Clear Sky Science · en
H3K4me1 directs H3K36me2 and H3K36me3 deposition in land plants
How plants remember and respond
Plants cannot run away from drought, heat, or changing seasons. Instead, they rely on a hidden layer of chemical marks on their DNA-packaging proteins to fine-tune which genes turn on or off. This study uncovers how one such mark in land plants acts like a molecular signpost, helping place another mark that shapes growth, flowering time, and responses to the environment.

A code written on DNA packaging
Inside every plant cell, DNA is wrapped around proteins called histones, forming bead-like structures along the genetic thread. These histones carry tiny chemical tags that together act as a code for gene activity. The team focused on two of these tags on histone H3, known by shorthand as H3K4me1 and H3K36me2/3. In animals, H3K4me1 is famous for marking stretches of DNA that boost gene activity, called enhancers. But in plants, its role has been puzzling. By comparing algae, mosses, flowering plants, yeast, flies, mice, and humans, the researchers showed that land plants have a distinctive pattern: H3K4me1 spreads across plant gene bodies and shows a unique relationship with how strongly genes are expressed.
A reader protein that spots the first mark
Chemical tags on histones only matter if other proteins can “read” them. The scientists searched plant genomes for proteins with a particular type of reader unit called a PHD finger, known to recognize H3K4 modifications. They homed in on a rice protein called Early heading date 3 (Ehd3), already known to influence flowering time. Biochemical tests using artificial histone pieces showed that Ehd3 strongly prefers H3K4me1 over other related marks. High-resolution structural work then revealed why: Ehd3 uses a pair of tightly linked PHD fingers to form a narrow pocket that fits a single methyl group on lysine 4, but would clash with bulkier versions. This unusual twin-pocket design makes Ehd3 highly selective for H3K4me1.
From reading to rewriting the code
The next question was what happens after Ehd3 binds H3K4me1 on chromatin. Using protein fishing experiments in rice, the team found that Ehd3 physically associates with SDG724, an enzyme that adds methyl groups at another position on histone H3, creating H3K36me2 and H3K36me3. Plants lacking either Ehd3 or SDG724 flowered late and showed similar shifts in gene activity across the genome. Mapping histone marks revealed that losing Ehd3 or SDG724 leaves H3K4me1 largely intact but strongly reduces H3K36me2/3 over affected genes. In test-tube assays, SDG724 showed little activity on its own, but worked far better on H3K4me1-marked nucleosomes when Ehd3 was present, indicating that Ehd3 not only brings SDG724 to the right place but also boosts its catalytic power.
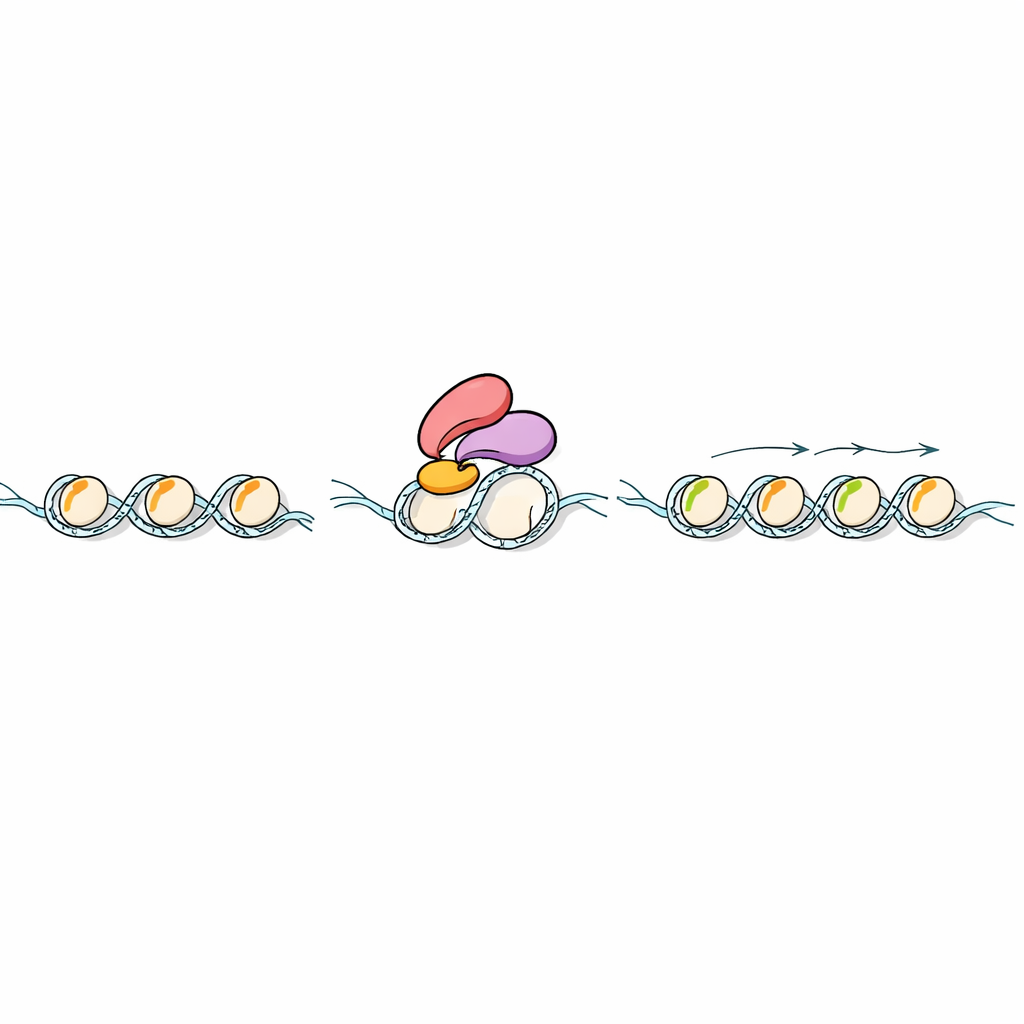
An evolutionary thread across land plants
The researchers extended their analysis beyond rice. They showed that close Ehd3 relatives in other land plants share similar PHD finger structures and, in many cases, the same preference for H3K4me1. Genome-wide data from moss, Arabidopsis, rice, and additional species revealed a recurring pattern: wherever H3K4me1 is found along genes, H3K36me2 and H3K36me3 tend to sit in the same regions. This tight pairing is much weaker or absent in algae and animals, suggesting that, as plants colonized land, they evolved a dedicated H3K4me1 reader to help establish H3K36 methylation. The result is an integrated marking system that likely helps plants fine-tune transcription during development and under stress.
Why this hidden system matters
For non-specialists, the key message is that plants use a layered chemical code on their DNA-packaging proteins to coordinate when genes are active. This work shows that one mark, H3K4me1, acts as a starting signal that recruits a reader protein, Ehd3, which in turn activates an enzyme to install a second mark, H3K36me2/3, along genes. Together, these linked marks shape plant growth, flowering time, and stress responses. Understanding this chain of events opens doors to breeding or engineering crops that can better adapt to changing environments by tweaking how they read and rewrite their own chromatin code.
Citation: Wu, J., Wang, J., Du, K. et al. H3K4me1 directs H3K36me2 and H3K36me3 deposition in land plants. Nat Commun 17, 2831 (2026). https://doi.org/10.1038/s41467-026-69632-5
Keywords: plant epigenetics, histone methylation, chromatin regulation, flowering time, stress-responsive genes