Clear Sky Science · en
Mathematical analysis of a stochastic delay model for respiratory syncytial virus dynamics
Why this study matters for everyday health
Respiratory syncytial virus (RSV) is best known for filling pediatric wards every winter, but the same virus also threatens older adults and people with weak immune systems. Health agencies need reliable forecasts of how RSV moves through a community and how control measures might change its course. This paper tackles a subtle problem behind those forecasts: most mathematical models ignore randomness and delays in how infection unfolds, which can make computer predictions misleading just when they are needed most. The authors build a more realistic model of RSV spread and, crucially, a safer way to simulate it on a computer so that predictions remain trustworthy even under uncertain conditions.

Breaking the outbreak into simple groups
The study begins by dividing the population into five groups: people who can catch RSV, those who have been exposed but are not yet contagious, those with ordinary infections, those who spread the virus much more than average, and those who have recovered. The model also builds in a waiting period between the moment someone is infected and the time they start passing the virus on. During this delay, people can still die from other causes, so the authors account for the chance that an infected person simply does not live long enough to become contagious. With these elements, they write a set of equations describing how people move between groups over time and show that the total population remains within realistic bounds and never becomes negative—a basic but essential check for any disease model.
Pinpointing the tipping point of an epidemic
Using their delay-based framework, the researchers calculate the “basic reproduction number,” a threshold that tells whether RSV will die out or continue circulating. This number blends together how quickly people mix, how long they stay contagious, how likely they are to become super-spreaders, and how long they live. If the reproduction number is below one, infections eventually vanish; if it is above one, the virus settles into a steady presence. The team proves mathematically that their model behaves exactly this way: one equilibrium corresponds to a community free of RSV, the other to an ongoing, but stable, level of infection. They also show that these outcomes remain stable even when the delay and other parameters vary within reasonable ranges.
Finding what matters most for control
Because many model inputs are uncertain, the authors test how sensitive the reproduction number is to each one. They find that factors increasing the rate of new infections—such as how often people come into contact or how quickly exposed individuals become infectious—push the system toward larger outbreaks. In contrast, factors that reduce the infectious period or increase natural death rates pull the reproduction number downward. This type of sensitivity ranking helps public health planners see which levers are most powerful: for example, strategies that effectively shorten how long people are infectious, or reduce opportunities for super-spreading, can have an outsized impact on RSV transmission.
