Clear Sky Science · en
Distilling population specific expertise into a unified model for generalizable brain tumor segmentation
Why one model for everyone matters
Brain tumors do not look the same from one person to another. Children, adults, patients in different countries, and people with different tumor types can all have very different patterns on MRI scans. Yet doctors and hospitals would prefer a single, reliable computer tool that can outline tumors automatically for every patient, instead of juggling many separate models. This study introduces a new way to train such a unified system so that it works well across five very different kinds of brain tumors, potentially bringing more consistent care to patients worldwide.
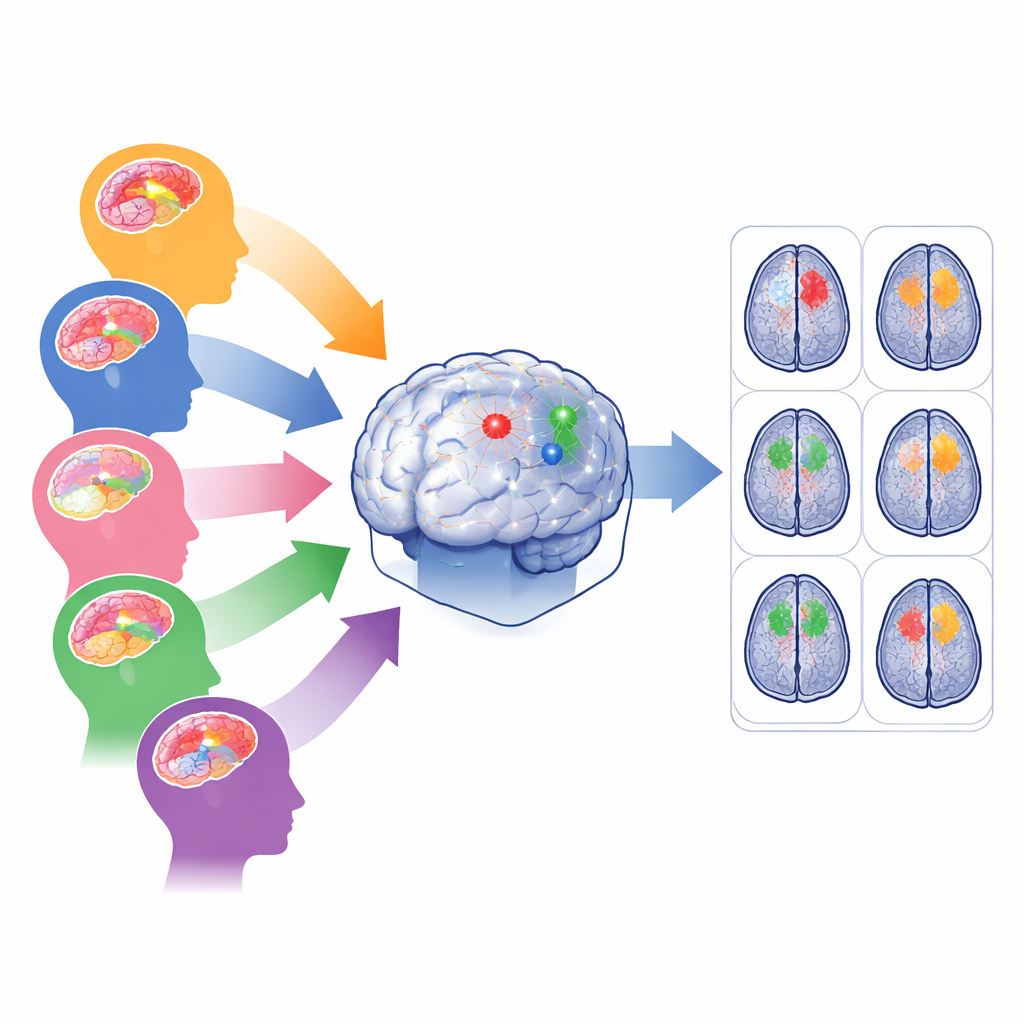
The challenge of many kinds of brain tumors
Modern artificial intelligence tools can trace brain tumors on MRI scans with impressive accuracy—but usually only when the data look similar to what they were trained on. In real hospitals, scans come from different machines and protocols, and tumors vary in size, shape, and location. Adult gliomas often lie deep within the brain, Sub-Saharan African gliomas are captured with lower-quality scanners, pediatric gliomas rearrange internal tumor regions, meningiomas sit on the brain’s surface, and metastases can appear as many scattered spots. Training one model on all of these cases at once tends to favor the most common group and neglect rare or noisy populations, risking poor performance exactly where help is most needed.
Why previous fixes fell short
Researchers have tried several workarounds. One option is to fine-tune a model for each new dataset or hospital, but this requires maintaining many versions and can cause the system to “forget” earlier knowledge. Another strategy is to use ensembles, where several specialized models vote on the final answer; while often accurate, this is slow and computationally expensive. Curriculum learning, which feeds data in a carefully chosen order from easier to harder cases, can help but is tricky to design and still may not capture all tumor varieties. Foundation models like “segment anything” promise general-purpose segmentation, but they usually need human prompts and are not fully automatic—limiting their usefulness in routine clinical workflows.
