Clear Sky Science · en
Multimodal data-based graph convolutional networks for predicting outcomes in ovarian cancer receiving neoadjuvant chemotherapy
Why this research matters to patients
For women facing advanced ovarian cancer, choosing the right treatment path can be a life‑altering decision. Many receive chemotherapy before surgery to shrink tumors and make operations safer, but doctors still struggle to predict who will benefit most and who might need a different strategy. This study presents a new, more accurate way to forecast both survival and surgical success using information that hospitals already collect, potentially bringing more tailored care within reach of everyday clinics.
Seeing the whole patient, not just one number
Traditionally, doctors have leaned on a few single measurements to guide treatment before surgery. One popular example is a blood test pattern called the KELIM score, which tracks how quickly a tumor marker called CA‑125 falls during early chemotherapy. Other tools look at scan findings or simple lab ratios such as white blood cell counts. While useful, each of these focuses on only one slice of a patient’s story. The new approach recognizes that real patients are more complex: blood tests, scan details, age, and tumor features all interact in subtle ways that can shape how someone responds to treatment.
Linking similar patients into a learning network
The researchers analyzed data from 853 women with advanced ovarian cancer who all received chemotherapy before interval surgery at three hospitals. For every patient, they used routine laboratory results and detailed features extracted from pretreatment CT scans, such as the shape and texture of visible tumors. Instead of treating each person as an isolated case, they built a "population graph" in which each patient is a point, and connections link patients with similar lab profiles. A specialized form of artificial intelligence, called a graph convolutional network, then learned patterns across this web, using the CT‑based features as the main descriptors for each node while the connections carried information about how patients resembled each other.
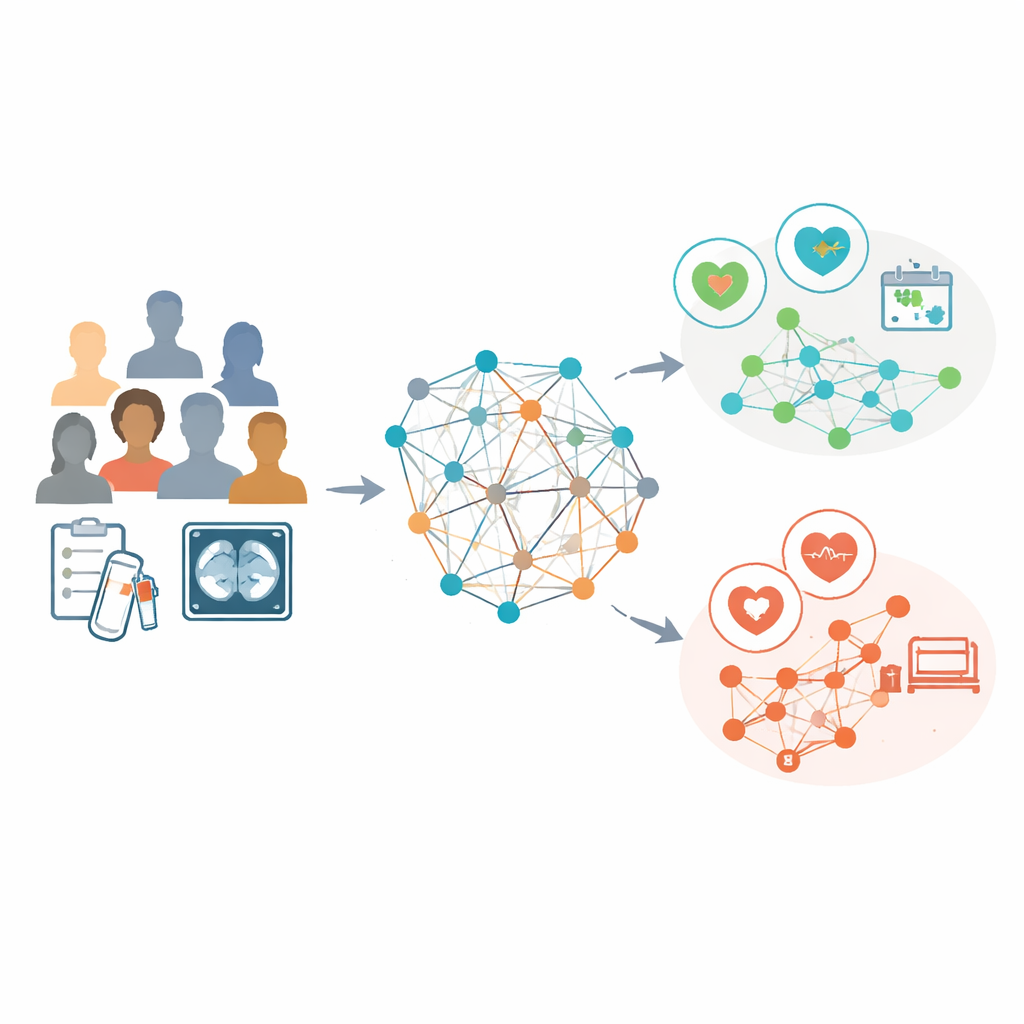
Outperforming current prediction tools
The team asked their model to do two things that matter greatly to patients and surgeons: predict overall survival over several years and estimate the chance of achieving an operation that leaves no visible tumor behind (called R0 resection). They compared its performance against four established approaches built from clinical data, scan features alone, both combined, and the KELIM score, as well as a commonly used surgical scoring system. Across the main hospital and two independent test hospitals, the graph‑based model consistently produced more accurate survival forecasts. It also did a better job than the other methods at distinguishing which patients were likely to end surgery tumor‑free and which were not, even when traditional models showed little separation.
