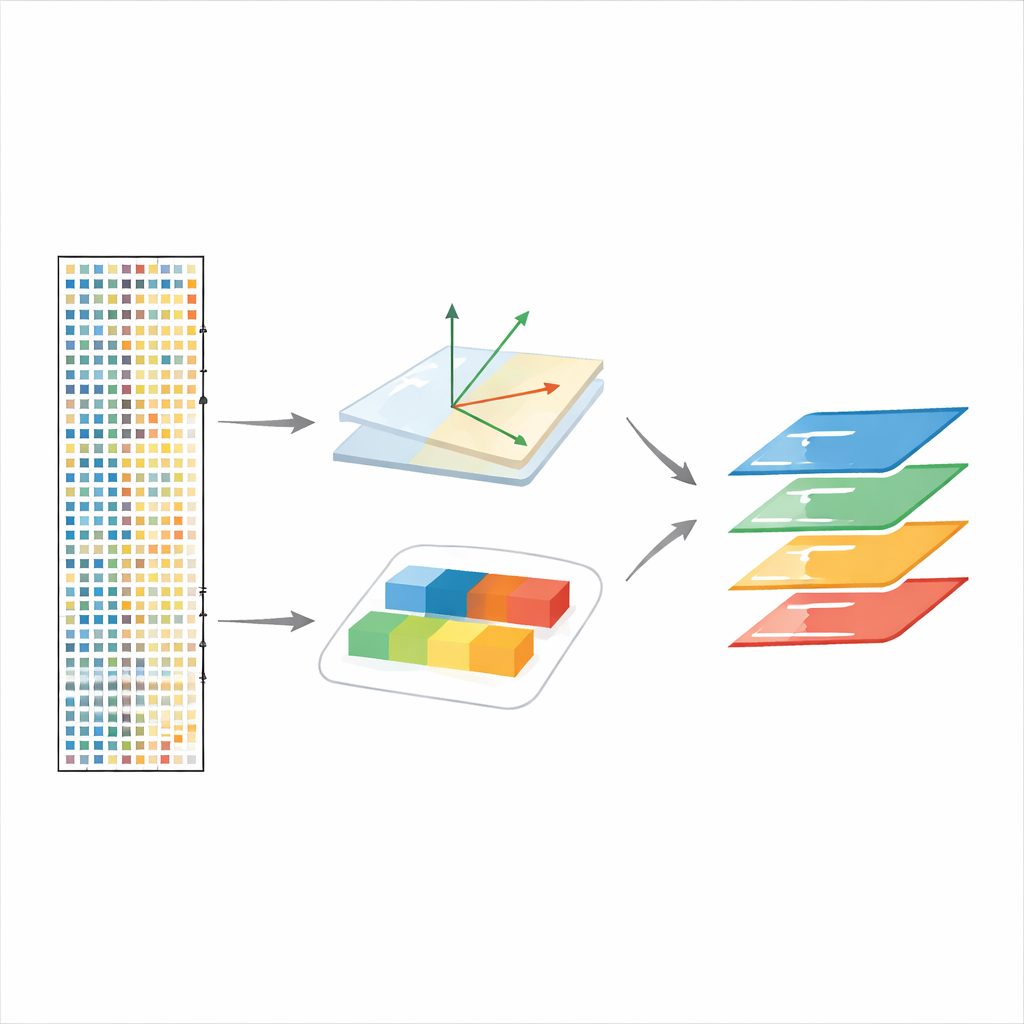
Following cell responses over time and across perturbations
scPCA also handles experiments with more than two conditions, such as neurons in mouse visual cortex exposed to light after a period in darkness. By treating time points as different levels of a condition, the method recovers early and late waves of gene activity across several brain cell types, separating rapid “early-response” factors from slower “late-response” programs. In a zebrafish experiment where a key developmental gene, chordin, is knocked out, scPCA successfully integrates embryos despite big shifts in cell-type composition and reveals transcriptional changes consistent with altered body patterning, including genes that were not emphasized in the original analysis.
What this means for future single-cell studies
In plain terms, scPCA gives researchers a clearer lens for looking at single-cell data collected under different conditions. It produces integrated maps where similar cells line up across treatments, and it highlights which genes within each shared pattern are turned up or down in response to a stimulus, aging, or genetic change. While the method assumes that underlying structure is largely shared and is best used for exploratory work that still requires follow-up validation, it offers a more transparent and interpretable alternative to many black-box models. This should help scientists draw more reliable conclusions from increasingly complex single-cell experiments.
Citation: Vöhringer, H. Joint modeling of cellular heterogeneity and condition effects with scPCA in single-cell RNA-seq. Commun Biol 9, 492 (2026). https://doi.org/10.1038/s42003-026-09651-6
Keywords: single-cell RNA sequencing, dimensionality reduction, cellular heterogeneity, gene expression changes, batch effects