Clear Sky Science · en
tRF-3005a regulates exon skipping of SPAG4 by interacting with RALY to drive gastric cancer progression
Why this stomach cancer study matters
Stomach cancer remains one of the world’s deadliest cancers, in part because it is often discovered late and current treatments do not work for everyone. This study uncovers a previously hidden layer of control inside tumor cells, involving tiny RNA fragments and a protein that edits gene messages. By revealing how this molecular partnership helps stomach tumors grow, the work points to new early-warning markers and treatment targets that might one day improve patient survival.
Tiny RNA pieces with big influence
Researchers have long thought of transfer RNAs as simple helpers in building proteins. In recent years, however, small pieces cut from these molecules—called tRNA-derived fragments—have emerged as powerful regulators in cancer. The authors focused on one such piece, tRF-3005a, after sequencing tumor samples from patients with gastric cancer. They found that tRF-3005a was much more abundant in cancer tissue than in nearby healthy stomach tissue, and higher levels were linked with larger tumors, deeper invasion, lymph node spread, and poorer overall survival. Imaging experiments showed that, unlike in normal stomach cells, tRF-3005a in cancer cells sits mainly in the nucleus, the command center where gene messages are processed.
Driving growth, invasion, and spread
To test what tRF-3005a actually does, the team increased or blocked its levels in a panel of gastric cancer cell lines. When tRF-3005a was boosted, cancer cells grew faster, formed more colonies, and became more capable of invading through artificial membranes and migrating across surfaces—behaviors that mirror tumor expansion and spread in the body. When tRF-3005a was suppressed, these aggressive traits were sharply reduced. In mice, implanted tumors made from cells with lowered tRF-3005a stayed smaller and showed reduced activity of key growth-related proteins, confirming that the RNA fragment promotes tumor growth not only in dishes but also in living organisms.
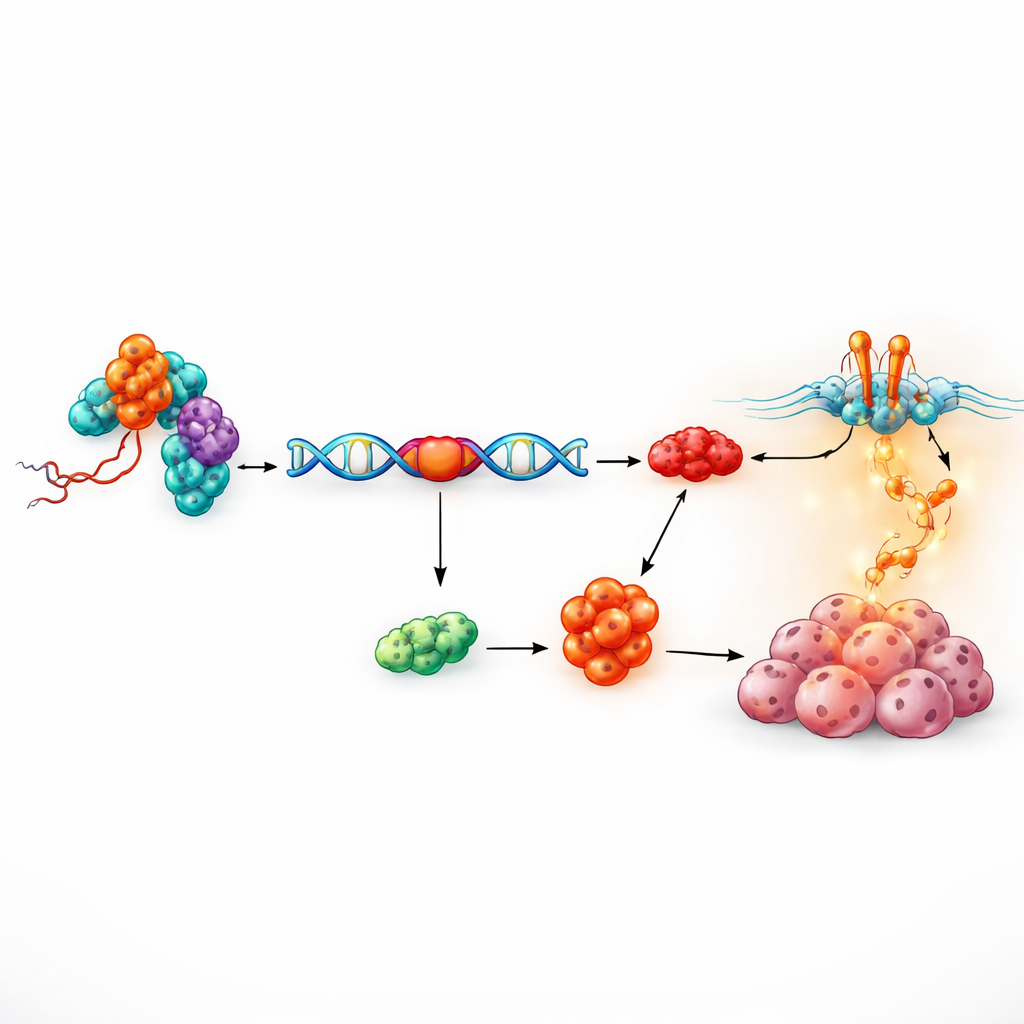
A protein partner that edits gene messages
The next question was how a 17-nucleotide RNA fragment could have such strong effects. Using a biochemical fishing approach, the researchers pulled down proteins that physically bind to tRF-3005a. Among 81 candidates, one nuclear protein—RALY—stood out. RALY is part of a family of factors that shape how raw gene messages are cut and stitched together, a process known as alternative splicing. Detailed mapping showed that tRF-3005a grips a flexible middle region of RALY without changing its overall amount, suggesting it tunes RALY’s behavior rather than simply turning it on or off. Functional rescue experiments, where RALY levels were raised or lowered, showed that the growth-promoting effect of tRF-3005a strictly depends on RALY, defining a tRF-3005a–RALY axis that fuels cancer cell proliferation.
