Clear Sky Science · en
MechFind: a computational framework for de novo prediction of enzyme mechanisms
Why Understanding Enzymes Matters
Every living cell runs on tiny molecular machines called enzymes. These proteins speed up chemical reactions that power our bodies, grow crops, and manufacture medicines. While we know the overall “before and after” chemistry for tens of thousands of enzyme reactions, we rarely know the exact step‑by‑step moves they use to get from starting materials to final products. This paper introduces MechFind, a computer framework that can automatically propose detailed enzyme reaction steps using only basic chemical information, helping scientists design better enzymes for medicine, industry, and sustainable chemistry.
From Missing Steps to a Digital Detective
Most biochemical databases list what goes into an enzyme‑driven reaction and what comes out, but not the sequence of bond‑breaking and bond‑making events in between. Fewer than a thousand reactions in the literature are annotated with full mechanisms, leaving a large “mechanism gap.” Previous computational tools tried to fill this gap but often required detailed 3D structures of enzymes or advance knowledge of which amino acids in the protein do the chemistry, sharply limiting their reach. MechFind tackles the problem in a different way: it ignores the full 3D protein and instead focuses on how small chemical pieces are rearranged, letting it work on any reaction where the input and output molecules are known.
Breaking Chemistry into Simple Building Blocks
At the heart of MechFind is a “moiety‑based” view of chemistry. Rather than tracking whole molecules, the method breaks them down into tiny labeled fragments based on each atom’s immediate neighbors. It then represents any reaction as gains and losses of these fragments. Using a large, curated collection of known enzymatic steps, MechFind assembles chains of small fragment changes that collectively reproduce the overall reaction. It prefers the simplest explanations, searching for mechanisms with as few steps as possible and enforcing strict conservation of atoms and charge so that nothing appears or disappears illegally along the way.
Checking Accuracy and Discovering New Paths
To see whether its predictions make sense, the authors first tested MechFind on hundreds of reactions whose mechanisms were already documented in a trusted database. Using only the starting and ending molecules as input, MechFind recovered the accepted mechanism as its top choice in nearly two‑thirds of cases and placed it within the top ten in 85% of them. The team then challenged the system with six recently published enzyme mechanisms it had never seen before. MechFind still identified the correct step sequence among its leading candidates, often by reusing chemical patterns derived from unrelated species, such as building a human enzyme mechanism from steps originally learned from yeast or plant enzymes.
Scaling Up to Tens of Thousands of Reactions
After validating its accuracy, the authors unleashed MechFind on two major biochemical reaction collections, covering nearly 38,000 distinct reactions. For more than half of the reactions in each database, the tool produced at least one plausible multistep mechanism, often suggesting up to ten alternative paths. This effort generated over 18,000 new mechanistic hypotheses, a more than tenfold jump in the number of reactions with proposed detailed steps. At the same time, the large‑scale test highlighted where the method currently fails—either because the optimization becomes too complex within the allotted computing time, or because the necessary fragment types never appeared in the training data.
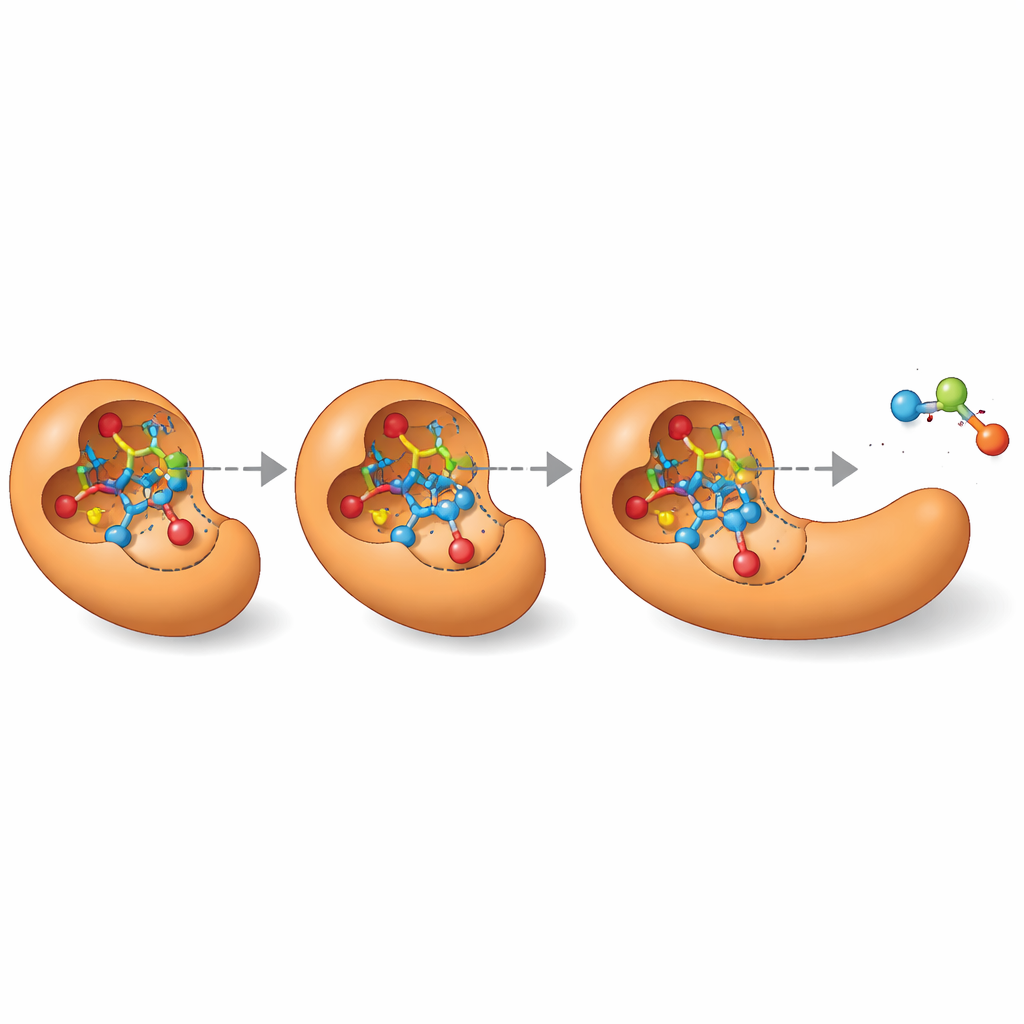
Opening Doors for Enzyme Design
Beyond listing a single “best guess,” MechFind can map out entire networks of different plausible routes that an enzyme might use for a given reaction. This landscape of alternatives is especially valuable for designing new enzymes from scratch. Modern protein design tools need a precise three‑dimensional picture of the fleeting transition states that reactions pass through. By laying out stepwise mechanisms, MechFind points directly to those key intermediates and transition states, turning vague design goals like “build an esterase” into concrete targets such as “stabilize this particular high‑energy structure.” Although its predictions still require expert review and follow‑up calculations or experiments, MechFind substantially expands our catalog of possible enzyme chemistries and charts a practical path toward more systematic, data‑driven enzyme engineering.
Citation: Hartley, A.D., Upadhyay, V., Boorla, V.S. et al. MechFind: a computational framework for de novo prediction of enzyme mechanisms. Nat Commun 17, 3903 (2026). https://doi.org/10.1038/s41467-026-71957-0
Keywords: enzyme mechanisms, computational biochemistry, enzyme design, metabolic reactions, reaction prediction