Clear Sky Science · en
Identification and verification of the key genes, CCR1 and EGR2, in diabetes-associated lipophagy
Why fat and sugar can overwhelm our cells
Type 2 diabetes is usually described as a problem of sugar, but fat plays an equally important and tightly linked role. When our bodies are flooded with excess calories, fat droplets build up inside organs such as the liver and muscle. Healthy cells can "clean house" by breaking down these stored fats, but in diabetes this clean‑up system falters. The study summarized here asks a simple but powerful question: which genes sit at the crossroads between fat overload, faulty cellular clean‑up, and rising blood sugar—and could they become new warning signs or treatment targets?
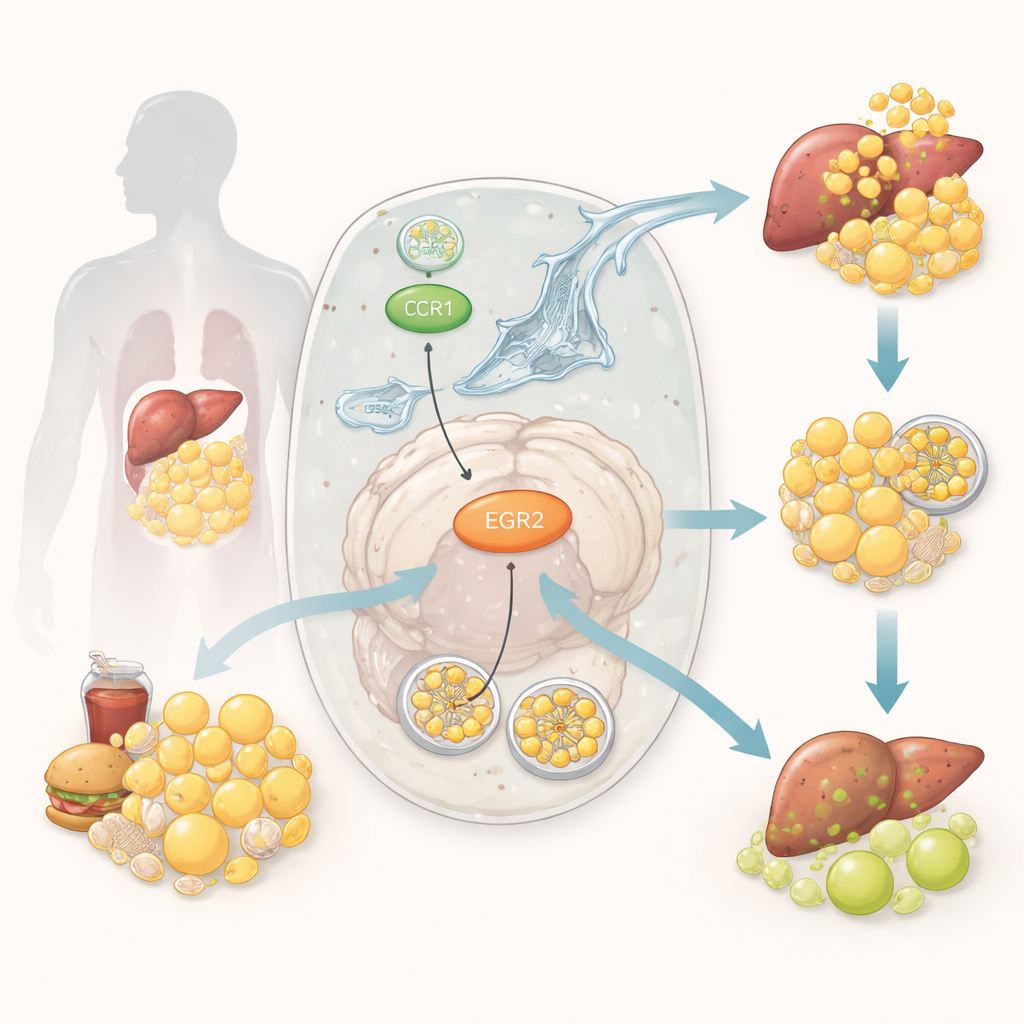
A cellular clean‑up crew for fat
Inside our cells, a specialized recycling pathway, called lipophagy, helps package and digest tiny fat droplets. Think of it as a cellular garbage truck that collects excess fat and delivers it to a disposal center so the contents can be safely reused. When lipophagy runs smoothly, fat stores stay in balance and cells respond properly to insulin. When it is disrupted, fat piles up in tissues such as the liver and muscle, fueling inflammation and making it harder for insulin to do its job. Although scientists know that both faulty fat handling and faulty cellular recycling are involved in diabetes, the precise molecular switches that connect these processes have been unclear.
Hunting for key switches in human data
The researchers began by mining large public databases of gene activity from the blood of people with and without diabetes. Using statistical tools and network analysis, they first narrowed thousands of genes down to a small group whose activity changed consistently in diabetes and that were already linked to fat metabolism or cellular recycling. They then applied several machine‑learning methods—techniques that allow computers to find robust patterns in complex data—to pinpoint which genes best distinguished diabetic from non‑diabetic samples. Across these independent approaches, two genes repeatedly rose to the top as central hubs: CCR1 and EGR2. Both were more active in the diabetic samples, and their expression levels tended to rise and fall together.
