Clear Sky Science · en
Identification and clustering analysis of drug-responsive temporally varying genes through high-frequency longitudinal RNA sequencing
Why watching genes over time matters
When we take a medicine, our bodies don’t respond in a single instant. Cells adjust, fight back, repair damage and recover over hours and days. Yet most lab tests look only before and after treatment, missing what happens in between. This study shows that by taking frequent small blood samples and reading out gene activity day by day, researchers can uncover hidden waves of response to drugs that would otherwise be invisible. The work focuses on drugs that stress the liver in rats, but the approach points toward more precise, time-aware medicine for people.
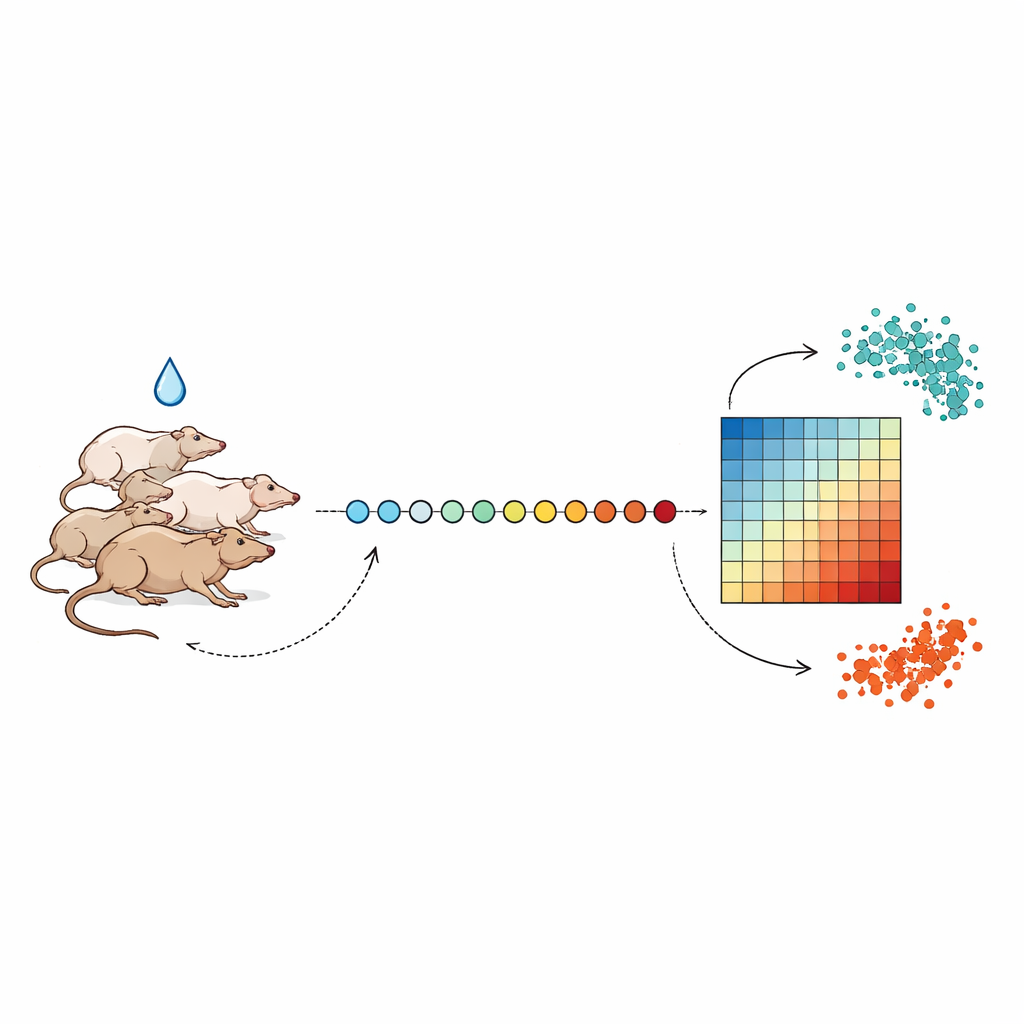
Taking daily snapshots of the body’s reaction
The researchers followed groups of rats for up to three weeks, drawing a very small amount of blood at nearly the same time each day to avoid normal day–night swings in biology. After a few days of baseline sampling, animals received a single dose of one of several liver-toxic compounds, including the antibiotic tetracycline and the tuberculosis drug isoniazid, at different strengths. From each blood sample, the team extracted RNA—the messenger molecules that reflect which genes are turned on or off—and used high-throughput sequencing to measure the activity of thousands of genes at once. By lining these measurements up across days, they could watch how gene activity moved away from a healthy baseline and then, in most cases, gradually returned.
Finding genes that change with time, not just on average
Standard analyses of such data usually compare one time point before treatment with one time point afterward, flagging genes with large average changes as “different.” But that snapshot view can miss genes that briefly spike or dip, or that respond later and then normalize. To tackle this, the authors used a framework to identify “temporally varying genes”–genes whose activity fluctuates in a meaningful way over the course of treatment, even if their starting and ending levels look similar. By scoring how strongly each gene’s levels rose and fell across days, and filtering out routine noise such as mild effects of repeated bleeding, they uncovered thousands of genes whose behavior contained rich timing information about the drug response.