Finding hope where older scores see risk
One of the most striking findings was that the new model uncovered a sizeable group of women—about 16 percent of the total—who had low KELIM scores (which usually signal poor response) but actually went on to live as long as those with favorable KELIM values. In other words, older tools would have labeled them as high‑risk, while the graph‑based method correctly recognized their good outlook. The model also showed a smooth, stepwise relationship between its score and outcomes: as the score rose, both survival time and the chance of complete tumor removal climbed steadily, from almost no R0 surgeries at the lowest scores to nearly all patients achieving R0 at the highest. This steady pattern was not seen with KELIM alone, suggesting the new score captures a clearer biological signal.
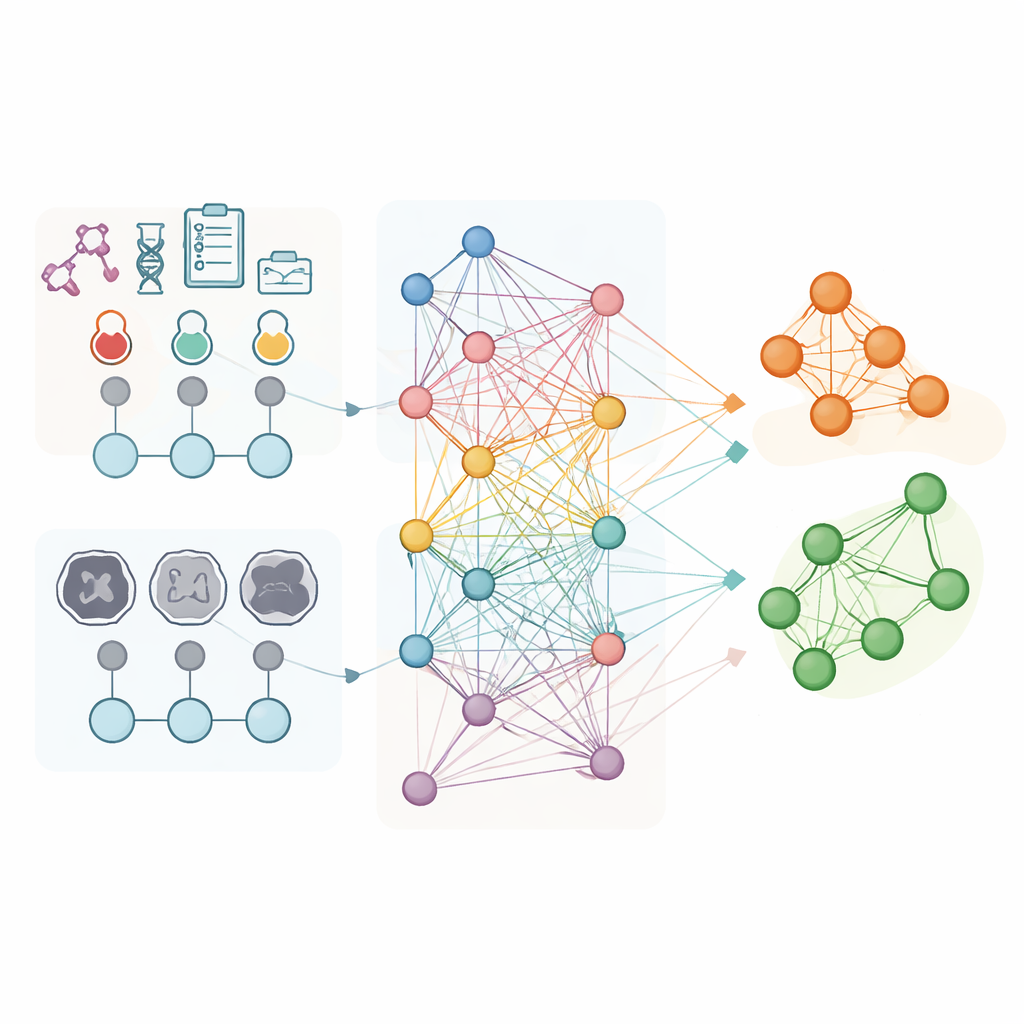
Designed for real‑world hospital use
Many advanced AI tools demand powerful graphics processors that are rarely available in standard hospital computers. In contrast, this graph‑based model was built to run entirely on ordinary central processors, making it more practical for everyday clinical use. It relies on lab tests and CT scans that are already part of routine care, and the authors showed that its predictive value remained strong even after accounting for age, tumor stage, type, and the KELIM score. Although the study was retrospective and did not yet include genetic markers such as BRCA status, its multicenter validation suggests that the method is robust across different hospitals and patient groups.
What this could mean for future care
For patients, the core message is straightforward: by learning from patterns across many similar individuals, this new tool can more accurately identify who is likely to benefit from chemotherapy before surgery and who may need a different plan. A high score signals a strong chance of longer survival and successful tumor removal, while a low score could prompt doctors to reconsider timing of surgery or explore alternative therapies rather than persisting with a strategy unlikely to help. With further prospective testing and integration into hospital software, graph‑based models like this one could help move ovarian cancer treatment away from one‑size‑fits‑all decisions and toward care that is more precisely matched to each woman’s unique profile.
Citation: Zhang, S., Liu, Y., Liu, Z. et al. Multimodal data-based graph convolutional networks for predicting outcomes in ovarian cancer receiving neoadjuvant chemotherapy. npj Precis. Onc. 10, 157 (2026). https://doi.org/10.1038/s41698-026-01346-9
Keywords: ovarian cancer, neoadjuvant chemotherapy, graph neural networks, radiomics, treatment prediction