Clear Sky Science · en
Identification of hot spring Obelisk-like RNA replicons and expanded diversity of the Obelisk superfamily
Hidden circles in extreme hot springs
Near-boiling, acidic hot springs seem like places where life should fall apart. Yet these harsh pools are teeming with microbes – and, as this study shows, with tiny ring-shaped molecules of RNA that quietly copy themselves. The researchers set out to see whether a mysterious class of RNA circles, known as Obelisks, also lives in such extreme settings. In doing so, they uncovered new members of this strange family and showed that these microscopic “circles of code” are far more common and varied in Earth’s ecosystems than anyone realized.
How tiny RNA rings differ from familiar viruses
Most people know viruses as particles carrying DNA or RNA that encode several proteins. Viroids and related agents are different. They are short loops of RNA that often fold into rod-like shapes and may not encode any protein at all. They hijack the machinery of their host cell to make copies, sometimes harming crops but otherwise remaining obscure. In recent years, high-throughput sequencing of environmental RNA has revealed a zoo of such circular molecules in places with few plants or animals, hinting that microbes can also host these minimal genetic elements. Among them are Obelisks, circular RNAs about one thousand building blocks long that do encode a single protein with a previously unknown three-dimensional shape.
Finding new RNA circles by reading double-stranded intermediates
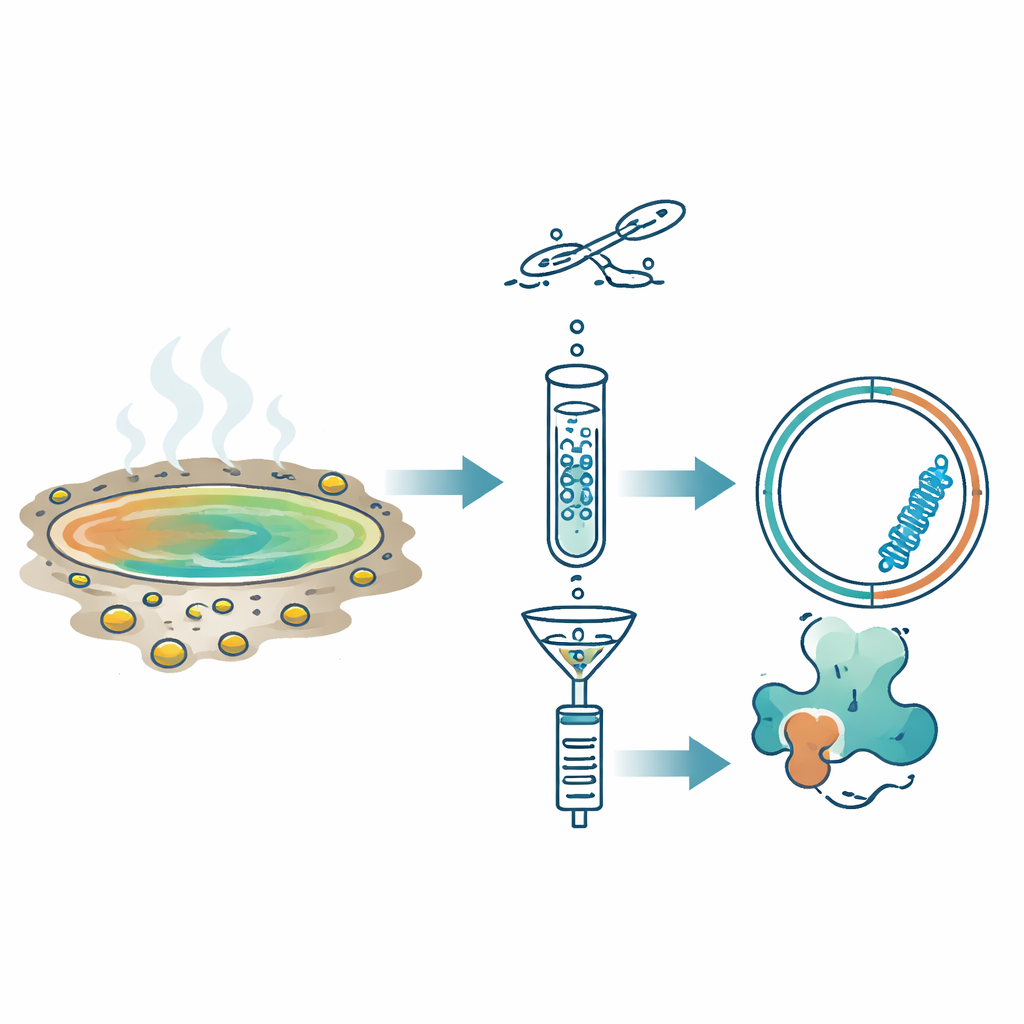
Instead of searching directly for known sequences, the team used a different hallmark of RNA replicons: during copying, they briefly form double-stranded RNA. They applied a method called Fragmented and primer-Ligated DsRNA Sequencing (FLDS), which enriches these double-stranded forms from environmental samples. After purifying double-stranded RNA from eleven acidic hot springs in Japan, they sequenced it and used a stepwise filtering pipeline. Only molecules that appeared circular, were well covered by sequencing reads, and folded into stable, rod-like structures were kept. To validate the approach, they first confirmed that it could rediscover a known plant viroid in infected chrysanthemum leaves. Then, applying the same logic to hot spring samples, they isolated a standout candidate they named Hot spring Obelisk (HsOb).
