Clear Sky Science · en
Immunoinformatic-based design of a multi-epitope subunit vaccine against Ruminococcus torques using subtractive proteomics and molecular dynamics simulations
Why this gut microbe matters
Hidden deep in the human intestine lives a bacterium called Ruminococcus torques that helps break down mucus lining the gut. When it grows out of balance, this microbe is linked to intestinal disorders such as inflammatory bowel disease and irritable bowel syndrome, and even infections outside the gut in vulnerable patients. Yet there is currently no vaccine that targets it. This study uses computer-based tools to design a candidate vaccine intended to nudge the immune system against R. torques while preserving the broader community of helpful gut bacteria. 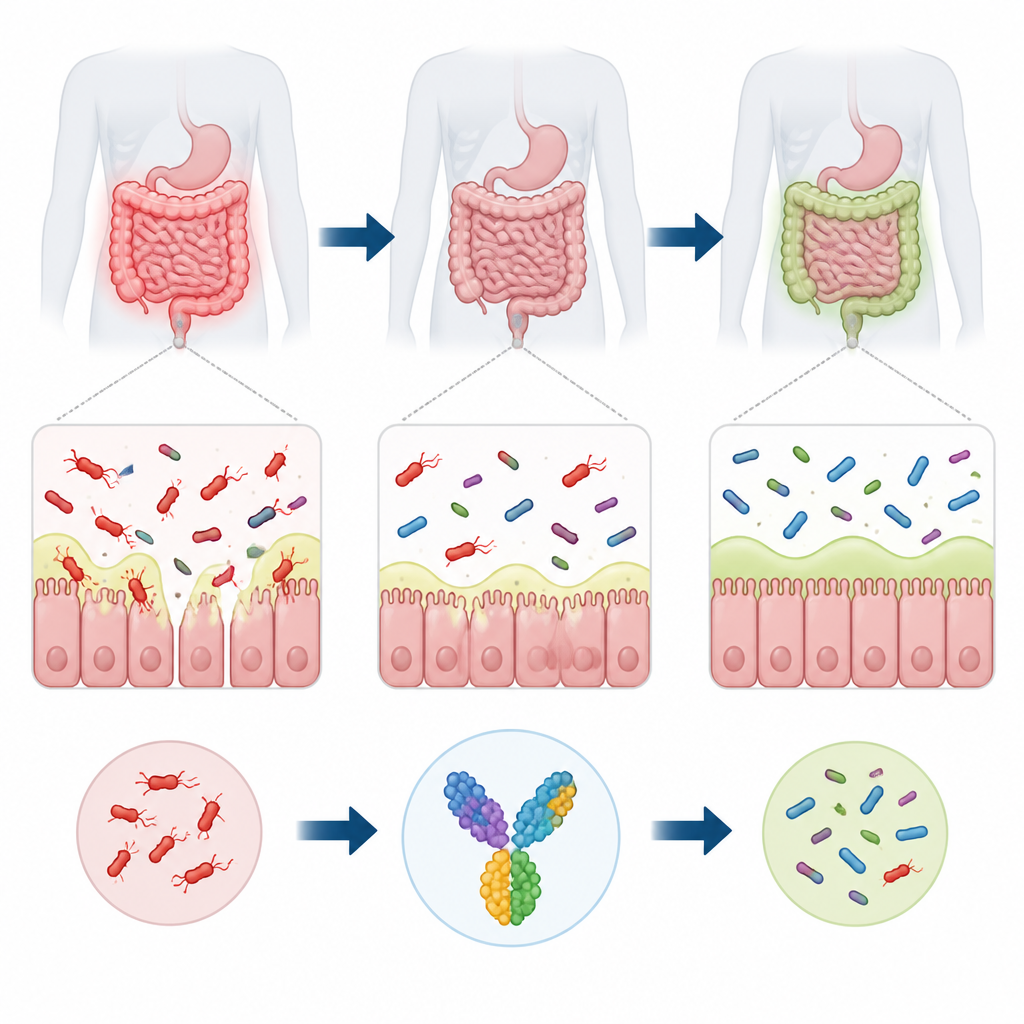
A troublemaker in the gut lining
The mucus layer that coats the intestine acts as a soft but vital shield, keeping most microbes at a safe distance from our cells. R. torques specializes in chewing through this mucus using powerful enzymes that dismantle the sugary side chains of mucin molecules. In a balanced gut, that activity can support a healthy ecosystem. But when R. torques becomes too abundant, it can thin the mucus barrier, increase leakiness of the intestinal wall, and expose immune cells to a constant stream of microbial fragments. This persistent irritation is associated with chronic inflammation in conditions like Crohn’s disease and ulcerative colitis, and may also contribute to obesity and metabolic problems.
Using computers to scout vaccine targets
Because R. torques is difficult to study in the lab, the researchers turned to its complete protein catalog, or proteome, which is publicly available in online databases. They used a stepwise “subtractive” strategy to narrow down nearly 2800 proteins to just a few promising vaccine targets. First, they identified proteins essential for the bacterium’s survival. Next, they removed any proteins that closely resemble human proteins or those from beneficial bacteria, to reduce the risk of unwanted cross-reactions. The remaining candidates were scored for how likely they are to trigger an immune response without being toxic or allergenic. This process spotlighted three key proteins involved in DNA protection and cell wall building as attractive targets.
