Clear Sky Science · en
Variant U1 snRNAs contribute to cell cycle and differentiation control of human iPS cells
Why tiny RNA pieces matter for stem cells
Stem cells fascinate doctors and engineers because they can turn into almost any cell in the body. That promise depends on very strict internal control. This study looks at an overlooked group of tiny RNA molecules inside human induced pluripotent stem cells and shows that they quietly help decide how fast stem cells divide and how ready they are to become specialized cells.
Small helpers in the cell’s message factory
Inside every cell, genes are first copied into RNA messages, which are then trimmed and stitched together before being read as instructions for making proteins. A key part of this trimming machinery uses small RNAs called U1. For years, extra copies of U1-like RNAs were thought to be useless leftovers. The authors focus on two of these look-alikes, called variant U1s, that are especially active in early human tissues and stem cells. Because their sequences differ slightly from the main U1, they might recognize different cut sites in RNA and subtly change which protein versions a cell can make.
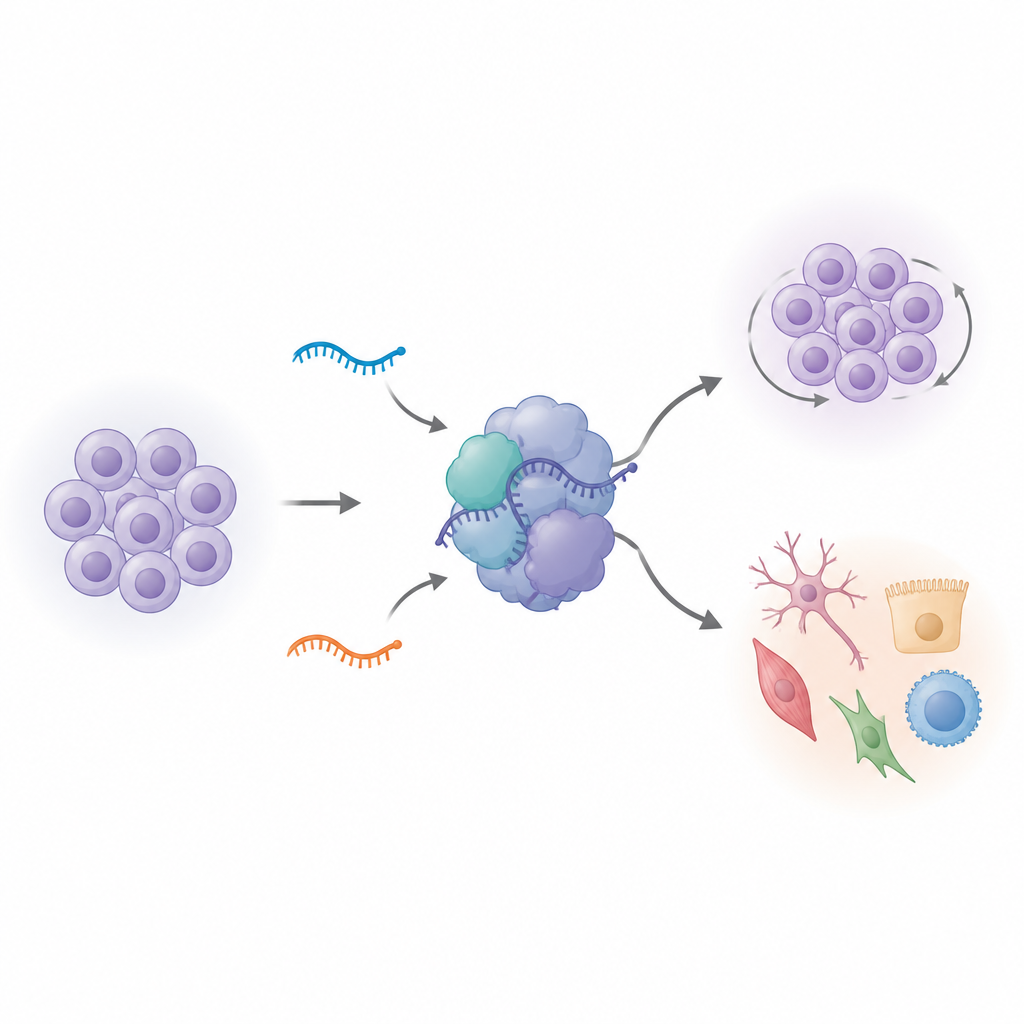
Switching off variant RNAs reshapes stem cell behavior
To see what these variant U1s actually do, the team used genome editing to remove either of the two variants in human induced pluripotent stem cells. At first glance the edited cells looked normal. But when the researchers measured which genes were turned on or off, they found sweeping changes, especially when one variant, vU1.8, was deleted. Hundreds of genes involved in cell division and stem cell identity shifted their activity. The cells’ internal clocks changed as well: more cells piled up in a later phase of the cell cycle, and overall growth rates increased when the cells were nudged to start differentiating. Genes that usually mark stemness and early brain development no longer followed their normal patterns, suggesting that these small RNAs help keep stem cells in a well-tuned state between self-renewal and specialization.
