Clear Sky Science · en
tRNA-deacylase-directed discovery of biosynthetic pathways
Hidden chemistry inside living cells
Many of today’s medicines, from painkillers to antibiotics and cancer drugs, began life as small molecules that microbes or plants make for their own purposes. Yet even after decades of searching, scientists suspect that most of nature’s chemical tricks are still undiscovered. This study reveals a new way to spot hard‑to‑find pathways in microbial DNA that build unusual amino acids and short peptides—molecules that could become tomorrow’s drugs or research tools.
Why special building blocks matter
Amino acids are best known as the building blocks of proteins, but cells also reshape them into an enormous variety of other molecules. Some are non‑standard amino acids that contain rare elements like fluorine or arsenic, or unusual linkages between nitrogen atoms. Others are stitched into complex antibiotics, neuromodulators such as psilocybin, or peptide hormones like insulin. Because these structures are so diverse, the genes that make them often do not resemble the classic enzyme families that current genome‑search tools look for. As a result, many of the underlying pathways remain invisible in the vast piles of microbial DNA now being sequenced.
Quality‑control enzymes as signposts
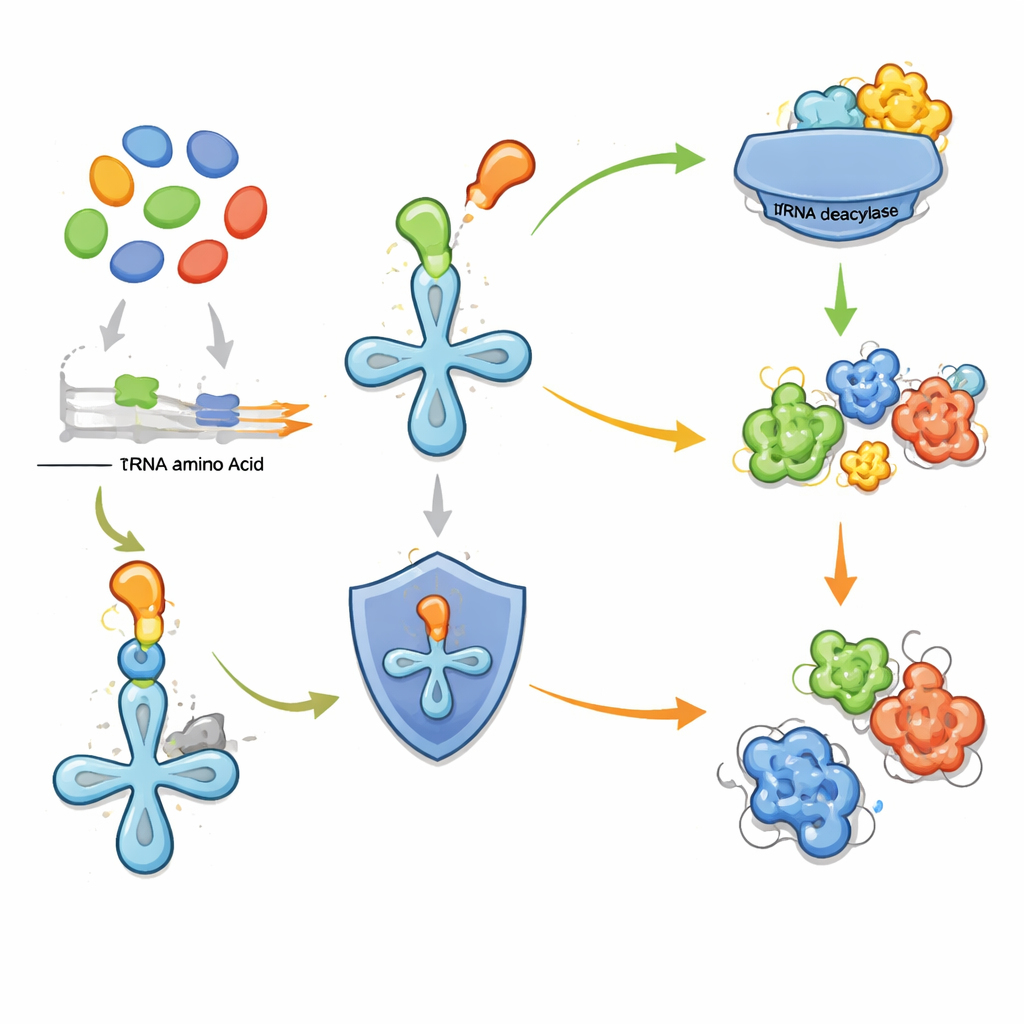
The key insight of the new work is that certain cellular “proofreading” enzymes can act as beacons for nearby pathways that make strange amino acids. When a cell produces a non‑standard amino acid, there is a risk it will be mistakenly inserted into proteins, damaging their function. To avoid this, some microbes carry standalone tRNA deacylases—enzymes that remove incorrectly attached amino acids from transfer RNA, the adaptors that feed building blocks into the protein‑making machinery. The authors had previously shown that one such deacylase protects bacteria from misusing a fluorinated threonine. Here they extend that idea: if a deacylase gene sits next to a cluster of other metabolic genes, it may be there to guard against a locally made, potentially harmful amino acid. That makes the deacylase a handy marker for an otherwise cryptic biosynthetic gene cluster.
Mining genomes with a new lens