Confirming patterns in blood and liver
To check that these patterns were not just quirks of the original datasets, the team tested fresh blood samples from people with and without type 2 diabetes. Using a standard lab assay, they found that both CCR1 and EGR2 proteins were indeed higher in the blood of individuals with diabetes. Next, they turned to a well‑established mouse model of the disease that develops obesity, fatty liver, and high blood sugar. In these animals, the liver accumulated large fat deposits, and markers of the cellular recycling pathway showed signs of being blocked. In the same livers, CCR1 and EGR2 levels were clearly elevated, again echoing the changes seen in human blood. Together, these findings suggest that the two genes are tightly connected to the combination of fat build‑up, impaired recycling, and poor sugar control.
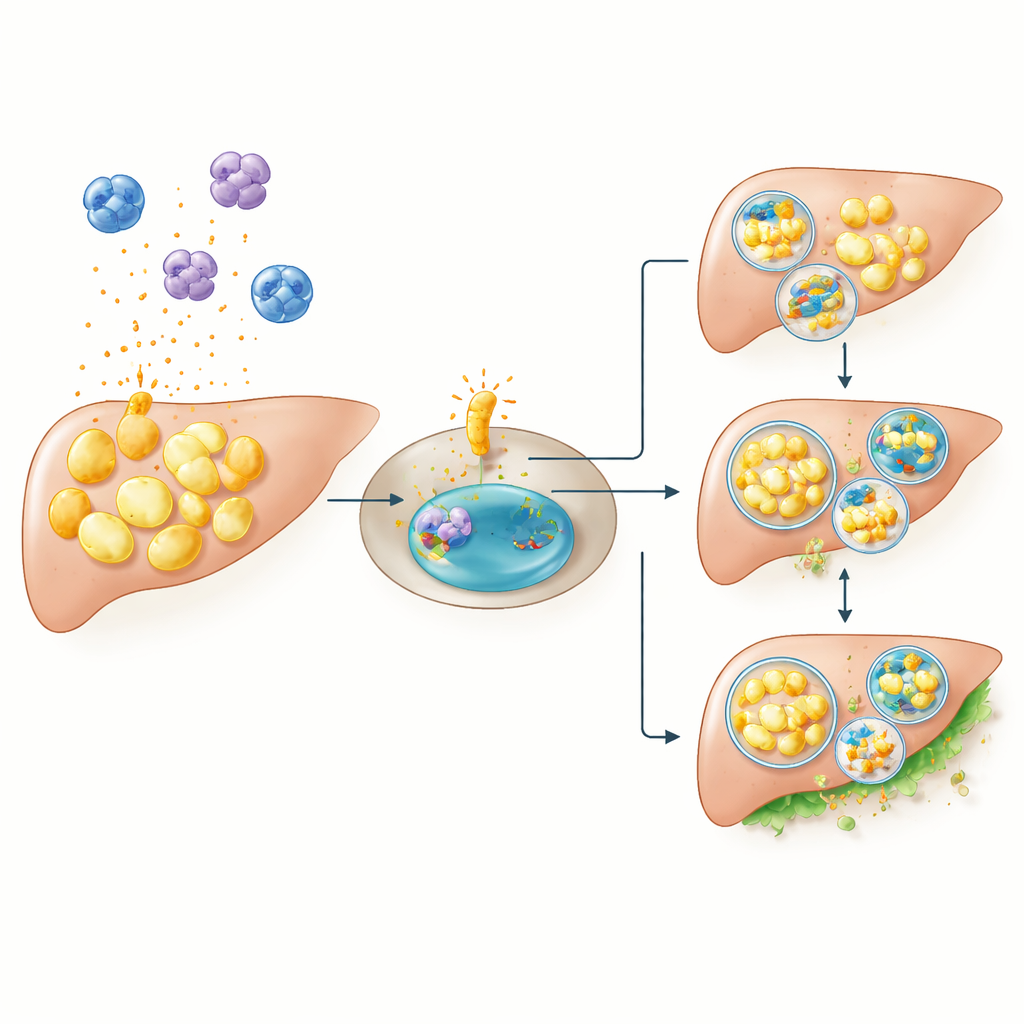
Probing cause and effect in mice
Correlation alone cannot prove that these genes help drive diabetes. To test cause and effect, the researchers generated mice lacking CCR1 and exposed them to a high‑fat diet, a stress that normally pushes blood sugar sharply upward. As expected, ordinary mice on the rich diet developed high fasting and after‑meal glucose levels. In striking contrast, mice without CCR1 were largely protected: under the same high‑fat conditions, their blood sugar stayed close to that of normally fed knock‑out animals. This suggests that CCR1 is not required to maintain everyday glucose balance, but becomes a critical player when the body is challenged by excess dietary fat, likely by amplifying inflammation and interfering with proper fat clearance inside cells.
What this means for people living with diabetes
By combining big‑data analysis, human blood measurements, and targeted mouse experiments, this study highlights CCR1 and EGR2 as potential molecular "dials" that tune how cells handle fat during diabetes. Higher levels of these genes track with fatty, inflamed liver tissue and with signs that the fat‑clearing lipophagy system is clogged. Importantly, removing CCR1 in mice blunts the rise in blood sugar normally triggered by a high‑fat diet, pointing to a possible new strategy for treatment. While much work remains to clarify exactly how CCR1 and EGR2 control fat recycling in different tissues, they now stand out as promising biomarkers to flag diabetes earlier and as starting points for drugs aimed at restoring the cell’s ability to clean up excess fat.
Citation: Liu, J., Zhang, X., Wang, Y. et al. Identification and verification of the key genes, CCR1 and EGR2, in diabetes-associated lipophagy. Sci Rep 16, 14274 (2026). https://doi.org/10.1038/s41598-026-43737-9
Keywords: type 2 diabetes, lipophagy, fatty liver, inflammation, biomarkers