Blending many experts into one student
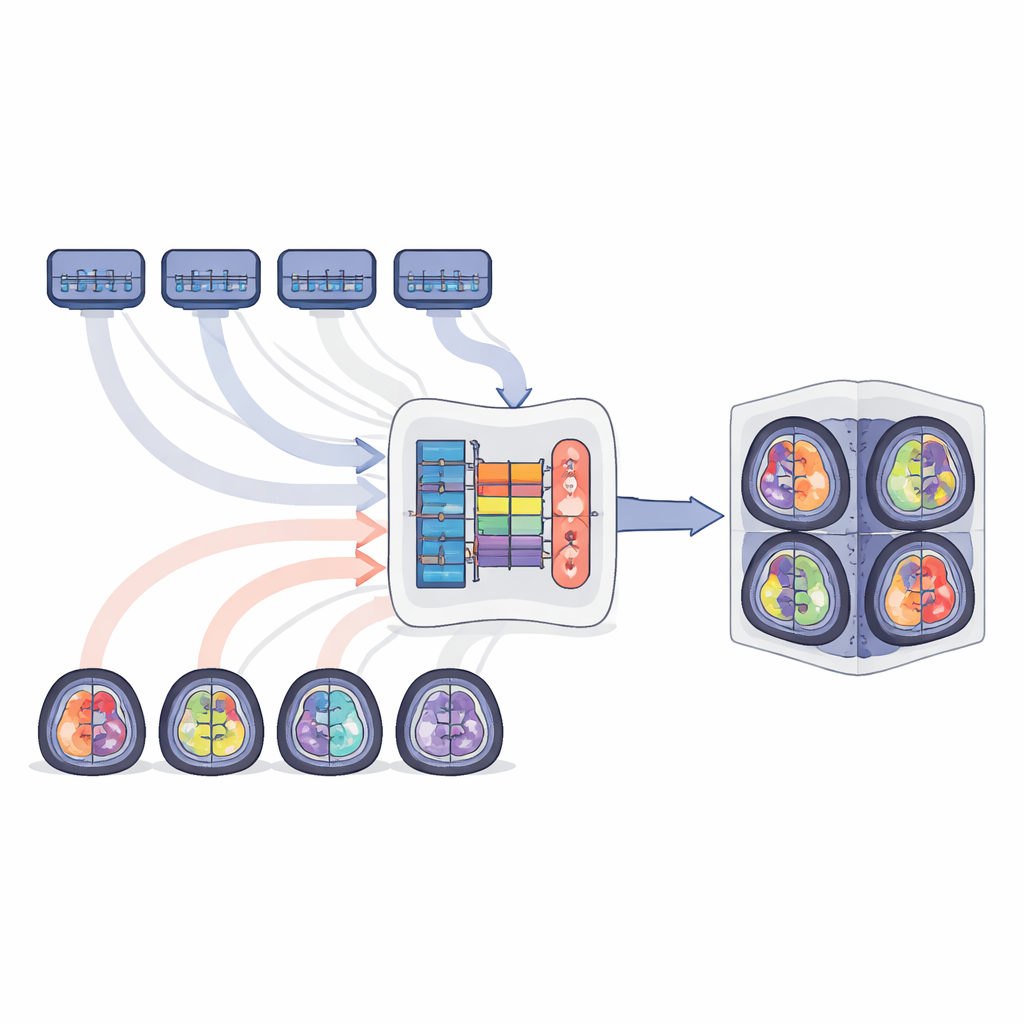
The authors propose MTSS-KDNet, a “multi-teacher, single-student” approach inspired by how a trainee doctor might learn from several specialists. First, five expert models are trained separately, each focusing on one tumor population: adult gliomas, Sub-Saharan African gliomas, pediatric gliomas, meningiomas, and metastases. These teacher networks become highly tuned to the features of their own group. Then a single student model is trained to imitate them—but in a smart, population-aware way. For each training scan, the matching teacher and the student process the image in parallel. The student is nudged to match the teacher’s internal patterns and final predictions while also being corrected using the true tumor outlines. A fairness-focused sampling scheme ensures that each training batch always includes one case from each population, so rare groups are seen as often as common ones.
How the unified model performs in practice
After training, only the student model is used for new patients, keeping deployment simple and fast. The researchers evaluated it on all five tumor populations using standard measures of how closely the predicted tumor volumes and boundaries match expert labels. Across whole tumor, tumor core, and actively growing regions, the unified model achieved strong scores that matched or exceeded not only its individual teachers but also powerful baselines like fine-tuned models, curriculum learning, and multi-model ensembles. Gains were especially striking for difficult settings such as pediatric tumors, low-quality Sub-Saharan African scans, and scattered brain metastases, where the student clearly outperformed population-specific experts. Visual inspections showed cleaner, more complete outlines, and internal feature maps revealed that the model learned to separate populations in its internal representation without needing labels at test time.
What this means for future patient care
To a non-specialist, the key idea is that the authors have found a way to pour the know-how of many narrow, expert systems into one general model, without losing what makes each expert good at its own job. Their MTSS-KDNet framework keeps the practical advantages of a single, automatic tool while preserving performance across diverse patients, scanners, and tumor types. Although it still demands substantial training resources and depends on good initial teacher models, this approach points toward “foundation” segmentation systems that can serve global populations more fairly. In the long term, such unified models could help ensure that patients with rare tumors, children, or those in under-resourced regions benefit from the same level of imaging precision as those in major medical centers.
Citation: Elzayat, A., Hanafy, N., Magdy, M. et al. Distilling population specific expertise into a unified model for generalizable brain tumor segmentation. Sci Rep 16, 12969 (2026). https://doi.org/10.1038/s41598-026-35627-x
Keywords: brain tumor segmentation, medical imaging AI, knowledge distillation, model generalization, MRI