Clear Sky Science · en
A slight mismatch between a gene’s codon usage and the cellular tRNA supply is beneficial
When "Silent" DNA Choices Aren’t So Silent
Our genes contain many ways to write the same instruction, using different three-letter DNA “words” that all mean the same amino acid. For decades, these alternative spellings—called synonymous codons—were thought to be mostly trivial details. This study upends that view. It shows that how closely a gene’s codon “accent” matches the cell’s translation machinery can make the difference between sluggish growth and thriving, and that a slight mismatch can actually be best.
How Cells Read Genetic Instructions
To turn genes into proteins, cells use small adaptor molecules called tRNAs that recognize codons on messenger RNA and deliver the right amino acids. Each amino acid (with two exceptions) has several possible codons, but cells do not use them equally. Over evolution, organisms develop a characteristic pattern of codon usage, and they stock their tRNA pool accordingly. The traditional view assumes that the ideal situation is a perfect match: codons used in genes mirror the available tRNAs, giving fast, accurate translation. Any deviation from this perfect match was thought to persist only because of random genetic drift and mutation, not because it was helpful.
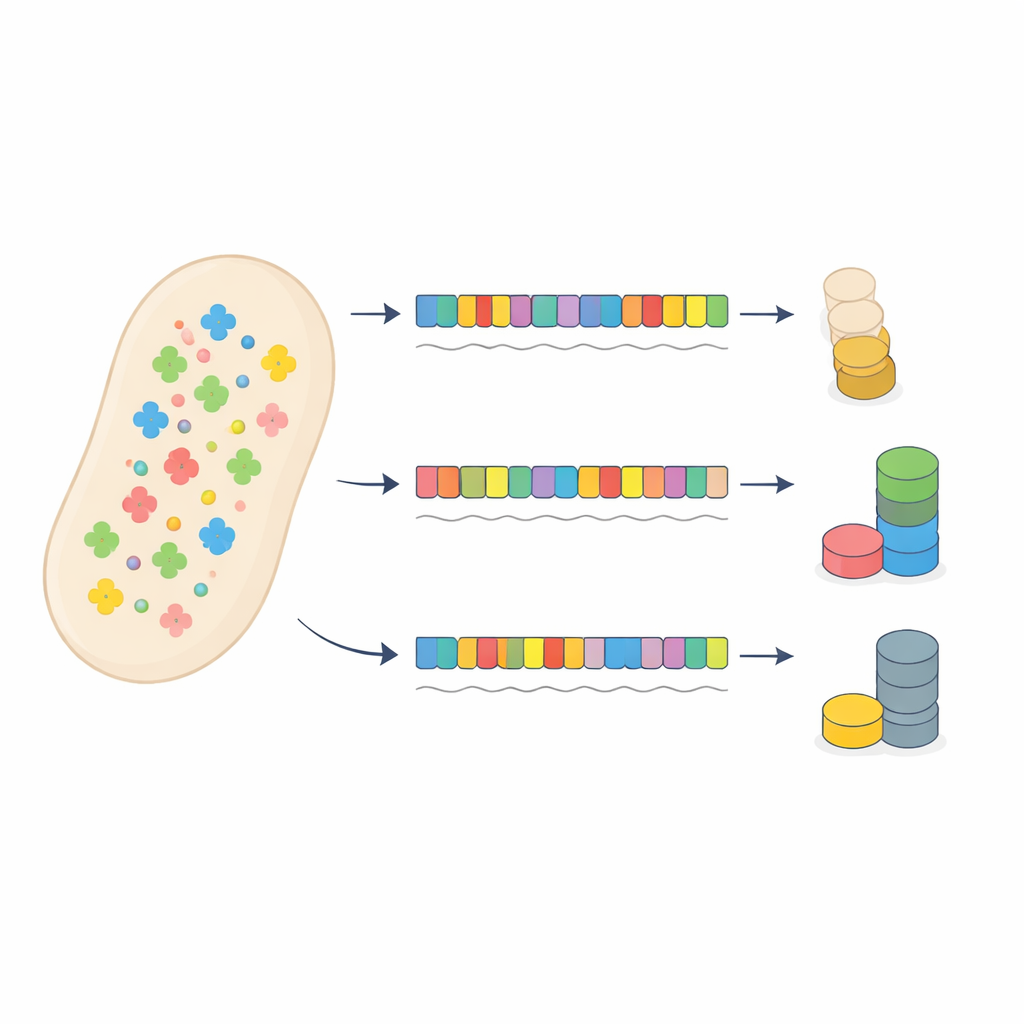
A Test Case Using Antibiotic Resistance
The authors challenged this conventional picture by carefully redesigning an antibiotic resistance gene in bacteria. They created 53 versions of a gentamicin resistance gene that all encoded the same protein but used different patterns of synonymous codons, ranging from a near-perfect match to the cell’s tRNA pool to a strong mismatch. Each version was placed on a plasmid and fused to a fluorescent marker so protein output could be measured. A second fluorescent protein on the same plasmid acted as a sensor of how hard the translation system was working overall, because heavy use of tRNAs by the resistance gene would slow translation of other proteins.
Finding the Sweet Spot for Growth
When these engineered bacteria grew in medium containing gentamicin, genes whose codons matched the tRNA pool extremely well produced the most resistance protein—but they also imposed the greatest burden on the cell’s translation machinery. Bacteria carrying such “over-optimized” genes grew more slowly than those with a modest mismatch. In other words, while better matching boosted the benefit of the resistance protein, it also raised the cost of using shared translation resources, and beyond a point the cost won. The fastest-growing cells were those whose gene versions showed a small, not minimal, mismatch with the tRNA supply, indicating that the true optimum balances payoff and cost rather than maximizing either alone.
