Clear Sky Science · en
A single-nucleotide enhancer mutation overrides chromosomal sex to drive XX male development
A tiny change with big consequences
Our biological sex is usually thought to be written in our chromosomes: XX for females and XY for males. This study reveals a surprising twist to that story. The researchers show that altering just a single “letter” in a stretch of DNA that regulates a key gene can override chromosomal sex in mice. With this minute tweak, animals carrying two X chromosomes develop male reproductive organs, even though they lack the Y chromosome traditionally required for testis formation.
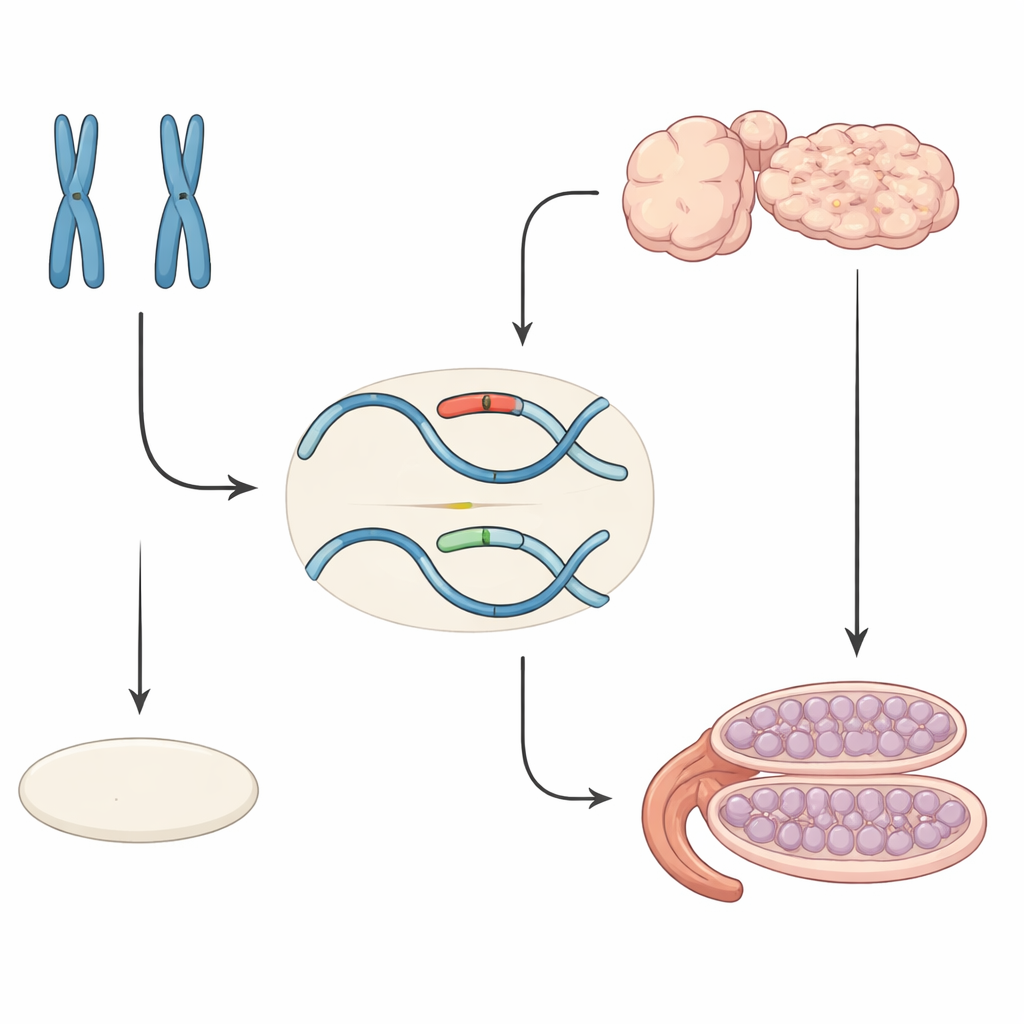
How sex is normally decided before birth
In mammals, sex is determined in several stages. At first, embryos possess undifferentiated gonads that can become either testes or ovaries. In XY embryos, a gene on the Y chromosome called Sry switches on another gene, Sox9, which drives the cells toward forming testes. In XX embryos, Sry is absent and a group of “pro-female” factors keep Sox9 off, allowing ovaries to develop. The level of Sox9 acts like a molecular threshold: if it rises high enough at the right time, testis development is launched; if it stays low, ovarian development proceeds.
The hidden switch far from the gene
The team previously identified a short DNA element, called Enh13, lying more than half a million DNA letters away from the Sox9 gene. Although distant, Enh13 works as a powerful on-switch for Sox9 in developing testes. Deleting Enh13 in mice or humans reduces Sox9 activity so sharply that XY individuals can develop ovaries instead of testes. Intriguingly, some XX people with differences of sex development carry small duplications that include the human version of Enh13, hinting that extra copies might inappropriately activate Sox9 and push XX gonads toward a testis fate.
Single-letter edits that flip XX gonads to testes
In this study, the researchers engineered extremely subtle changes inside Enh13 in mice: either a three-letter deletion or a one-letter insertion within a short sequence where the SOX9 protein can bind. XX mice that inherited two copies of these altered enhancers developed as males. As adults, they looked male externally and internally, with testes instead of ovaries, although the testes were small and infertile because they lacked Y-linked genes needed for sperm production. When the scientists examined embryos, they found that XX gonads initially formed a patchwork of ovarian and testicular tissue—an “ovotestis”—before ultimately resolving into testes as development progressed.
