Clear Sky Science · en
MIrROR release 02: Expanded and refined 16S-ITS-23S rRNA operon dataset
Why tiny microbes matter to us
Microbes shape our health, our environment, and even the climate, but identifying exactly which microscopic species are present in a soil sample, a river, or the human gut is surprisingly hard. This article introduces an upgraded reference dataset called MIrROR release 02, which helps scientists read long stretches of microbial DNA more precisely so they can tell closely related species apart and better understand how microbial communities work.
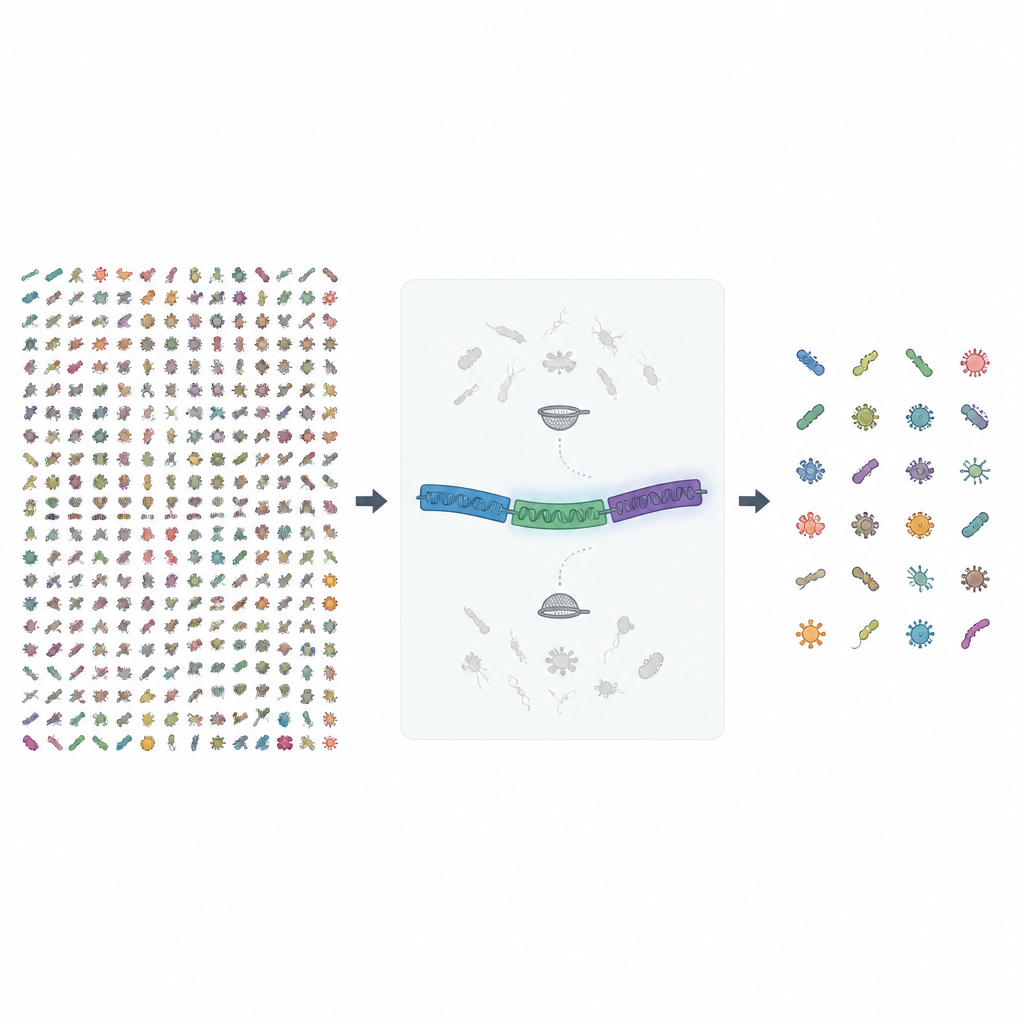
Looking beyond a single genetic landmark
For years, microbiologists have relied on short snippets of a single gene, known as 16S rRNA, to spot and count bacteria and archaea in a sample. That method is fast and cheap, but it often blurs the picture, treating different species as if they were the same. Even with newer long read sequencing machines that can read the full 16S gene, some species remain indistinguishable because this gene is too similar across close relatives. The MIrROR project tackles this by using a longer stretch of DNA that covers the full rRNA operon, including 16S, a spacer region, and another rRNA gene called 23S, giving many more sequence details to tell look alike microbes apart.
Building a bigger and cleaner reference map
In this new release, the authors gathered nearly 1.7 million bacterial and archaeal genomes from a public archive and searched them for complete rRNA operon sequences of reasonable length. They then put these raw sequences through several rounds of quality checks. Genomes lacking clear species names were dropped, exact duplicates across species were removed, and sequences with too many uncertain DNA letters were filtered out. Finally, highly similar sequences were clustered, and groups that mixed species were carefully inspected and cleaned, including manual checks with sequence comparison and evolutionary tree building tools to weed out contamination.
Adding neglected branches of the tree of life
A major advance in MIrROR release 02 is the inclusion of archaea, a broad group of microbes that thrive in environments ranging from hot springs to the human gut. The dataset now covers over a thousand archaeal species, among them medically and industrially important organisms. At the same time, the authors updated the names and groupings of many microbes using a modern genome based taxonomy. This reclassification affected about half of all genomes in the dataset and brought in nearly nineteen thousand additional bacterial species, including rare environmental microbes, clinically relevant pathogens, and species important in biotechnology and food production.
