Clear Sky Science · en
Enabling global-scale nucleic acid repositories through versatile, scalable biochemical selection from room-temperature archives
Why storing genetic material matters
Imagine being able to track new viruses, study rare diseases, and preserve entire ecosystems’ genetic footprints without needing rows of energy-hungry freezers. Today, most DNA and especially fragile RNA must be kept extremely cold, which is costly and impractical for many clinics and laboratories around the world. This paper presents a new way to store and search through vast collections of genetic samples at room temperature, while still being able to quickly find just the samples scientists need for testing.
From freezer farms to tiny capsules
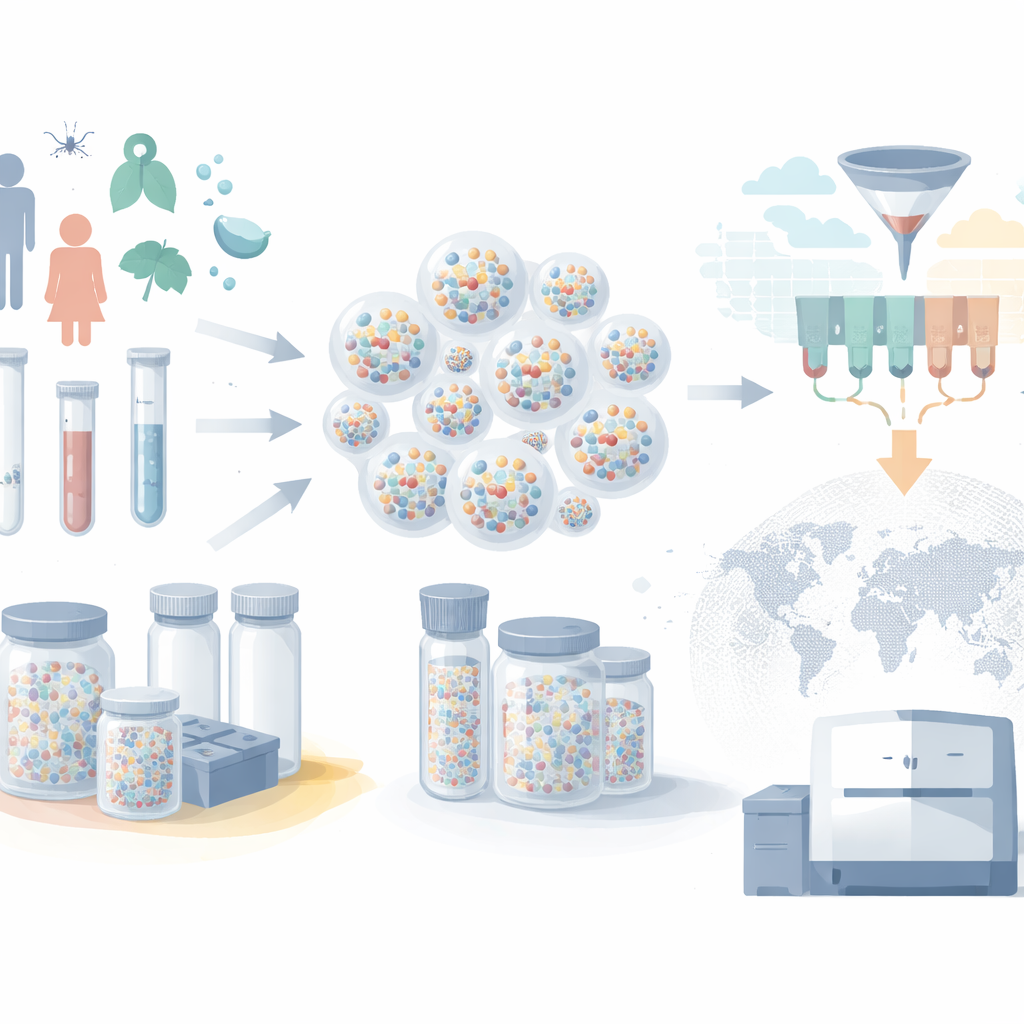
Modern biobanks hold millions of blood, saliva, and tissue samples, but typically each sample lives in its own labeled tube inside a large freezer. As collections scale up, this “one tube per sample” approach becomes expensive and slow. Freezers demand constant electricity, robotic systems move samples one by one, and RNA can degrade if anything goes wrong. The authors instead use microscopic silica shells—microcapsules—to protect genetic material. DNA and RNA are trapped inside these durable particles, which remain stable at room temperature. Many different nucleic acids can be stored together in the same capsule, and huge numbers of capsules can be pooled into a single small tube, slashing storage space and energy needs.
Turning tubes into a searchable database
Storing samples is only half the problem; researchers must also be able to pick out the right subset when new questions arise. The key innovation in this work is a clever barcoding scheme that treats each capsule like a tiny database record. Short pieces of DNA attached to the outside of each capsule act as “tags” describing the sample: for example, patient age, city of origin, month and year of collection, vaccination status, and whether the person was symptomatic. Instead of assigning one unique tag to each possible value, the system reuses a compact set of tags and combines them in different ways, much like how numbers are written with digits or how categories can be encoded using combinations. This “type-aware” design allows simple physical operations on tags to mimic the powerful range and filter queries that digital databases perform.
