Clear Sky Science · en
SMC-LUD:Large-Scale B-Mode Liver Ultrasound Dataset for Hepatocellular Carcinoma and Hemangioma Classification
Why this matters for everyday health
Liver cancer is one of the world’s deadliest cancers, in part because it is hard to spot and classify early enough for effective treatment. Doctors often rely on ultrasound, a quick and affordable scan, to look for suspicious spots in the liver. But on these grainy black-and-white images, dangerous tumors and harmless growths can look confusingly similar, even to experts. This article introduces a large new image collection designed to help computers learn to tell the difference, potentially making liver checks more accurate and more widely available.
A closer look at liver spots
Two common findings on liver scans sit at opposite ends of the risk spectrum. Hepatocellular carcinoma is a primary liver cancer that can be life-threatening if missed or discovered late. Hemangiomas, by contrast, are usually harmless clusters of blood vessels that rarely require treatment. On basic ultrasound images, however, these two can be hard to distinguish, especially when the liver tissue is already damaged or when the spots are small. Today, doctors often need to order extra tests such as CT or MRI scans to be confident, which adds cost, time, and radiation or contrast exposure.
Building a large library of liver images
To tackle this problem, researchers at Samsung Medical Center in Seoul assembled SMC-LUD, a new public collection of 5,385 liver ultrasound images from 1,021 real patients scanned between 2015 and 2024. Each image shows a liver lesion that has been carefully classified into one of two groups: cancer (hepatocellular carcinoma, 2,716 images) or benign hemangioma (2,669 images). Cancer cases were confirmed by examining tissue removed during surgery or biopsy, while hemangiomas were diagnosed using characteristic scan features interpreted by experienced radiologists. All images were anonymized, grouped by patient, and double-checked by specialists to ensure reliable labels. 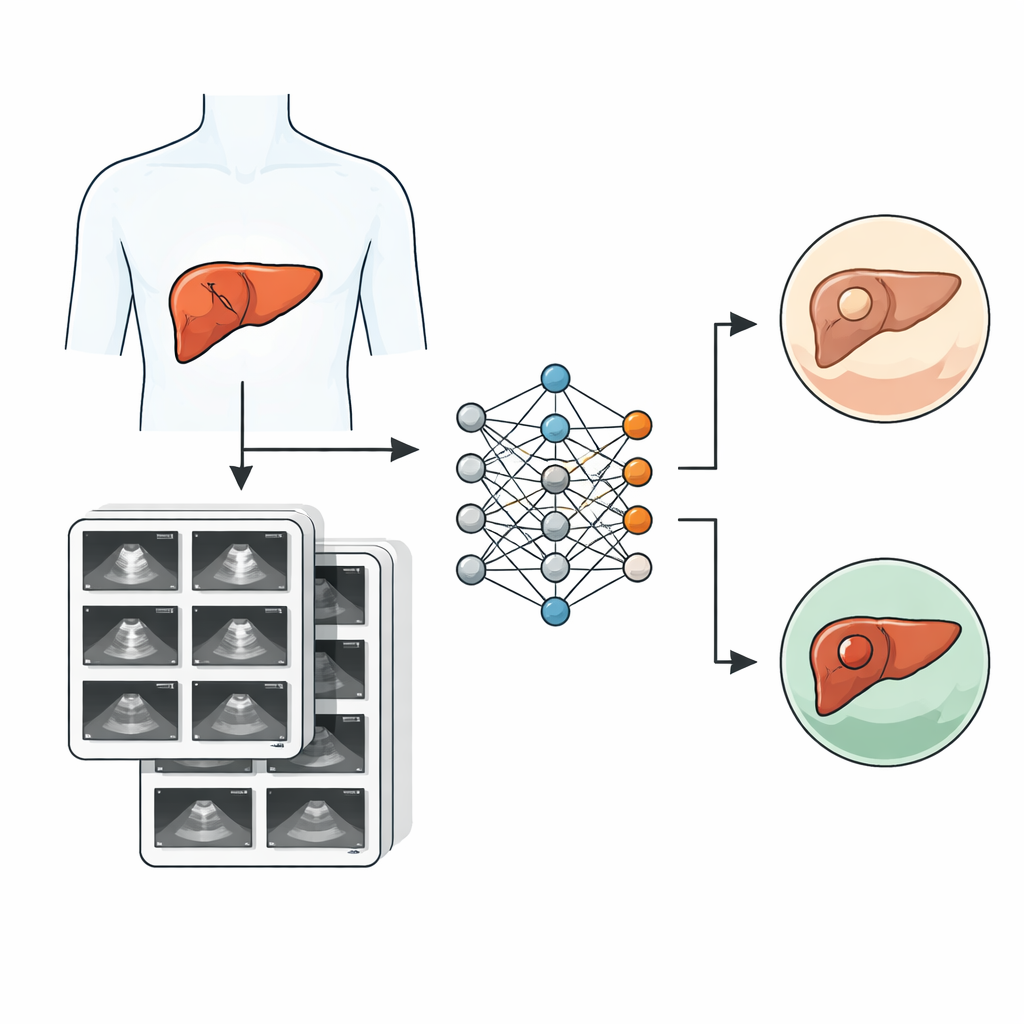
Preparing the data for smart algorithms
The team organized the dataset to make it easy for artificial intelligence researchers to use. Images were resized and standardized so that computer models see them in a consistent format, and the data were split into separate groups for training, fine-tuning, and final testing, with care taken so that images from the same patient never appear in more than one group. The dataset also comes in two flavors: a “Clean” version without measurement markers and a “Caliper” version that includes them. For fair testing, the researchers trained their models only on the Clean images to avoid the risk that algorithms might cheat by learning to associate marker patterns with certain diagnoses instead of focusing on the lesions themselves. A companion table of patient information, such as tumor size and stage for cancer cases, allows deeper clinical analyses.
