Clear Sky Science · en
Population differences of chromosome 22q11.2 duplication structure predispose differentially to microdeletion and inversion
Why this chromosome hotspot matters
Some people are born missing a small piece of chromosome 22, a change that can lead to heart defects, learning difficulties, facial differences, and a higher risk of schizophrenia. This condition, called 22q11.2 deletion syndrome, is one of the most common known genetic disorders. Yet it is puzzlingly less frequent in people of African ancestry. This study digs into the fine structure of the DNA in this chromosome region, showing how subtle differences between populations can change the chances of dangerous breaks, flips, and losses of genetic material.
A fragile stretch of repeated DNA
The authors focus on a 5-million-letter stretch of DNA on chromosome 22 called 22q11.2. This region is packed with large, nearly identical repeated segments of DNA, known as low-copy repeats. These repeats act like similar-looking paragraphs in a book: when cells make eggs or sperm, the copying machinery can accidentally line up the wrong repeats and swap pieces between them. That process, called non-allelic homologous recombination, can delete sections, duplicate them, or flip them around. Most 22q11.2 deletion syndrome cases arise when two particular repeat blocks, named A and D, mispair and remove a roughly 3-million-letter segment that contains dozens of genes important for development and brain function.
Reading the region end-to-end
Until recently, this part of chromosome 22 was too repetitive and complex to read cleanly with standard DNA sequencing. The researchers used new long-read technologies that can follow stretches of DNA tens of thousands of letters long, combined with improved assembly software, to rebuild 135 complete versions of this chromosome segment from people around the world, plus several ape species. They discovered 63 distinct structural "haplotypes"—different ways the repeat blocks in region A are arranged and sized—varying more than eleven-fold in length. Much of this diversity comes from a core repeated unit about 105,000 letters long, bracketed by shorter repeated pieces. This core unit has multiplied and shuffled specifically in humans over the last million years, making our version of 22q11.2 far more elaborate than that of chimpanzees, gorillas, or orangutans.
Population differences in a genetic minefield
When the team compared human groups, they found that individuals of African ancestry tended to have longer, more repeat-rich versions of block A than non-Africans. Counterintuitively, these longer structures often place key repeats in opposite orientations relative to their partners in block D. In that arrangement, mispairing tends to produce harmless flips, or inversions, of DNA segments rather than large deletions that remove genes. Using both within-chromosome and between-chromosome comparisons of the core 105,000-letter repeats, the authors built a ranking of haplotypes that are more likely to yield deletions versus those that are more likely to yield inversions. African haplotypes were significantly enriched for architectures predicted to protect against the classic deletion, while East Asian haplotypes were enriched for structures predicted to favor deletions.
Tracking real-world flips and losses
To test these predictions, the researchers searched for large inversions—multi-million-letter segments flipped end to end—in hundreds of genomes. They identified several distinct inversion types spanning the 22q11.2 region. Each was rare overall, but most were found in people of African or admixed American ancestry, matching the idea that their local DNA layout favors inversions over deletions. The team then zoomed in on four families in which a child had a new 22q11.2 deletion not present in either parent. By fully assembling the parents’ and children’s chromosome 22 segments, they could pinpoint where the break and faulty repair occurred. In every case, the deletion breakpoints landed within the 105,000-letter repeat unit, but the exact positions varied, falling in small stretches of almost perfectly matched sequence that are especially prone to mispairing.