Adding randomness and safer computer simulations
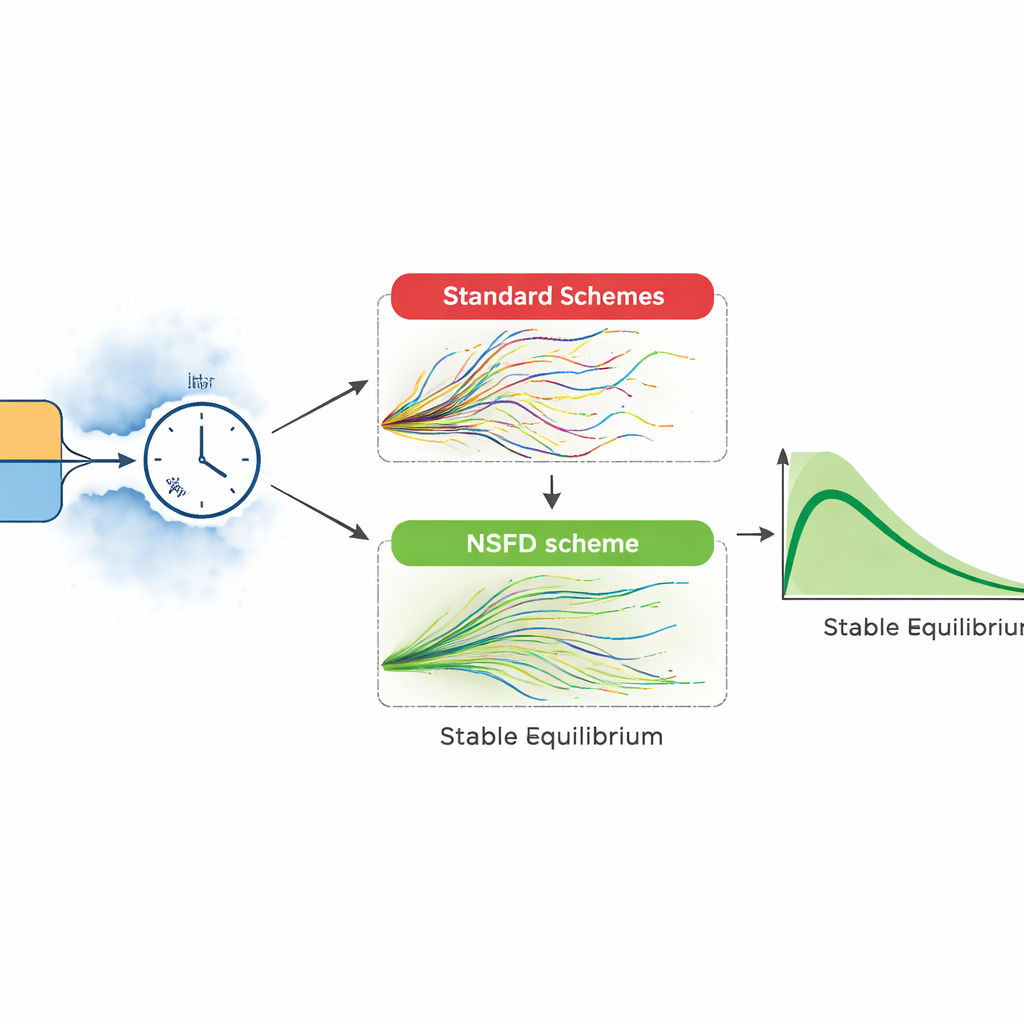
Real epidemics do not unfold as smooth curves. The authors therefore extend their model to include random environmental fluctuations and uncertainty in key processes, such as transmission and recovery. They construct two stochastic versions: one built from a detailed list of possible events (like an infection or recovery happening in a short time window) and another that perturbs each group with random noise. These richer models better reflect irregular outbreaks and unexpected surges. However, standard computer methods used to solve such models can behave badly: for large time steps, they may produce negative numbers of patients or explosive swings that have no biological meaning.
A robust new numerical tool
To overcome these pitfalls, the paper introduces a stochastic “nonstandard finite difference” scheme designed to respect the biology of the problem. In this approach, the drift, or average motion of the system, is handled carefully so that the simulated numbers of susceptible, infected, and recovered people always stay nonnegative and within sensible bounds, even when randomness is strong. The authors prove that this scheme remains stable for any size of time step, then test it against widely used methods such as Euler–Maruyama and stochastic Runge–Kutta. In head‑to‑head numerical experiments, the conventional methods work only for very tiny steps; when the step size grows, they generate unrealistic oscillations and even negative populations. The new scheme continues to track the expected course of infection smoothly, matching the theoretical equilibria of the model.
What this means for understanding RSV
For non-specialists, the key message is that the way we simulate an epidemic can be just as important as the equations themselves. This work delivers a detailed RSV model that captures both delays and randomness, and a companion computational method that keeps predictions physically meaningful under a wide range of conditions. Such a tool can help researchers and policymakers explore “what‑if” scenarios—like changing contact patterns or targeting super-spreaders—without worrying that numerical glitches are driving the results. In the long run, approaches like this could support more reliable planning for RSV vaccination campaigns and other interventions aimed at keeping seasonal surges under control.
Citation: Raza, A., Lampart, M., Shafique, U. et al. Mathematical analysis of a stochastic delay model for respiratory syncytial virus dynamics. Sci Rep 16, 10022 (2026). https://doi.org/10.1038/s41598-026-39783-y
Keywords: respiratory syncytial virus, epidemic modeling, stochastic dynamics, numerical simulation, disease transmission