Focusing on one deacylase family called AlaX, the team sifted through more than 23,000 related sequences. They built a visualization tool, tR3D, that maps how each deacylase gene is surrounded by its neighbors in the genome. Most AlaX enzymes appeared alongside housekeeping genes involved in routine amino‑acid and protein metabolism. But about 11 percent sat in more exotic company, near transporters, regulators, and unusual enzymes typical of secondary metabolism—the specialized chemistry microbes use for defense or signaling. This short list of promising clusters included only a modest fraction that belonged to well‑known natural‑product classes, suggesting that many others represent previously overlooked chemistry.
New molecules from marked clusters
To test the strategy, the researchers experimentally probed two very different gene clusters flagged by nearby deacylases. The first, a compact four‑gene unit found in over a hundred species, encodes a simple enzyme that adds a hydroxyl group to methionine, a common amino acid, creating a non‑standard variant. The partner deacylase selectively strips this modified methionine from transfer RNA, protecting the cell from misreading it as the normal building block. The second cluster, from a soil microbe, is more elaborate. It combines enzymes that generate a reactive nitrogen–nitrogen unit with others that join three amino‑acid‑derived pieces together. By reconstituting the pathway in laboratory bacteria and tracing labeled starting materials, the team uncovered a new hydrazide‑linked tripeptide they call fuscazine. Its backbone is built from arginine, alanine, and a ring‑shaped derivative of lysine, connected in an unusual way that had previously been seen only in more complex systems.
Opening doors to future discoveries
These case studies show that tRNA deacylases are more than simple quality‑control tools—they also highlight neighborhoods of DNA where evolution has pushed amino‑acid chemistry into new territory. By following these signposts, the authors identified thousands of candidate clusters, many of which fall outside standard categories used in current genome mining. Their tR3D platform can be applied to other enzyme families as well, helping researchers zoom in on genes likely to encode unusual bond‑forming reactions. For non‑experts, the takeaway is that living cells hide a vast library of chemical solutions, and smart ways of reading genomes—guided by built‑in safety valves like deacylases—are beginning to reveal it. Each newly mapped pathway not only expands basic understanding of how life builds molecules, but also offers fresh starting points for designing drugs and biocatalysts.
Citation: Millar, D.C., Zhou, Y., Marchand, J.A. et al. tRNA-deacylase-directed discovery of biosynthetic pathways. Nat. Chem. 18, 863–871 (2026). https://doi.org/10.1038/s41557-026-02126-5
Keywords: natural products, noncanonical amino acids, genome mining, tRNA quality control, biosynthetic gene clusters