Clear Sky Science · en
Environmental dissemination of multidrug-resistant Escherichia coli across one health interfaces in Mymensingh, Bangladesh
Why germs in water matter to everyone
Antibiotic-resistant “superbugs” are often discussed in hospitals, but they do not stay there. This study shows how dangerous strains of Escherichia coli, a common gut bacterium, are spreading through water used by people, animals, and farms in one district of Bangladesh. By tracing these microbes across hospitals, livestock farms, fish ponds, and a river, the researchers reveal how everyday activities can quietly help resistant germs move through the environment and back to humans.
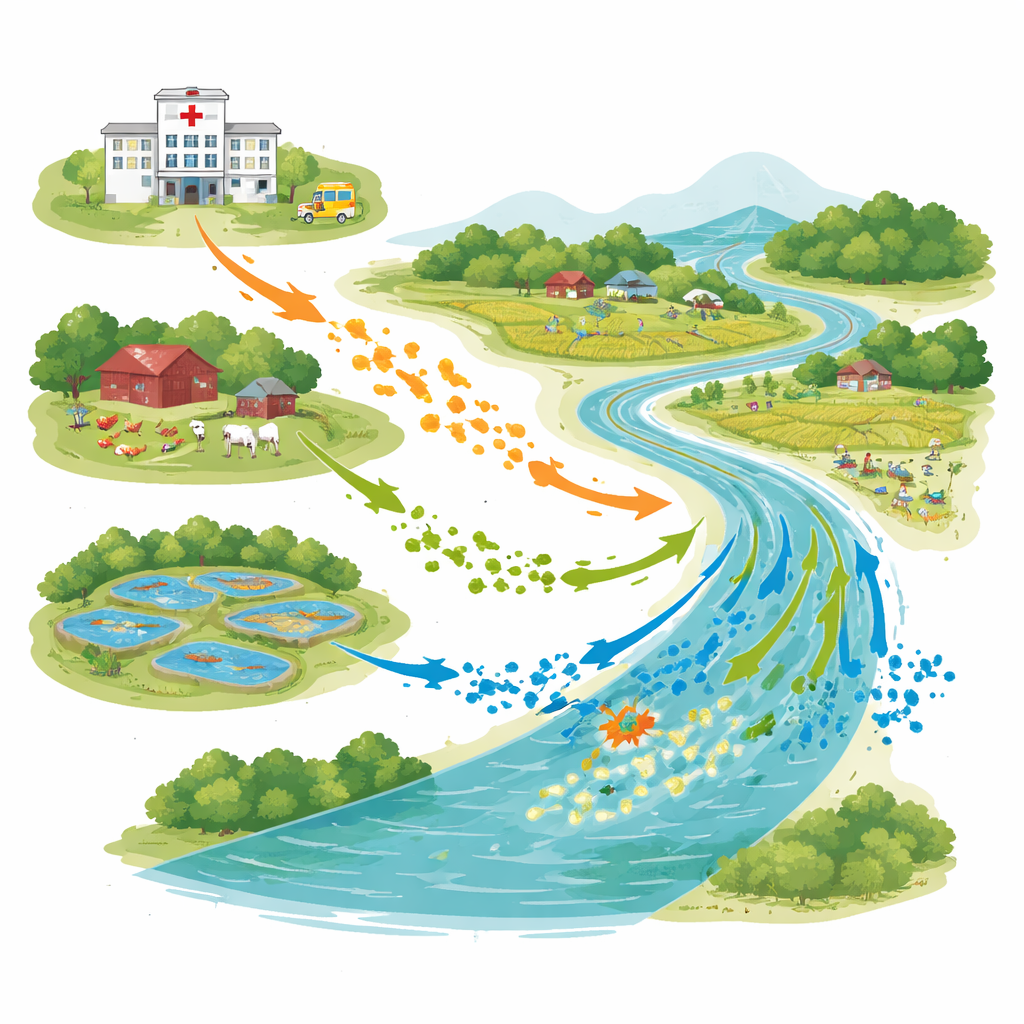
A connected landscape of people, animals, and water
The research took place in Bhaluka Upazila, a semi-urban area where hospitals, poultry and cattle farms, aquaculture ponds, and the Khiro River all sit within the same landscape. In May 2022, scientists collected 28 water samples from hospital sewage outlets, farm effluents, fish ponds, and two points along the river, upstream and downstream from hospital discharges. Using a standard World Health Organization protocol, they counted E. coli in each sample and focused on strains that can defeat powerful antibiotics used for serious infections.
High levels of resistant bacteria in waste and river water
The team found E. coli in every sample and discovered that 86% contained strains resistant to widely used “third-generation” antibiotics. Levels were highest in hospital wastewater and poultry and cattle farm effluents, where both total E. coli and resistant types were extremely abundant. Carbapenem-resistant strains—able to withstand last-resort drugs—were less common but more alarming. They appeared only in hospital sewage and in the river water downstream from hospital discharge points, and were completely absent from livestock and aquaculture ponds. Fish ponds overall showed much lower contamination, suggesting fewer antibiotic inputs there.
Reading the DNA fingerprints of resistance
To understand how these bacteria are related, the researchers sequenced the genomes of 26 representative isolates. They detected 93 different resistance genes, with a handful occurring again and again across sites. One gene family, known as blaCTX-M-15, which blocks many important beta-lactam antibiotics, dominated in all environments. Another gene, blaNDM-5, confers resistance to carbapenems, some of the strongest drugs in modern medicine. This gene turned up in hospital samples and in a downstream river isolate, linked to small, mobile DNA loops called plasmids that can shuttle between bacteria.
Genetic links between hospital drains and the river
By comparing tiny DNA differences—single-letter changes in the genome—the scientists built a family tree of the E. coli they collected. Most bacteria clustered by source: poultry isolates grouped together, cattle isolates formed their own branch, and aquaculture strains sat apart with fewer resistance traits. Hospital isolates were more diverse but, crucially, one hospital strain and a downstream river strain shared an almost identical genetic backbone, with zero differences in the core of their genomes. Although classified as slightly different sequence types, this near-perfect match, combined with the presence of the same carbapenem-resistance gene on plasmids, points strongly to recent spread from hospital effluent into the river.
What this means for health and policy
This work offers a close-up snapshot of how resistant bacteria move across the “One Health” interface linking people, animals, and the environment. It shows that hospital wastewater is a major hotspot, releasing highly resistant E. coli into a river that local communities depend on. Livestock systems add large numbers of resistant strains as well, though not yet those resistant to last-resort drugs. Even with a modest number of samples taken in a single month, the findings support urgent action: better treatment of hospital sewage, more careful antibiotic use in farms and clinics, and routine monitoring of rivers and other shared water sources. For lay readers, the key message is that choices about antibiotic use and waste management do not just affect hospital patients—they shape the invisible flow of superbugs through the environment that we all share.
Citation: Rahman, A., Roy, S., Afreen, N. et al. Environmental dissemination of multidrug-resistant Escherichia coli across one health interfaces in Mymensingh, Bangladesh. npj Antimicrob Resist 4, 27 (2026). https://doi.org/10.1038/s44259-026-00202-x
Keywords: antimicrobial resistance, environmental contamination, wastewater, Escherichia coli, One Health