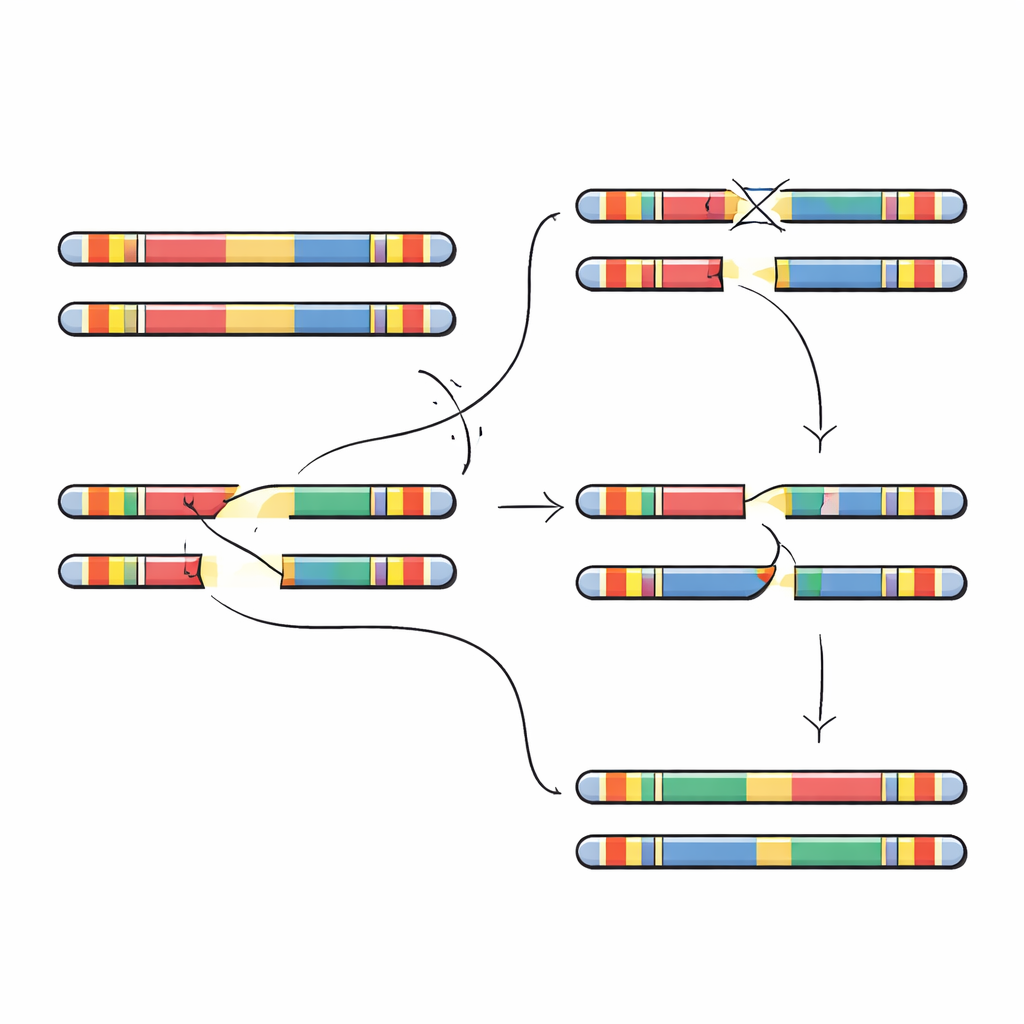
What this means for risk and diagnosis
Taken together, the findings show that not all versions of this chromosome region are equally fragile in the same way. Some architectures bias the system toward flips of DNA, which often leave the overall gene content intact, while others more readily produce gene-losing deletions. Because the protective architectures are more common in people of African ancestry, this helps explain why 22q11.2 deletion syndrome has been observed less often in these populations. At the same time, architectures predicted to be more deletion-prone are more frequent in East Asian genomes, suggesting that careful screening in these groups could be particularly important. More broadly, the work demonstrates how reading complex regions of the genome from end to end can reveal hidden structural differences that shape who is more or less likely to experience serious genetic disorders.
Citation: Porubsky, D., Yoo, D., Koundinya, N. et al. Population differences of chromosome 22q11.2 duplication structure predispose differentially to microdeletion and inversion. Nat Commun 17, 3701 (2026). https://doi.org/10.1038/s41467-026-71905-y
Keywords: 22q11.2 deletion syndrome, chromosome structural variation, segmental duplications, population genomics, genome instability