Clear Sky Science · en
Mobile intron RNA from a bacterial predator accumulates in dead archaeal cells
Tiny Predators and Hidden Messages
In the dark, oxygen‑free corners of our planet, microscopic predators hunt other microbes in slow‑motion battles that shape how carbon is recycled and methane is produced. This study looks at an unexpected traveler that moves during those encounters: a piece of genetic RNA from a bacterial predator that ends up inside dead cells of a different domain of life, the archaea. The work offers a rare real‑time glimpse of how genetic elements may jump between distant branches of the tree of life, feeding long‑standing ideas about “horizontal” gene transfer and the ancient RNA world.
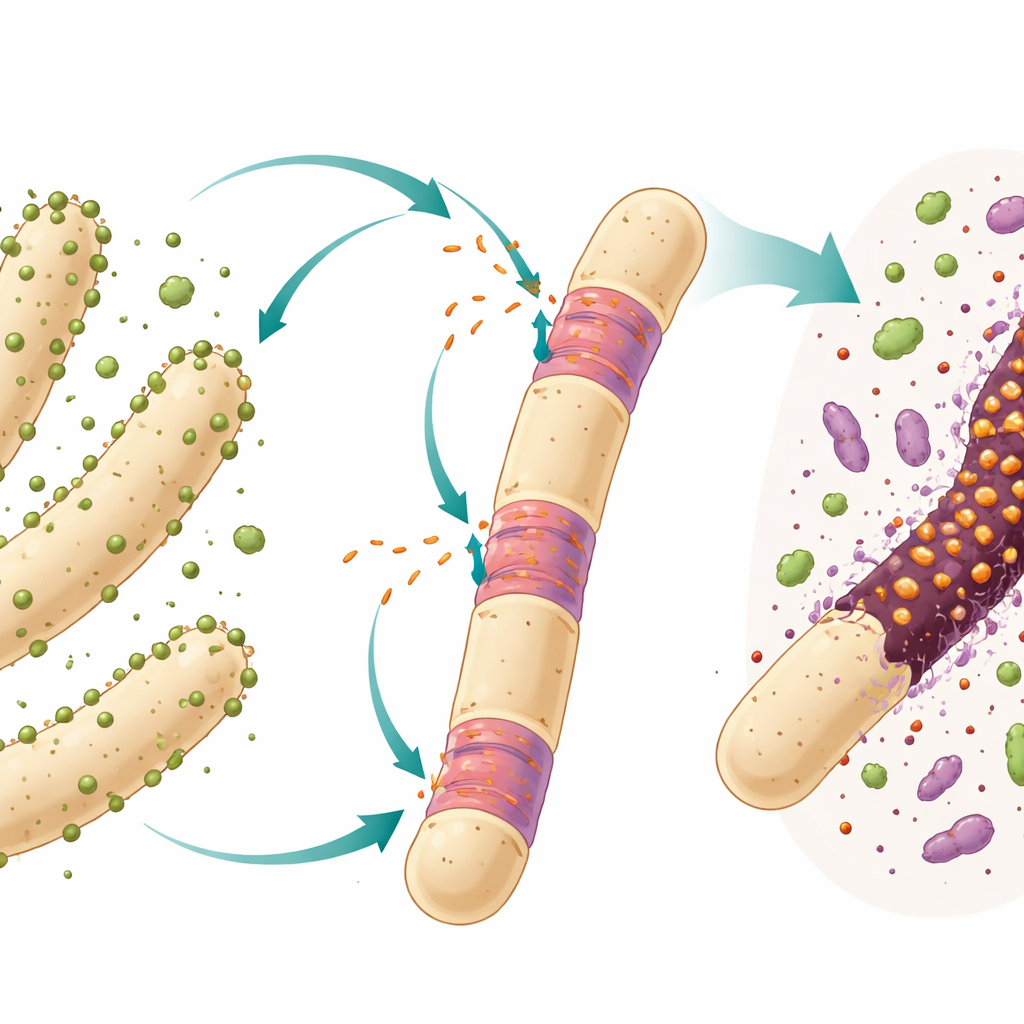
Jumping Genes in Simple Cells
Many bacteria and archaea carry genetic stowaways called introns, stretches of DNA that are copied into RNA and then cut out again before a working molecule is produced. In complex organisms, introns are routine features of genes, but in simple cells they are rarer and often behave like mobile freeloaders that can move around and invade new spots in the genome. The focus here is a particular intron of “group I” type sitting inside the 23S ribosomal RNA gene of an ultratiny bacterial predator named Candidatus Velamenicoccus archaeovorus. This bacterium lives stuck to long filament‑shaped archaeal cells in a long‑running laboratory culture that breaks down the plant compound limonene and slowly produces methane.
