Clear Sky Science · en
Integrative structural and physicochemical characterization of chalcone synthase enzymes from medicinal plants using AlphaFold, molecular docking, and molecular dynamics
How Plants Make Helpful Natural Chemicals
Many of the health-boosting compounds in teas, herbs, and fruits come from a single crucial plant enzyme called chalcone synthase. This molecular workhorse helps plants manufacture flavonoids—natural chemicals with antioxidant, anti-inflammatory, and even anticancer activity. The study behind this article uses modern computer tools, including AlphaFold, to examine chalcone synthase from a range of medicinal plants, asking a deceptively simple question: how similar are these enzymes, and what does that mean for harnessing them in nutrition, medicine, and biotechnology?
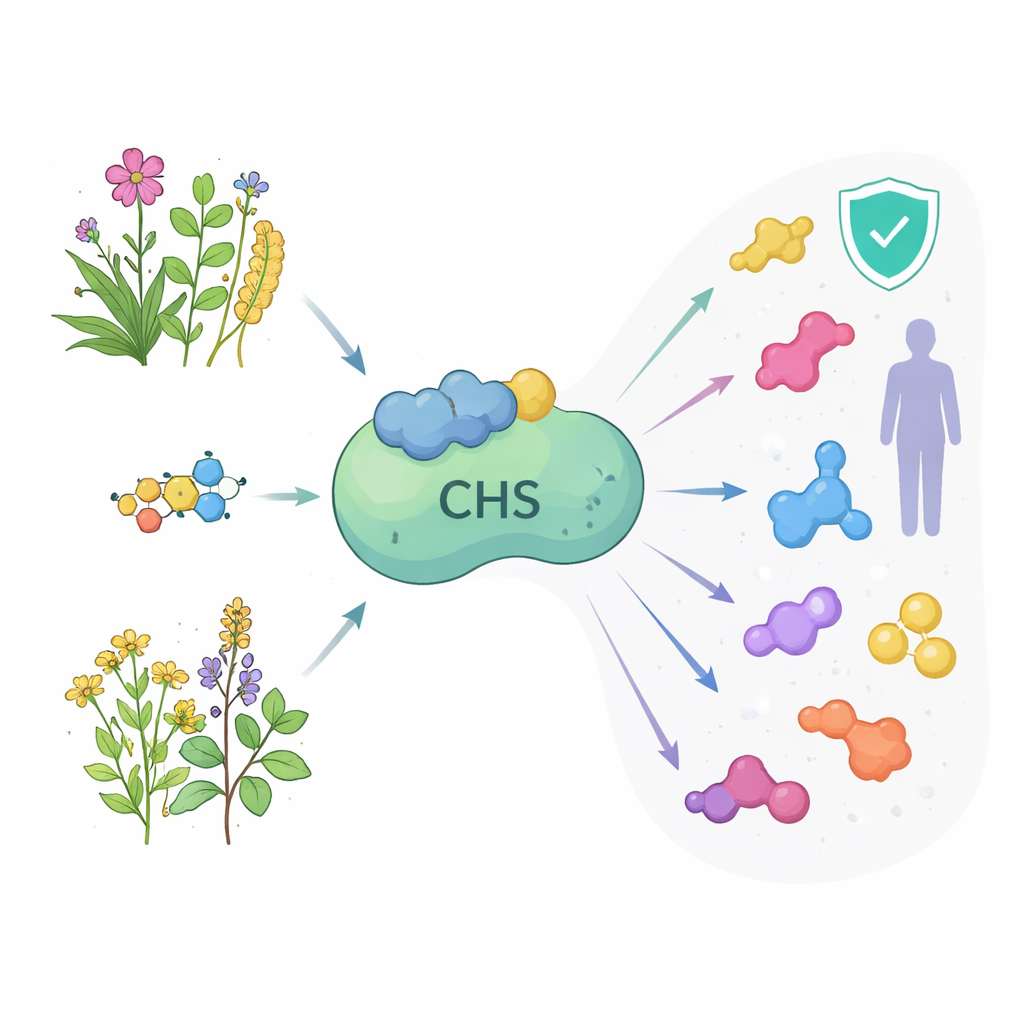
The Enzyme at the Start of a Busy Assembly Line
Chalcone synthase sits at the first committed step of the flavonoid assembly line. It takes in a starter molecule derived from common plant metabolism and stitches it together with three small building blocks to form naringenin chalcone, the gateway to many different flavonoids. These downstream products color flowers and fruits, shield leaves from ultraviolet light, defend against microbes, and act as chemical signals with soil bacteria. In humans, the same molecules are studied for roles in heart health, brain protection, anti-infective therapies, and cancer treatment. Because this one enzyme controls how much material enters the pathway, understanding its form and behavior across medicinal plants could open new ways to boost or redirect valuable natural products.
Looking Across Medicinal Plants
The researchers gathered chalcone synthase protein sequences from 13 medicinal plants, plus the model species Arabidopsis as a reference. They aligned these sequences and built a family tree to see how related they are. Despite the plants coming from many different botanical families, the enzyme’s key features were strikingly conserved: a three-part "catalytic triad" of amino acids and a short signature motif that shape the active tunnel where chemistry happens. Most of the differences between species appeared at the protein’s ends or in loop regions on its surface, not in the core catalytic machinery. This pattern suggests that evolution has tightly protected the basic reaction, while allowing subtle adjustments that may tune how each plant handles its flavonoid chemistry.