When Gene Importance Changes, So Does the Optimum
The team then varied how crucial the resistance gene was by changing the antibiotic dose. At low gentamicin levels, the protein’s benefit to survival was modest, and the best-performing codon patterns showed a relatively large mismatch, which reduced wasteful strain on the translation system. As the antibiotic concentration rose and the gene became more important, the optimal codon pattern shifted toward a closer match with the tRNA pool, and at very high doses the nearly perfect match performed best. Similar experiments with a second resistance gene and with altering tRNA supply directly supported the same principle: the best codon usage depends on how much benefit a gene provides and how much translation cost it imposes.
Clues from Natural Genomes
To see if this trade-off is reflected in evolution, the authors examined thousands of genes from bacteria, yeast, fruit flies, and humans. For genes that matter more to survival—estimated from how much growth suffers when they are deleted—natural codon usage tends to match the tRNA pool more closely, especially among highly expressed genes. At the same time, among the most highly expressed genes, codon patterns with a perfectly tight match are actually less common; instead, these genes retain a modest mismatch, consistent with avoiding excessive translation cost. Mutation-accumulation experiments and precise fitness measurements of individual synonymous mutations in yeast further showed that both increasing and decreasing the mismatch are often harmful, implying that evolution maintains codon usage near an intermediate optimum rather than pushing endlessly toward a perfect match.
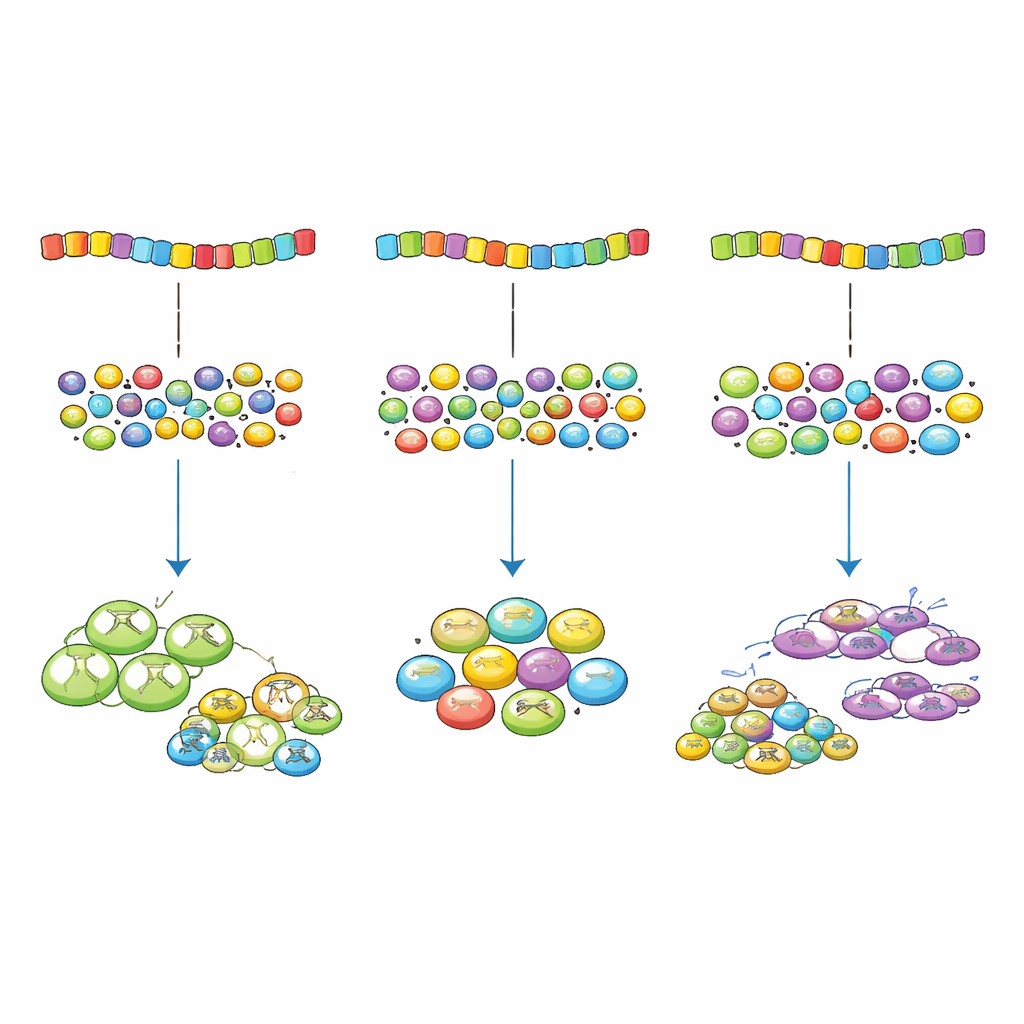
Why a Little Imperfection Helps
For a non-specialist, the key message is that there is such a thing as “too optimized” when it comes to gene spelling. A perfect match between codons and the cell’s translation tools can cause certain genes to hog the shared machinery, starving others and slowing overall growth. Evolution instead appears to favor a controlled, slight mismatch that delivers enough protein from important genes without overtaxing the system. This insight not only changes how we think about supposedly “silent” mutations, but also offers practical guidance for designing safer and more efficient genes in biotechnology and medicine.
Citation: Chen, F., Liu, Y., Zhou, Z. et al. A slight mismatch between a gene’s codon usage and the cellular tRNA supply is beneficial. Nat Commun 17, 3371 (2026). https://doi.org/10.1038/s41467-026-69643-2
Keywords: codon usage bias, tRNA supply, synonymous mutations, protein translation, antibiotic resistance