How to ask questions of molecules
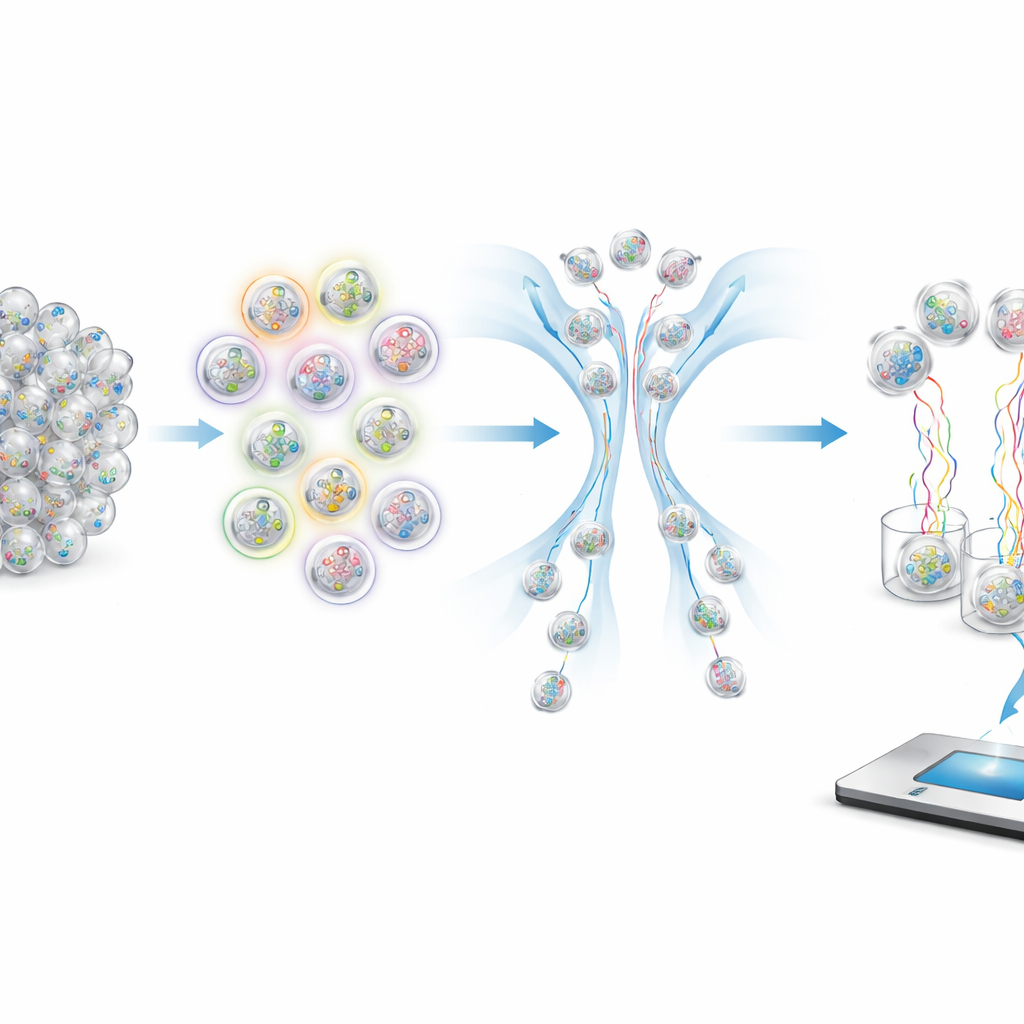
To retrieve specific samples from a crowded tube, the authors use fluorescent probes that stick only to capsules bearing matching tags. These probes carry colored dyes that can be read by a flow-based sorter, which sends brightly glowing capsules down one path and dim ones down another. By choosing which tags to light up and how to combine the colors, the system can implement logical operations like AND, OR, and NOT. For example, selecting “not symptomatic” means rejecting any capsule that glows when tested for the symptomatic tag. Numerical ranges such as age bands or date intervals are handled by ignoring some of the less important “digits” in the encoded age or date, so a single query can pull out all capsules whose tags fall within a chosen window.
Testing the system with a mock outbreak
To show that their approach works in practice, the researchers built a small but realistic test database of 96 synthetic SARS-CoV-2 samples, modeled as if they had been collected from airline passengers arriving in Boston. Each capsule carried fragments of viral RNA plus a unique internal identifier and external tags encoding made-up patient and flight information. They then carried out a series of queries that mirror real-world epidemiology: finding all asymptomatic passengers, selecting specific age ranges to look for variant patterns, and running a complex combined query that pulled out passengers from a particular city, during certain months in 2020, who were either symptomatic or unvaccinated. Sequencing the retrieved capsules confirmed that the right samples had been enriched with very high accuracy, even when the desired group was only a fraction of the whole pool.
Proof that real patient samples can be preserved
Beyond synthetic test cases, the team encapsulated genuine patient-derived SARS-CoV-2 samples carrying different Omicron sub-lineages. After storage and recovery, they sequenced the viral genomes and compared them to the same samples that had never been encapsulated. Despite working with very low amounts of viral RNA, the method still recovered the correct variants, showing that the silica protection does not erase subtle genetic differences. Because non-target capsules can be saved after each search, the pool can be reused repeatedly, and simple internal DNA identifiers allow ongoing health checks on the archive, much like integrity checks in digital storage.
What this means for the future
In plain terms, this study shows that genetic samples can be packed into tiny, durable capsules, piled together in a small space at room temperature, and later pulled apart using flexible, database-like searches. The method dramatically reduces dependence on freezers while allowing rich queries that can combine age ranges, locations, dates, and health states in a single operation. With further scaling, such room-temperature archives could support global pathogen surveillance, precision medicine, and even long-term ecological monitoring in places where cold-chain infrastructure is scarce. The work points toward a future where the world’s biological information can be stored compactly and accessed as easily as files in the cloud.
Citation: Berleant, J.D., Banal, J.L., Rao, D.K. et al. Enabling global-scale nucleic acid repositories through versatile, scalable biochemical selection from room-temperature archives. Nat Commun 17, 2807 (2026). https://doi.org/10.1038/s41467-026-69402-3
Keywords: nucleic acid storage, molecular database, DNA barcoding, pathogen surveillance, room temperature biobanking