A Slow, Closed Microbial World
The researchers worked with an anaerobic enrichment culture that has been maintained for over twenty years on limonene with only one transfer per year. In this closed system, different microbes cooperate and compete. One bacterial partner breaks down limonene into smaller compounds, and several methanogenic archaea, including the filament‑forming species Methanothrix soehngenii, turn those products into methane. The ultramicrobacterium Ca. Velamenicoccus archaeovorus lives as an epibiont, attached to the surface of these filaments. Earlier work suggested it acts as a predator: some filament cells appear dead yet still contain DNA and lipids, implying that the predator extracts key cellular materials while leaving a partial shell behind.
Seeing Foreign RNA Inside Dead Cells
To test whether the predator sends its intron RNA into its archaeal victims, the team used a sensitive imaging technique called CARD‑FISH, which uses short labeled DNA probes to light up matching RNA molecules inside preserved cells. They designed three probes that recognize the intron RNA and combined them with probes for ribosomal RNA of the predator and for DNA staining. Under the microscope, the intron signal appeared both in the tiny spherical predator cells and, crucially, inside certain segments of the large Methanothrix filaments. These filament cells showing intron signal had lost their own ribosomal RNA, a hallmark of death, but still contained DNA, confirming they were dead but not yet fully decomposed. Control probes with reversed sequences did not light up, which argues that the signal was specific to the intron.
Counting Rare RNA Travelers
To complement the images, the authors re‑examined a large RNA sequencing dataset previously generated from the same culture. Because ribosomal RNA had not been removed from this sample, they could directly compare how often the intron sequence appeared relative to the normal, fully processed 23S ribosomal RNA. They found that reads matching the intron were about 1 in 20,000 compared with the mature 23S RNA, meaning most intron copies are successfully cut out of the primary transcript. A handful of reads bridged the boundaries between intron and its neighboring sequence, indicating that a very small fraction of transcripts remained unspliced. Together with the imaging, this showed that excised intron RNA molecules exist in the culture and can be found outside their original host cell.