Building a multi-piece vaccine
Rather than using whole proteins, the team focused on short stretches within them, called epitopes, that immune cells recognize. They predicted epitopes for killer T cells, helper T cells, and B cells, then filtered them using several criteria: strong predicted immune stimulation, absence of toxicity or allergy signals, and no close similarity to human proteins. Selected epitopes were strung together like colored beads into a single “multi-epitope” vaccine chain, joined with short linkers so each piece can be properly displayed to the immune system. An immune-boosting component derived from a cholera toxin subunit was attached to the front of the construct to enhance its impact. Computer models suggested that the resulting protein is stable, soluble, and strongly antigenic. 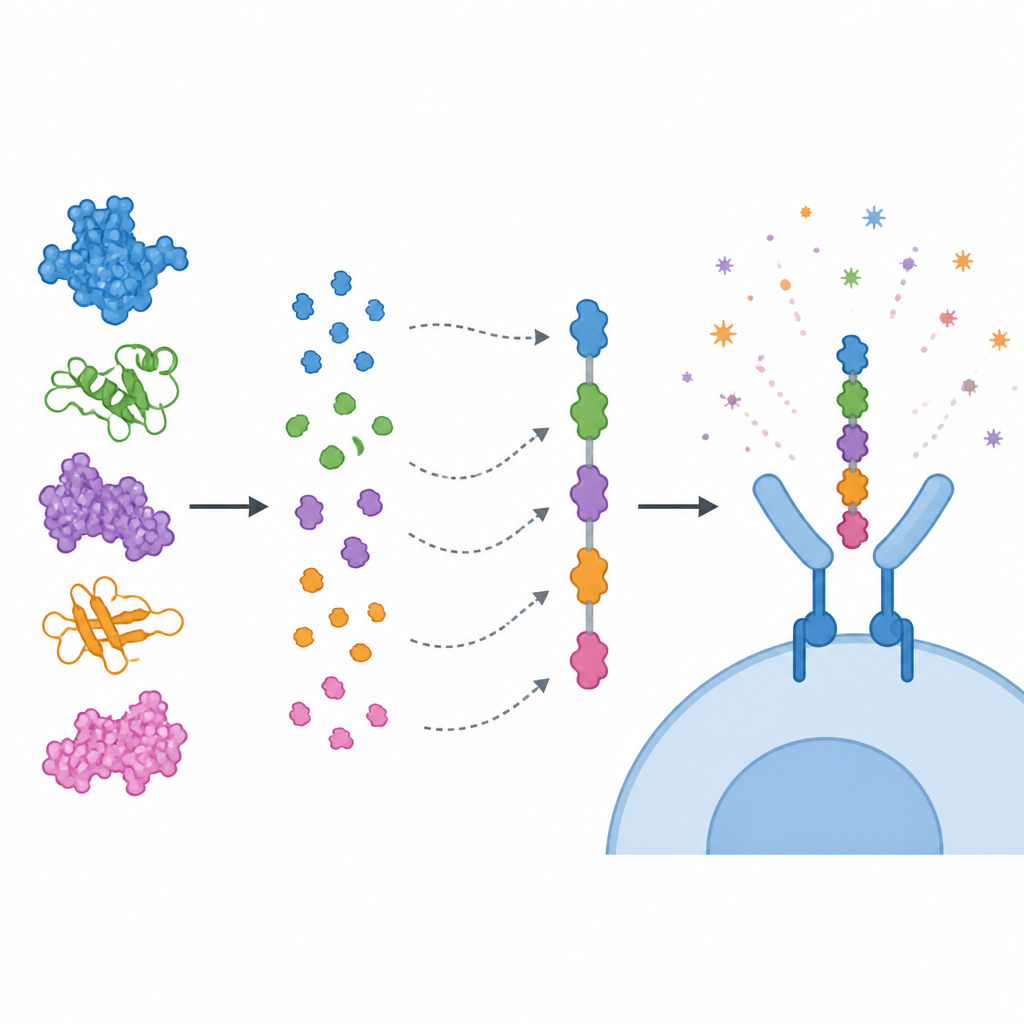
Testing fit and stability on screen
To see whether this designed molecule could talk to the human immune system, the authors simulated how it might bind to Toll-like receptor 4, a sentinel protein on immune cells that detects microbial threats. Docking studies indicated a snug interaction with many stabilizing contacts, and detailed motion simulations over 100 billionths of a second suggested the complex remains compact and stable in a watery environment. Additional simulations of the immune response predicted that the vaccine could stimulate both antibody production and long-lasting T cell responses. The gene sequence for the vaccine was also digitally adjusted to be easily produced in standard laboratory bacteria, a first step toward real-world testing.
What this could mean for future treatments
This work does not deliver a ready-to-use shot, but it lays out a carefully reasoned blueprint. Using only data and algorithms, the researchers designed a multi-part protein that appears safe, stable, and capable of awakening the immune system to R. torques. If future laboratory and animal studies confirm these predictions, such a vaccine could become part of a toolbox to prevent or lessen gut disorders linked to this mucus-eating microbe, ideally without disturbing the overall balance of the intestinal microbiome.
Citation: Kousar, S., Manzoor, I., Muhammad, S. et al. Immunoinformatic-based design of a multi-epitope subunit vaccine against Ruminococcus torques using subtractive proteomics and molecular dynamics simulations. Sci Rep 16, 15072 (2026). https://doi.org/10.1038/s41598-026-45572-4
Keywords: Ruminococcus torques, gut microbiome, multi-epitope vaccine, reverse vaccinology, immunoinformatics