New Obelisks thriving in near-boiling acid pools
The key RNA discovered in one spring called Oi was about 870 nucleotides long and formed an extended rod-like structure even when modeled at 80 °C, the water temperature at the site. It contained a single long open reading frame that would produce a protein of 213 amino acids, dubbed HsOblin-1. Although its amino acid sequence showed almost no similarity to known proteins, structural prediction tools revealed that HsOblin-1 folds into the same basic architecture as the original Obelisk protein, Oblin-1: a compact, α-helix-rich core followed by a flexible, positively charged tail likely suited for gripping RNA. A related RNA from another spring, H5, encoded a similar protein. The microbial communities in these samples were dominated by thermoacidophilic bacteria of the family Hydrogenobaculaceae, making these bacteria the leading candidates as hosts for the hot spring Obelisks, even though direct host assignment remains to be proven.
A global survey shows a vast and varied family
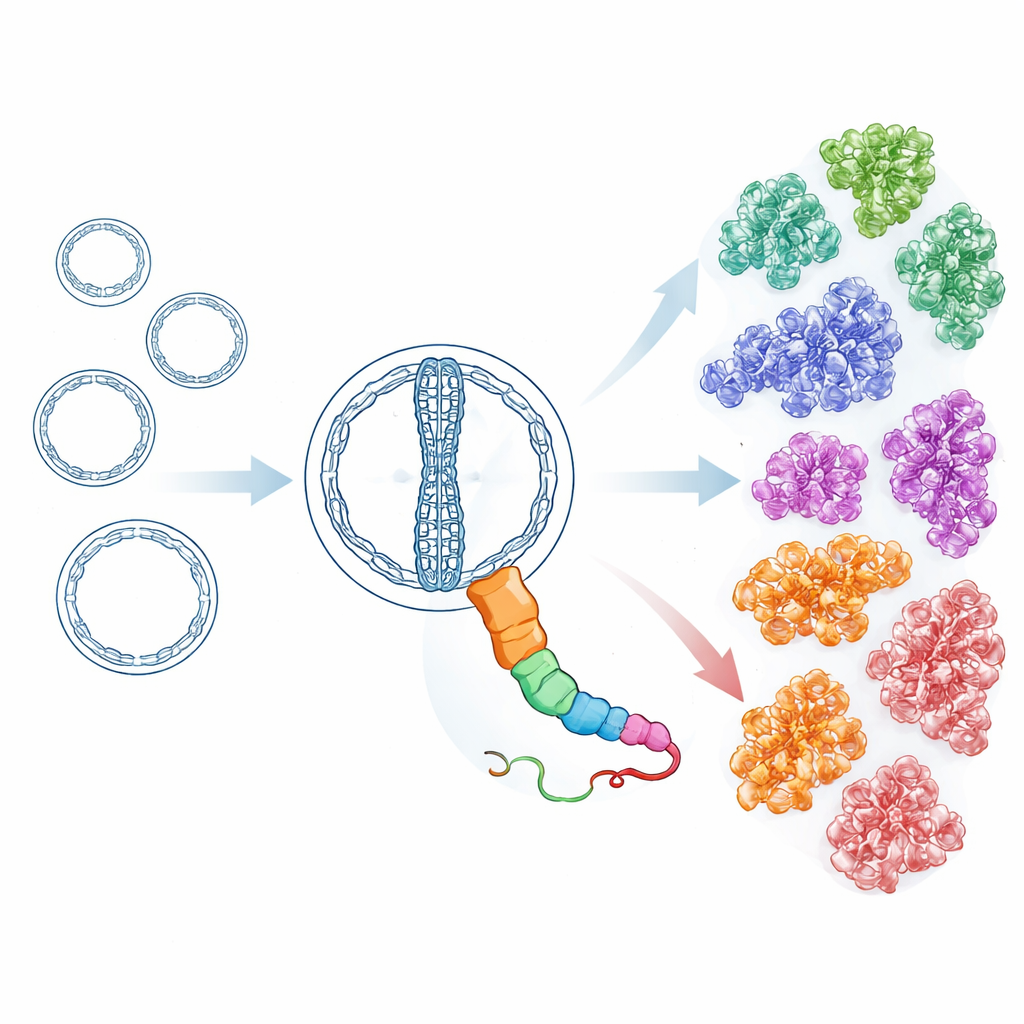
Armed with sensitive structure-aware search tools, the researchers then scanned millions of putative circular RNAs from over 7,000 environmental metatranscriptomes, along with additional datasets from public repositories. Starting from the hot spring HsOblin-1 proteins and the originally described Oblin-1, they iteratively built profiles that capture distant relatives even when sequence similarity is weak. This effort uncovered 6,443 distinct proteins with an Oblin-like fold, encoded by more than 8,000 Obelisk RNA genomes. Grouping these proteins by similarity and by predicted three-dimensional structure revealed 21 main subfamilies. All share the same core fold but differ in surface features and in parts of a long, flexible loop, suggesting that they have diversified to interact with different hosts or partners while keeping the basic replication toolkit intact. Some subfamilies also appear to carry extra small proteins with simple helical shapes, hinting at add-on functions that have evolved independently.