Making long read surveys work in real and test communities
To show that the expanded dataset is not just bigger but more useful, the team tested it on both laboratory made and computer simulated microbial mixtures. They compared MIrROR release 02 with earlier MIrROR data and with other common reference collections. In controlled tests, the new dataset was better at pinpointing species, including ones that older datasets missed entirely, such as a particular Prevotella species in a gut community standard. When archaeal species were added to a simulated gut community, the new MIrROR version could detect and classify them at both genus and species levels, while a widely used 16S only reference often produced vague labels like unexplained bacteria and struggled to assign reads to the correct species.
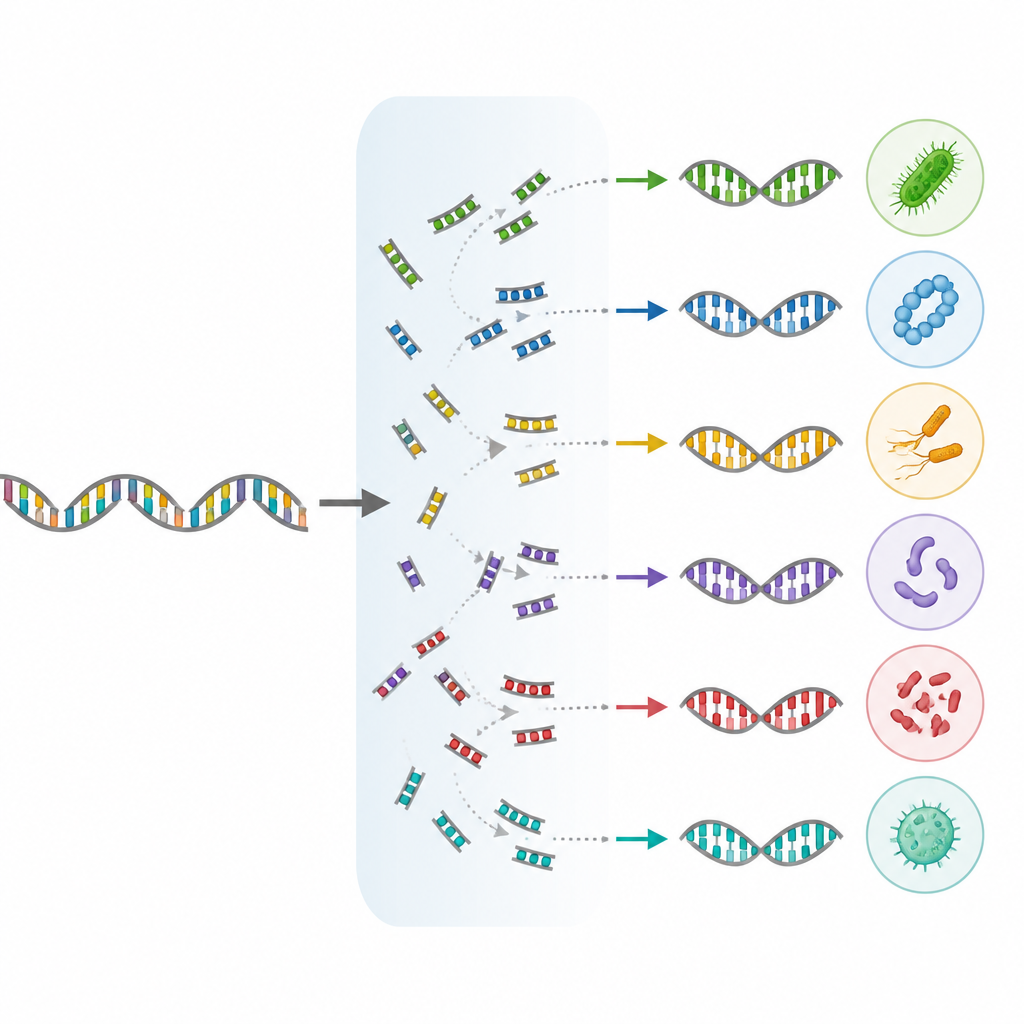
Helping scientists choose the right tools
Because long read sequencing relies on specific DNA starting points called primers, the authors also checked different primer pairs in computer simulations to see which could best capture both bacteria and archaea across the full operon. They recommend two primer sets that strike a balance between broad coverage and compatibility with long read platforms. At the same time, they point out known biological quirks, such as microbes that keep their rRNA genes unlinked or in multiple slightly different copies, which can bias counts and must be kept in mind when interpreting community data.
What this means for everyday questions
In simple terms, MIrROR release 02 is a much larger, better organized address book for microbes, built to work with modern long read DNA sequencing. It allows scientists to separate look alike species more reliably, to include archaea in their surveys, and to compare results across different studies with greater confidence. While it does not remove all challenges in reading microbial communities, it gives researchers a sharper lens for exploring how microbes influence human health, ecosystems, and industrial processes.
Citation: Lee, J., Hong, J., Seol, D. et al. MIrROR release 02: Expanded and refined 16S-ITS-23S rRNA operon dataset. Sci Data 13, 714 (2026). https://doi.org/10.1038/s41597-026-06729-y
Keywords: microbiome, rRNA operon, long read sequencing, microbial taxonomy, archaea