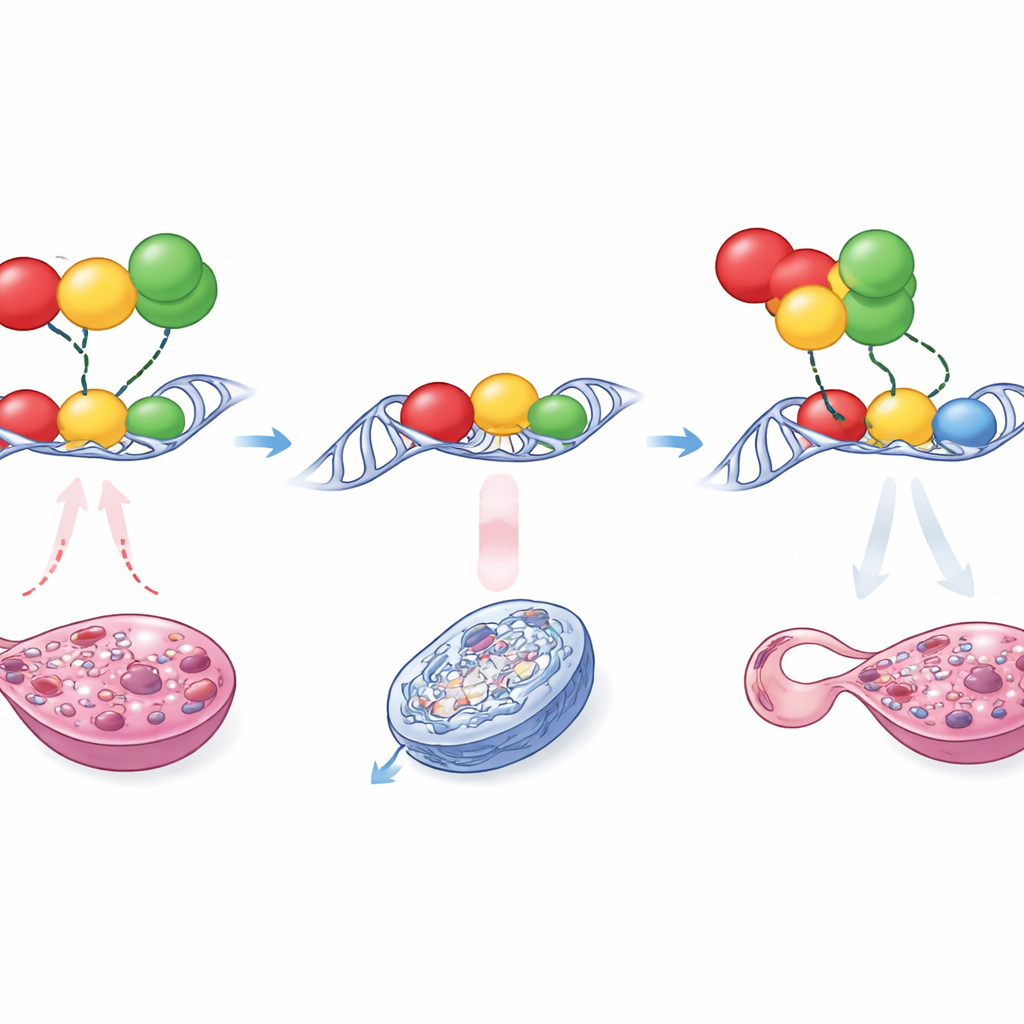
A tug-of-war on a tiny DNA strip
How can such a delicate change cause such a dramatic outcome? Detailed molecular tests showed that the mutations did not simply make SOX9 bind more strongly. Instead, Enh13 acts as a crowded docking site for several proteins that either push toward a testis fate or hold the system in an ovarian state. Among these are RUNX1, NR5A1, and GATA4, factors active in the early gonad. In the normal enhancer, the arrangement and spacing of their binding spots allow “pro-female” influences, especially RUNX1 in partnership with others, to dampen Enh13’s activity in XX embryos and keep Sox9 below the critical threshold. The small insertion or deletion subtly changes the spacing and local structure of this binding cluster. As a result, RUNX1 can no longer impose the same level of repression, and in one mutant a new site appears that lets GATA4 bind in a more activating configuration. These structural shifts let the enhancer become overactive even without Sry, nudging Sox9 expression just high enough that SOX9 begins to boost its own production, locking the gonad into the testis pathway.
Why this matters for sex development and disease
This work shows that Enh13 is not just an “on” switch for testes but also a key site where ovarian factors normally silence Sox9. In other words, the same tiny DNA element can either start or stop testis development depending on which proteins are bound. The study demonstrates that even a one-letter change in non-coding DNA—regions that do not encode proteins—can completely flip an animal’s sexual fate. This has important implications for understanding unexplained cases of human sex development differences and illustrates a broader principle: the three-dimensional arrangement of protein docking sites within enhancers can be just as crucial as the presence of the sites themselves. Slight shifts in spacing can rewire how regulatory proteins cooperate, turning a balanced genetic switch into a powerful driver of developmental change.
Citation: Abberbock, E., Ridnik, M., Stévant, I. et al. A single-nucleotide enhancer mutation overrides chromosomal sex to drive XX male development. Nat Commun 17, 3186 (2026). https://doi.org/10.1038/s41467-026-71328-9
Keywords: sex determination, Sox9, enhancers, gonadal development, XX male sex reversal