How the computer learns to see
To show what can be done with this resource, the authors built a deep learning model based on a well-known image-recognition framework and enhanced it with special “attention” modules. These components help the network focus on the most informative parts of the image and the most telling patterns of brightness and texture. The design uses two parallel branches that process the same features in slightly different ways before rejoining and then applies a spatial filter that highlights important regions, much like a radiologist mentally zooming in on a suspicious area. Trained on the SMC-LUD Clean subset, this model was compared with several popular neural network designs for medical imaging. 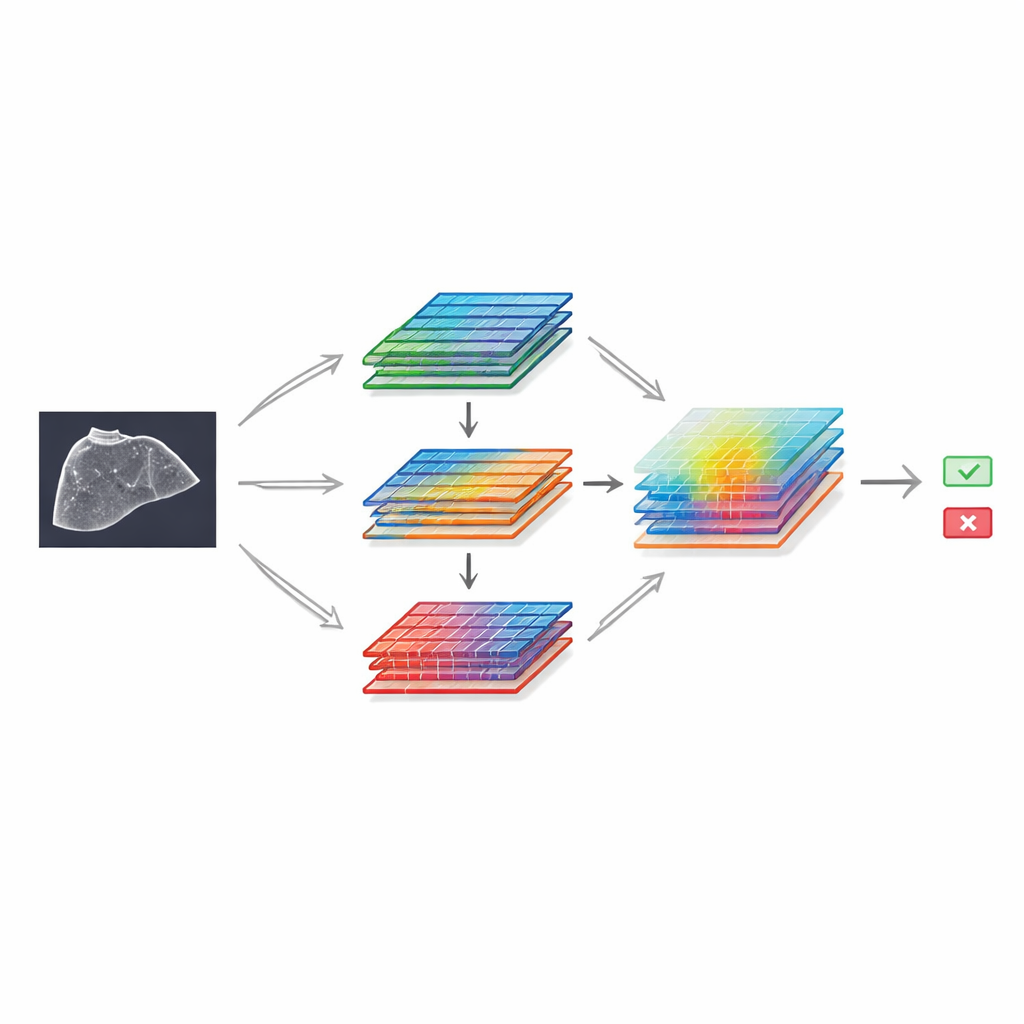
What the results show
On the held-out test images, the attention-enhanced model correctly distinguished between liver cancer and hemangioma nearly 99% of the time, outperforming the comparison models. Heat maps derived from the network’s internal calculations showed that it tended to focus on the lesions themselves rather than on unrelated image details, suggesting that it was learning medically meaningful cues. While this study evaluates only a two-way distinction between one malignant and one benign lesion type, its performance highlights both the quality of the image labels and the potential of ultrasound-based computer diagnosis when supplied with sufficiently large, well-organized data.
What this could mean for patients
For non-specialists, the key message is that this dataset does not by itself change how your doctor will read your next scan—but it lays important groundwork. By freely sharing a large, carefully verified library of ultrasound images, the authors give researchers worldwide the raw material needed to train and test smarter tools for liver screening. In the future, such tools could act as a second pair of eyes, flagging worrisome spots that deserve closer attention and reducing unnecessary follow-up for clearly benign findings. Ultimately, this may help shift liver cancer diagnosis toward earlier, more treatable stages while keeping costs and complexity in check.
Citation: Tak, J., Ko, RE., Kwon, R.D. et al. SMC-LUD:Large-Scale B-Mode Liver Ultrasound Dataset for Hepatocellular Carcinoma and Hemangioma Classification. Sci Data 13, 649 (2026). https://doi.org/10.1038/s41597-026-07023-7
Keywords: liver ultrasound, hepatocellular carcinoma, hemangioma, deep learning, medical imaging dataset