Clear Sky Science · en
AI caption generation model for digital pathology of adenocarcinoma in endoscopic histopathology using multi-instance attention mechanisms
Why turning slides into words matters
When doctors suspect stomach cancer, tiny samples of tissue are taken from the stomach lining and examined under the microscope. Interpreting these slides is a skilled and time‑consuming task for pathologists, and the written report they produce guides every later treatment decision. This study introduces an artificial intelligence (AI) system that does something surprisingly human: it looks at digital images of gastric biopsy slides and automatically writes short, structured descriptions similar to a pathologist’s report.
From glass slide to digital helper
Gastric adenocarcinoma, a common and deadly form of stomach cancer, is currently diagnosed and graded by pathologists who visually study biopsy samples. Over the last decade, many hospitals have begun scanning glass slides into very large digital images, opening the door for AI tools that can assist with diagnosis. Earlier systems focused mainly on yes‑or‑no questions, such as whether cancer is present, or on assigning a category. However, real pathology reports are narrative: they describe cell shapes, tissue patterns, and how aggressive the tumor appears. Only a few research efforts have tried to generate these descriptive captions directly from whole slide images, and many of them depend on extra labels that indicate predefined subtypes, which are expensive and not always available.
A new way for AI to read many small pieces
The authors present a captioning model called MIAC (Multi‑instance Attention Captioning) designed specifically for biopsy slides of gastric adenocarcinoma. A single digital slide is far too large to feed into a model at once, so it is divided into many small square images, or patches. MIAC uses an approach known as multi‑instance learning: instead of treating these patches as an ordered sequence, it views them as a set, allowing the number and arrangement of patches to vary from slide to slide as happens in routine practice. A powerful image network first extracts visual features from each patch. Then a self‑attention mechanism, built without position information, learns how strongly each patch should influence the final summary. The model combines these signals into a single compact representation of the whole slide, which then feeds a language module that generates a sentence‑length diagnostic caption, one word at a time.
Training on real reports, testing in a different hospital
To teach MIAC, the researchers used a public dataset called PatchGastricADC22, which contains nearly a thousand scanned biopsy slides from a Japanese hospital. Each slide is paired with a short diagnostic sentence taken from the original pathology report and converted into standardized terminology. The slides were broken into hundreds of patches, and the model was trained to produce the original caption when given a subset of these patches. Performance was measured with standard language comparison scores that evaluate how closely AI‑generated descriptions match expert text. MIAC outperformed a previous state‑of‑the‑art method across all metrics, especially when it was allowed to see more patches per slide during training, suggesting it was better at capturing the slide‑level picture from scattered local views.
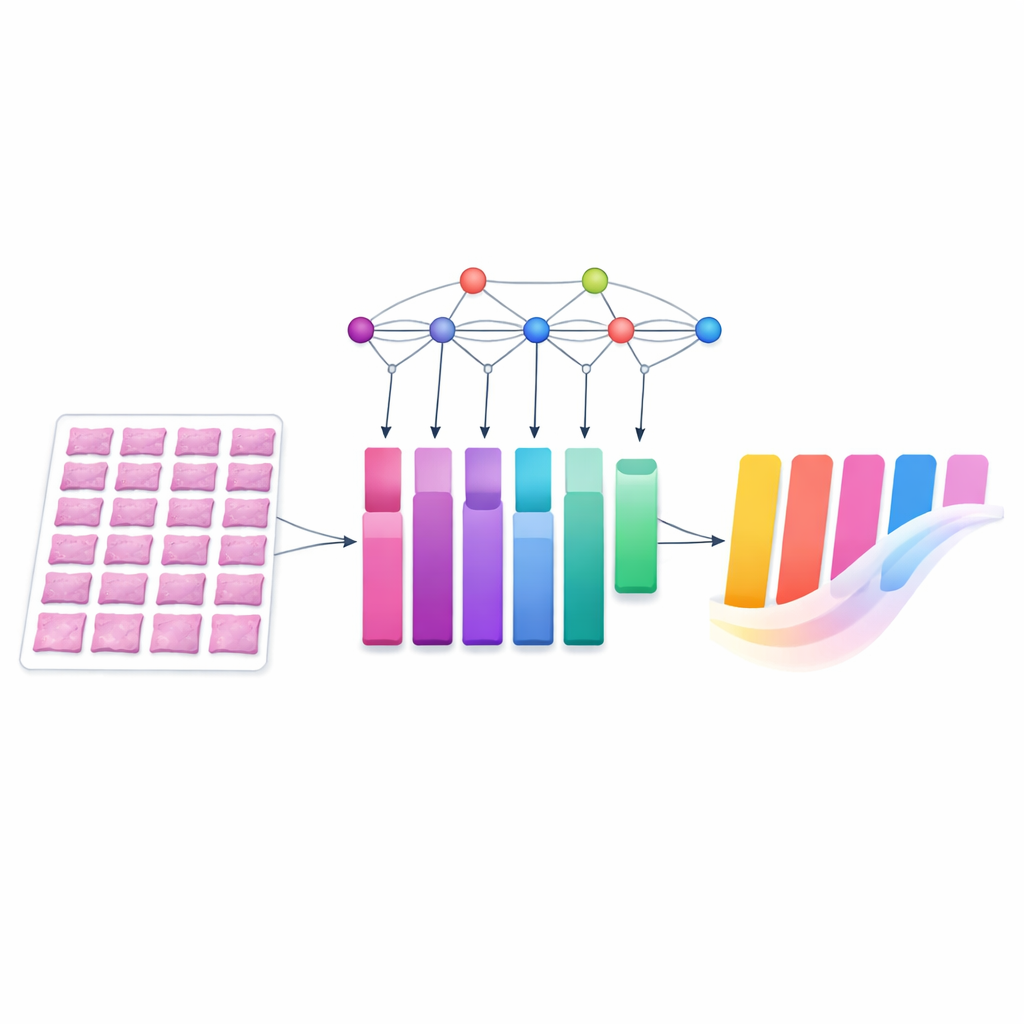

Handling differences between hospitals
A major question for any medical AI system is whether it still works when the data come from somewhere new. The team therefore tested MIAC on an independent set of gastric biopsy slides collected at a different hospital, with captions written by another pathologist. These images differed in color because of local staining and scanning practices, a common issue that can trip up image‑based models. The researchers applied a widely used color‑normalization technique, which adjusts the stain shades to more closely match those of the training slides. Even without any further fine‑tuning, MIAC produced clinically meaningful captions on this external dataset, and its performance improved when color normalization was applied and when more patches were sampled from each slide.
What this could mean for patient care
MIAC is not meant to replace pathologists; instead, it aims to draft concise, standardized descriptions that experts can quickly review and edit. In busy clinics or regions with limited specialist access, such a tool could shorten reporting time, reduce variability in wording, and flag cases with complex patterns for closer attention. The study also highlights current limits: performance still drops when the model is confronted with slides from new institutions or with unusual tissue patterns, and automated language scores do not fully capture clinical usefulness. The authors argue that richer, multi‑center datasets, smarter ways of choosing which patches to inspect, and direct feedback from pathologists will be needed before such caption‑generating systems can be safely woven into everyday diagnostic workflows.
Citation: Lee, Y., Bai, K., Kim, Y. et al. AI caption generation model for digital pathology of adenocarcinoma in endoscopic histopathology using multi-instance attention mechanisms. Sci Rep 16, 13244 (2026). https://doi.org/10.1038/s41598-026-37455-5
Keywords: gastric cancer, digital pathology, medical AI, image captioning, histopathology