What Computer Models Reveal About Shape and Stability
Using AlphaFold and related tools, the team predicted three-dimensional structures for each enzyme and compared them with high-quality crystal structures from two well-studied species. The matches were extremely close—down to less than a tenth of a nanometer in backbone position—confirming that the predicted models are reliable for detailed analysis. All versions of chalcone synthase adopted the same characteristic fold seen in this family of enzymes, but they showed small, species-specific variations in the shape and openness of the tunnel that holds the substrate. Simple calculations also suggested modest differences in features like predicted thermostability, overall charge, and water-loving versus water-repelling character. These traits may influence how easily each enzyme can be produced in the lab or how robust it is inside different cells and environments.
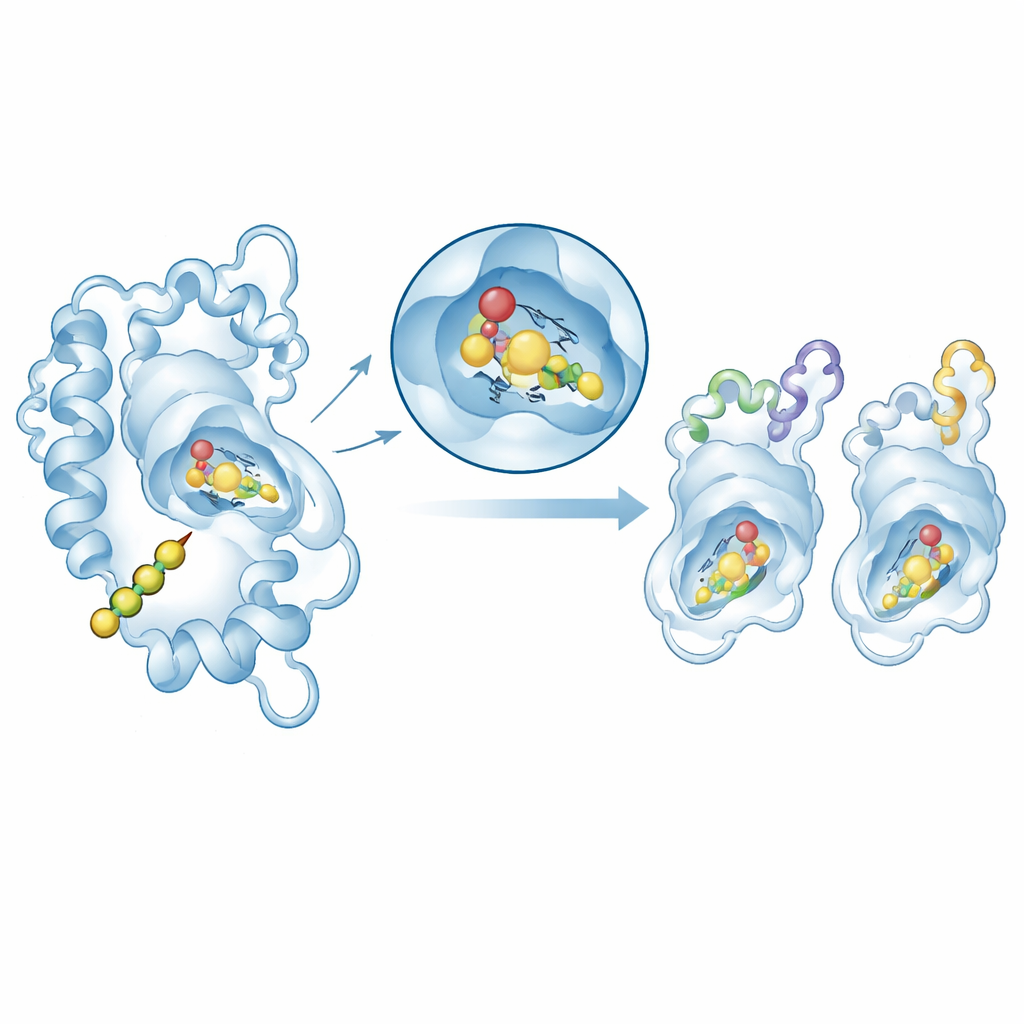
Testing How the Enzymes Grip Their Substrate
To connect structure with function, the authors used molecular docking to place the natural starter molecule, p-coumaroyl-CoA, into the active site of selected chalcone synthase models. In all cases, the substrate nestled into a similar pocket near the conserved catalytic triad, with moderately favorable binding energies that fall in the range typical for enzyme–substrate complexes. A more detailed follow-up on two representative enzymes—one from Arabidopsis and one from the ornamental plant Matthiola—used molecular dynamics simulations to watch the protein–substrate complexes move in virtual water for 100 billionths of a second. Both systems remained structurally stable, and the key active-site region barely wobbled. Energy calculations showed that snug, surface-to-surface contact (van der Waals forces) is the main contributor to binding, supported by electrostatic interactions.
Why This Matters for Future Medicines and Crops
Taken together, the work shows that chalcone synthase from diverse medicinal plants shares a deeply conserved catalytic core but differs in fine structural and physicochemical details around the active tunnel. Those small differences may help explain why different plants produce distinct bouquets of flavonoids, and they offer potential levers for engineering. Although everything here is based on computer models that still need experimental testing, the study delivers a ready-made framework for choosing promising enzyme variants, designing targeted mutations, and running virtual screens of new chalcone-like compounds. In practical terms, that could one day help scientists design crops with enhanced health-promoting flavonoids or streamline the microbial production of plant-derived drugs.
Citation: Muflikhati, Z., Mangindaan, D. & Enyi, C.U. Integrative structural and physicochemical characterization of chalcone synthase enzymes from medicinal plants using AlphaFold, molecular docking, and molecular dynamics. Sci Rep 16, 14624 (2026). https://doi.org/10.1038/s41598-026-45190-0
Keywords: chalcone synthase, flavonoid biosynthesis, medicinal plants, AlphaFold modeling, enzyme engineering