Clear Sky Science · en
A parallel UNet integrating KAN and mamba for medical image segmentation
Why this matters for patients and doctors
Behind every cancer screening scan or ultrasound image, a crucial step is teaching computers to outline where a lesion or organ actually is. This process, called segmentation, helps doctors measure tumors, track disease, and plan treatment. The paper introduces KMP-UNet, a new kind of compact image-analysis network that promises to trace these boundaries more accurately and efficiently, potentially making automated medical image reading more reliable and easier to deploy in real clinics.
Seeing the whole picture in a medical scan
Modern medical images are large and complex: important clues can be spread across a wide area, while the lesion itself may have fuzzy or irregular edges. Traditional convolutional neural networks, including the popular UNet family, are good at spotting local patterns but have trouble connecting distant regions in a scan. Transformer-based models can see the whole image at once but tend to be heavy and slow, which is problematic for high‑resolution clinical data and limited hospital hardware. The authors build on a newer family of sequence models, called state space models, and a fresh style of neural network known as Kolmogorov–Arnold Networks, to capture both broad context and subtle details without ballooning the computation.
Two brainy specialists working side by side
KMP-UNet keeps the familiar U-shaped encoder–decoder layout of UNet but changes what happens inside. Instead of relying on a single type of processing, it runs two parallel branches through the network. One branch is based on a model called Mamba, which excels at handling long sequences and is adapted here to scan across the image. This branch efficiently gathers long-range context, helping the network understand how far‑apart regions relate to one another. The second branch uses Kolmogorov–Arnold Networks, which replace fixed activation functions with flexible, learnable curves. This gives the model extra power to represent tricky, nonlinear patterns, such as subtle intensity changes along the edge of a skin lesion or polyp.
Smarter fusion of features and clearer boundaries
Simply bolting together different model types is not enough; the real challenge is how to combine their strengths. To this end, the authors introduce a Parallel Convolution Mamba Block (PCMB), which splits features into groups and processes them through several light Mamba-based and convolutional paths before fusing them back together. This design balances global context and fine detail while keeping the computational cost low. They also revisit the skip connections that tie the encoder and decoder together. A new Enhanced Spatial-Channel Attention (ESCA) module selectively emphasizes important locations and feature channels, while preserving useful information through residual connections. This makes the network less likely to be confused by distractions such as hair, glare, or noisy textures around a lesion.
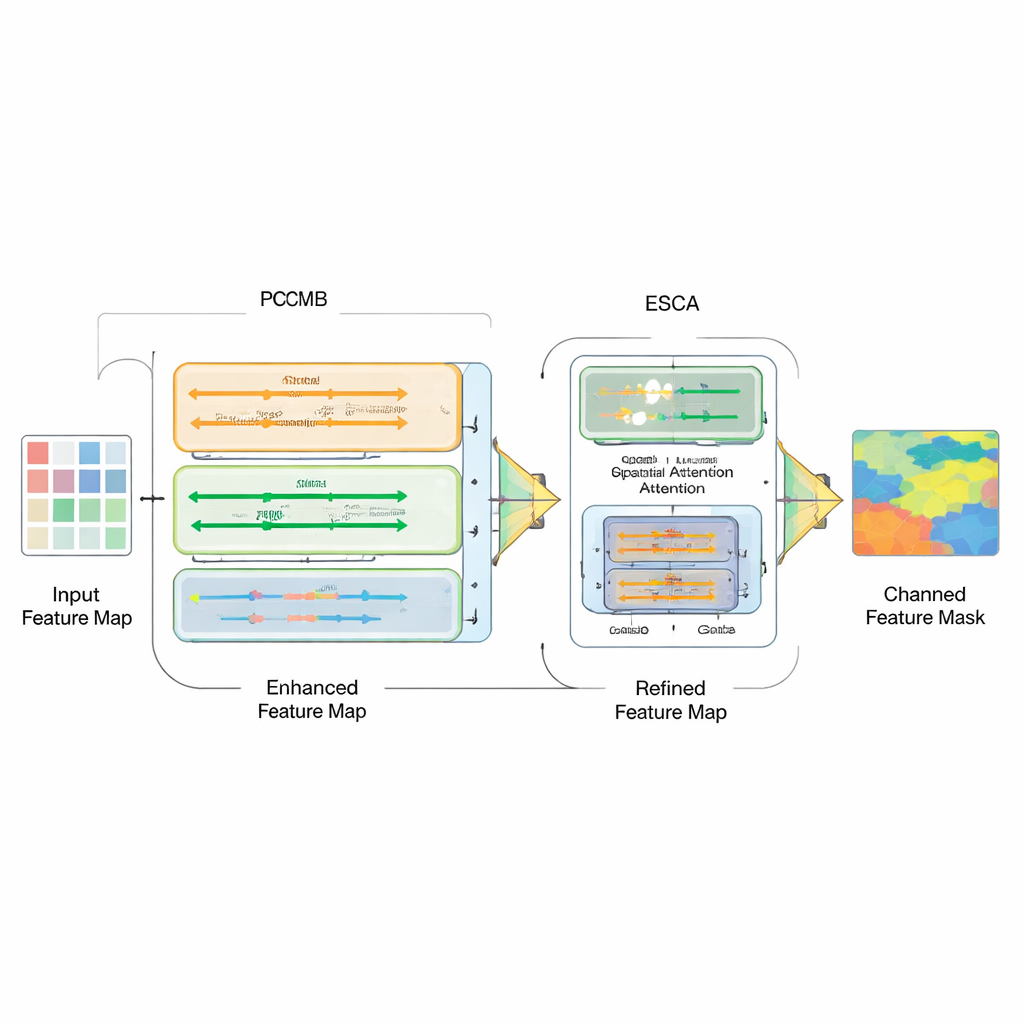
How well does the new approach work?
The team evaluates KMP-UNet on four public datasets covering skin lesions, colon polyps, and breast ultrasound images, and also tests transfer to another skin dataset. Across these benchmarks, the model consistently matches or outperforms a range of strong baselines, including classic UNet variants, Transformer-based designs, and other recent Mamba- and KAN-enhanced networks. For example, on a widely used skin lesion dataset (ISIC2018), KMP-UNet reaches a high Dice score and accuracy while using roughly one million parameters—far fewer than many competitors. Detailed experiments show that each component—Mamba-based context modeling, KAN-based nonlinear refinement, and ESCA-based skip refinement—contributes measurable gains, particularly in boundary quality and reduction of false positive regions.
What this means for future medical AI tools
For non-specialists, the key takeaway is that KMP-UNet offers a more compact yet capable way to teach computers to outline lesions and organs in images, by letting two complementary “experts” work in parallel and then carefully fusing their insights. While the method still struggles with the most challenging cases—very irregular shapes, extremely blurred borders, or heavy artifacts—it provides a strong foundation for accurate, efficient segmentation. With further work on handling noisy backgrounds and adapting across imaging centers, approaches like KMP-UNet could help bring dependable, fast image analysis closer to everyday clinical practice.
Citation: Liu, J., Wu, J., Xu, L. et al. A parallel UNet integrating KAN and mamba for medical image segmentation. Sci Rep 16, 13579 (2026). https://doi.org/10.1038/s41598-026-43127-1
Keywords: medical image segmentation, deep learning, UNet, state space models, Kolmogorov–Arnold networks