Editing messages by cutting and rejoining
The study then zoomed in on how variant U1s influence the detailed editing of RNA messages. Many human genes can be cut and rejoined in different ways, producing protein variants that differ in stability, location, or activity. By comparing normal and edited stem cells, the authors discovered thousands of changes in how RNAs were spliced. Some messages lost important protein-coding sections and instead produced shortened or noncoding versions. One variant favored changes at the start and end of messages, trimming the tail regions that can control how long an RNA survives and when it is used. Genes tied to cell division and neural development were especially affected, linking these tiny RNAs to processes that are central for both healthy growth and brain formation.
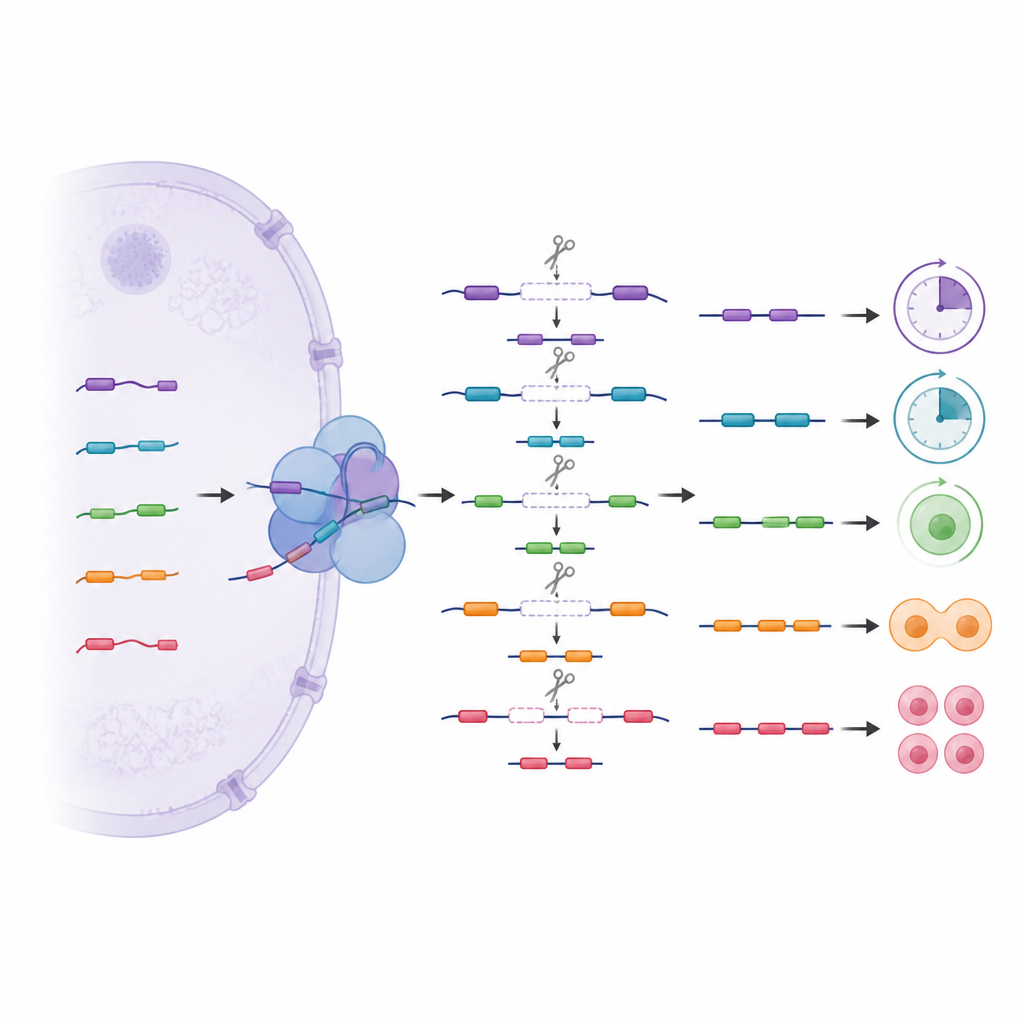
A hidden layer of stepwise cutting
Beyond ordinary splicing, the team explored a more complex pattern called recursive splicing, in which long gene segments are removed in several steps using hidden internal cut sites. These stepwise cuts were most common in cells from early development and in genes that manage DNA, RNA, and cell signaling. By tracking how often specific sites were used in normal and edited stem cells, the authors found that variant U1s help choose which hidden cut sites are recognized. Losing a variant caused some sites to be skipped and others to be used more often, which in turn altered nearby message pieces. Some of these hidden segments behave like cryptic mini-exons that are usually kept out of final messages but can slip in when control is lost, potentially leading to unstable or degraded RNAs.
What this means for stem cell science and disease
Overall, the work reveals that these variant U1 RNAs are not silent bystanders. They influence whether stem cells divide, which protein versions they can make, and how long their RNA messages last. By steering both ordinary and stepwise splicing, they add a fine-tuning layer to the gene control networks that sustain human pluripotent stem cells and guide their early steps toward becoming neurons and other cell types. Because genes containing these special splice sites also harbor genetic variants linked to human disorders, understanding how variant U1s work may eventually help explain why certain changes in noncoding DNA contribute to developmental and neurological disease.
Citation: Zhu, Y., Sofiadis, K., Mizi, A. et al. Variant U1 snRNAs contribute to cell cycle and differentiation control of human iPS cells. Nat Commun 17, 4334 (2026). https://doi.org/10.1038/s41467-026-73121-0
Keywords: stem cells, RNA splicing, cell cycle, pluripotency, neurodevelopment