Rewiring a single gene to favor a harmful form
To see which gene messages are reshaped by this axis, the team sequenced mRNA from cells where tRF-3005a was blocked and analyzed changes in splicing patterns. The most affected events involved skipping of specific exons, and one gene, SPAG4, emerged as a top hit. SPAG4 can be produced in two main forms: a long version that includes exon 8 and a short version that leaves this exon out. The researchers showed that RALY binds mainly to the region around exon 8 and suppresses its skipping, pushing cells to make more of the long SPAG4 variant. This long form, but not the short one, drove cancer cell growth. Importantly, tRF-3005a strengthened the physical interaction between RALY and SPAG4 RNA, further reducing exon 8 skipping and tipping the balance toward the tumor-promoting SPAG4-L form.
Link to a major growth pathway in cancer
Downstream of SPAG4-L, the study identified GRB14, a signaling adaptor protein, and the well-known PI3K/AKT pathway, which controls cell survival and growth in many cancers. When SPAG4-L or tRF-3005a was increased, levels of GRB14 and activated PI3K/AKT rose, along with the cancer-driving protein c-Myc. Silencing GRB14 or chemically blocking PI3K/AKT blunted the growth boost caused by tRF-3005a or SPAG4-L, while restoring GRB14 reversed this effect. These results support a chain of events: tRF-3005a binds RALY, RALY reshapes SPAG4 splicing to favor the long form, and SPAG4-L activates GRB14 and PI3K/AKT signaling, ultimately driving uncontrolled cell division.
What this means for patients
Taken together, the work reveals how a tiny RNA fragment can act as a master switch for a cancer-promoting pathway in gastric tumors. By fine-tuning a splicing protein, tRF-3005a steers a single gene toward a harmful version that feeds a powerful growth signal. Because tRF-3005a is elevated in tumors and linked with worse outcomes, it could serve as a biomarker to flag high-risk patients. In the longer term, drugs or RNA-based therapies that block tRF-3005a, disrupt its interaction with RALY, or shift SPAG4 back toward its safer form might offer new ways to slow or stop stomach cancer progression.
Citation: Cui, H., Yuan, Y., Yin, Y. et al. tRF-3005a regulates exon skipping of SPAG4 by interacting with RALY to drive gastric cancer progression. Cell Death Discov. 12, 169 (2026). https://doi.org/10.1038/s41420-026-03049-3
Keywords: gastric cancer, tRNA-derived fragments, alternative splicing, PI3K AKT signaling, biomarkers