Tracing hosts and lifestyles across the tree of life
To understand who these RNA circles live with, the team searched for matches between Obelisk sequences and CRISPR spacers – short genetic records that bacteria and archaea keep of past infections. They found dozens of convincing matches, especially in bacteria that inhabit the guts of humans and ruminants, strengthening the idea that Obelisks commonly infect or coexist with bacteria. Combined with earlier work that found Obelisks in the human mouth and in marine waters, these results suggest that Obelisks are not rare curiosities, but widespread members of microbial communities from ocean to hot springs to animal microbiomes. Yet many puzzles remain: it is not clear whether Obelisks behave more like plasmids (extra genetic elements that stay within a cell line) or like infectious agents that spread between hosts, and the details of how they copy themselves are still unknown.
What this means for our picture of life’s diversity
To a non-specialist, these findings show that even in places as inhospitable as near-boiling acid pools, evolution has filled the microscopic world with unconventional genetic entities. Hot spring Obelisks prove that circular RNA replicons with a shared protein fold can thrive at high temperatures and in very different habitats. By almost doubling the known diversity of Obelisks and mapping their likely bacterial partners, this study reveals a vast, previously unseen layer of genetic life. It suggests that simple RNA-based replicons are not just historical relics, but active, adaptable players in today’s biosphere, quietly shaping microbial ecosystems across the planet.
Citation: Urayama, Si., Fukudome, A., Mutz, P. et al. Identification of hot spring Obelisk-like RNA replicons and expanded diversity of the Obelisk superfamily. Nat Commun 17, 3041 (2026). https://doi.org/10.1038/s41467-026-71096-6
Keywords: circular RNA, hot springs, microbial ecosystems, RNA replicons, Obelisk elements