Patterns of early alarm, sustained stress, and late recovery
When the team grouped these time-varying genes by how their activity changed after a high dose of tetracycline, clear patterns emerged. Some genes jumped quickly and then settled back within a few days, acting like early alarm bells. Others rose and stayed high through the peak of drug impact, reflecting sustained stress on protein-making and other core cell processes. A third group turned on or off later, marking slower repair and cleanup phases. Similar, though not identical, timing patterns appeared with other liver-toxic drugs. By linking each pattern to known cellular tasks—such as immune defense, waste disposal, or DNA repair—the authors could sketch a stepwise story of how the liver first senses injury, then adapts, and finally tries to heal.
Shared warning signs and drug-specific fingerprints
Looking across four different toxic compounds, the researchers cataloged more than 4,000 unique time-varying genes. Only 186 of them changed in a coordinated way for every drug, forming a shared “core” signature of liver stress that was especially rich in genes handling iron and a form of cell death called ferroptosis. The rest were drug-specific, highlighting that different chemicals injure cells through distinct routes and trigger different recovery programs. The team also examined how gene responses depended on dose. Some genes reacted even at the lowest drug levels, making them promising early-warning markers, while others responded only to the strongest doses, pointing to pathways that engage when damage becomes severe. These dose thresholds may help explain why the same treatment can lead to mild discomfort in one individual and serious harm in another.
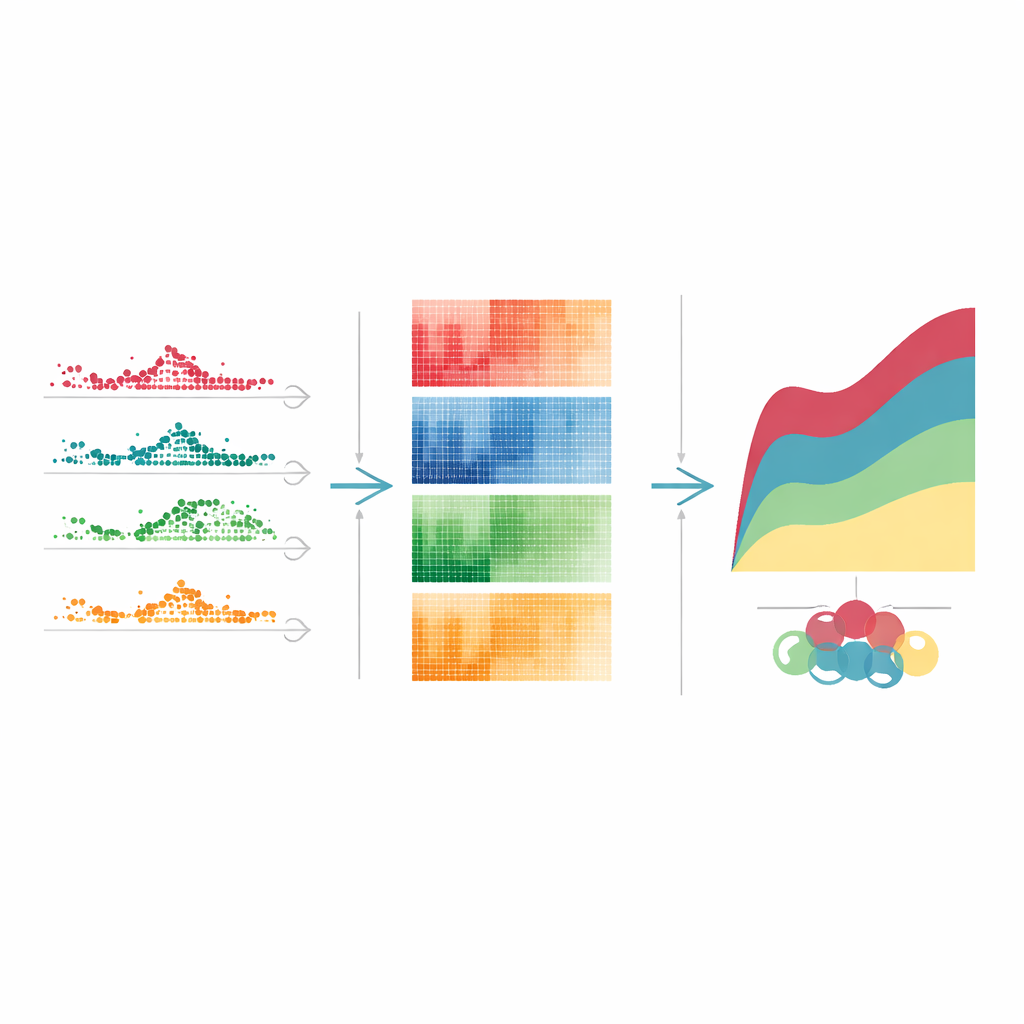
What this means for future treatments
Taken together, the study shows that watching gene activity unfold over time can reveal subtle, fast, or delayed responses that flat, single-time-point tests overlook. The authors argue that these time-varying genes offer a more sensitive way to distinguish direct drug effects from the body’s own attempts to adapt and repair, and to separate safe exposure levels from dangerous ones. While the work was done in rats and focused on liver-toxic compounds, the underlying idea—dense, repeated sampling combined with smart analysis—could be applied broadly. As sequencing becomes cheaper and data analysis more powerful, such longitudinal molecular tracking may help tailor drug dosing, detect side effects earlier, and better match treatments to each person’s unique pattern of response.
Citation: Jiang, Q., Weng, X., Chai, Y. et al. Identification and clustering analysis of drug-responsive temporally varying genes through high-frequency longitudinal RNA sequencing. Sci Rep 16, 14143 (2026). https://doi.org/10.1038/s41598-026-44560-y
Keywords: longitudinal RNA sequencing, drug-induced liver injury, gene expression dynamics, toxicology biomarkers, precision pharmacology