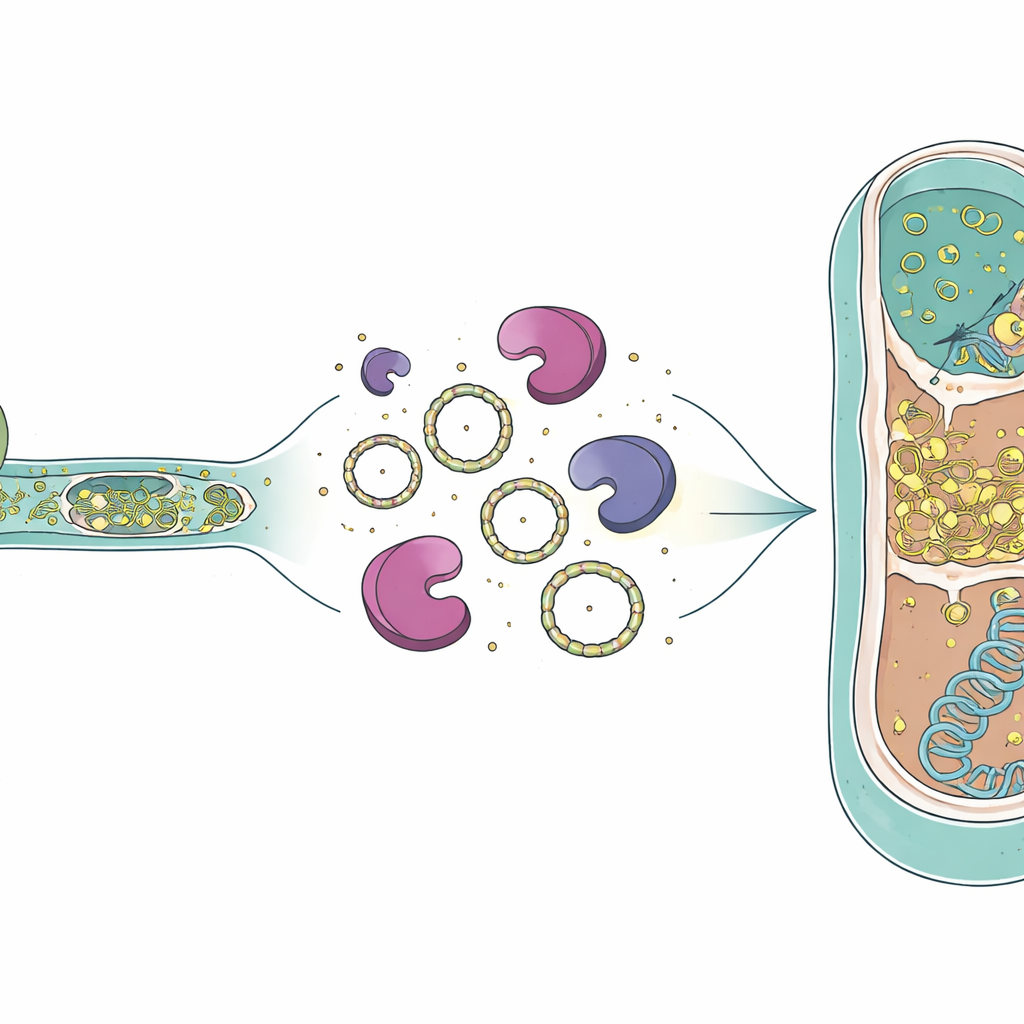
How the RNA May Survive and Move On
The presence of intron RNA inside dead archaeal cells raises questions about how these molecules persist long enough to matter. Previous work has shown that group I introns can form circular RNA rings after splicing, and such circles are more resistant to the enzymes that usually chew up RNA. The predator’s genome also encodes a special reverse transcriptase—an enzyme that can copy RNA back into DNA—which was previously detected as a protein in the same culture. Electron microscopy images from earlier studies showed open cytoplasmic contacts between predator and victim, suggesting that not only RNA but possibly this enzyme could pass into the archaeal cell. If circular intron RNA and a reverse transcriptase enter a victim cell together, they could in principle be copied into DNA and inserted into the archaeal genome, creating a new intron in a new host.
Why This Matters for Life’s Genetic Traffic
For non‑specialists, the main takeaway is that this study provides direct, microscopic evidence that a mobile piece of bacterial RNA can leave its home cell and accumulate inside the dead remains of a very different microbe. That movement is a key early step in horizontal gene transfer, the process by which genes and genetic elements spread across species lines. The work also broadens the known roles of extracellular RNA, which already includes signaling and growth inhibition, by adding mobile intron RNA to the mix. In the bigger picture, such mobile RNAs and their associated enzymes help explain how genetic innovations have been shuffled through microbial communities over evolutionary time, blurring the boundaries between separate branches of life.
Citation: Kizina, J., Lonsing, A. & Harder, J. Mobile intron RNA from a bacterial predator accumulates in dead archaeal cells. Sci Rep 16, 14654 (2026). https://doi.org/10.1038/s41598-026-51721-6
Keywords: mobile introns, extracellular RNA, microbial predation, horizontal gene transfer, methanogenic archaea