Clear Sky Science · en
Synthetic data-driven deep learning for label-free autonomous atomic force microscopy
Seeing the Tiny World Without Human Eyes
Our ability to engineer new materials, study clean energy devices, or probe living cells often depends on seeing structures that are a thousand times smaller than a human hair. Atomic force microscopes (AFMs) can map these tiny landscapes in 3D, but today they still rely heavily on expert operators who must choose where to look and how to interpret the images. This paper introduces SimuScan, a way to teach computers to run AFMs and recognize nanoscale features by training them on lifelike simulated images instead of painstakingly labeled experimental data.
Why Today’s Nanoscale Imaging Gets Stuck
AFMs have become central tools in materials science, energy research, and biology because they can feel surfaces with nanometer precision. Yet they are slow, cover only small areas at a time, and demand many decisions from a skilled user: where to scan, what settings to use, and which tiny shapes matter. Unlike optical or electron microscopes that can capture large survey images, AFMs build up surfaces line by line. On top of that, modern artificial intelligence methods thrive on enormous labeled image collections like those available for everyday photos, but such datasets simply do not exist for AFM. Each AFM image is influenced by subtle instrument quirks—noise, distortions, and probe shape—so generic computer vision tools trained on ordinary pictures often fail when applied directly.
Teaching a Microscope With Make-Believe Images
The authors’ key idea is to generate vast libraries of synthetic AFM images that look and behave like the real thing, complete with all the usual imperfections. Their SimuScan framework starts by defining the shapes that might appear on a sample: simple blocks and rods, delicate DNA assemblies, or whole bacterial cells. These shapes can come from mathematical descriptions, computer-aided design files, or even 3D surfaces extracted from a few real AFM images. SimuScan then places these objects on simulated substrates that can include steps, roughness, periodic patterns, and random debris, creating realistic, pre-distortion landscapes. Finally, it passes these surfaces through a detailed forward model of the microscope that adds the effects of a finite probe tip, feedback glitches, line-by-line corrections, and electronic noise. The result is an image that closely mimics what an AFM would measure, paired with an exact “ground truth” map of every object’s outline.
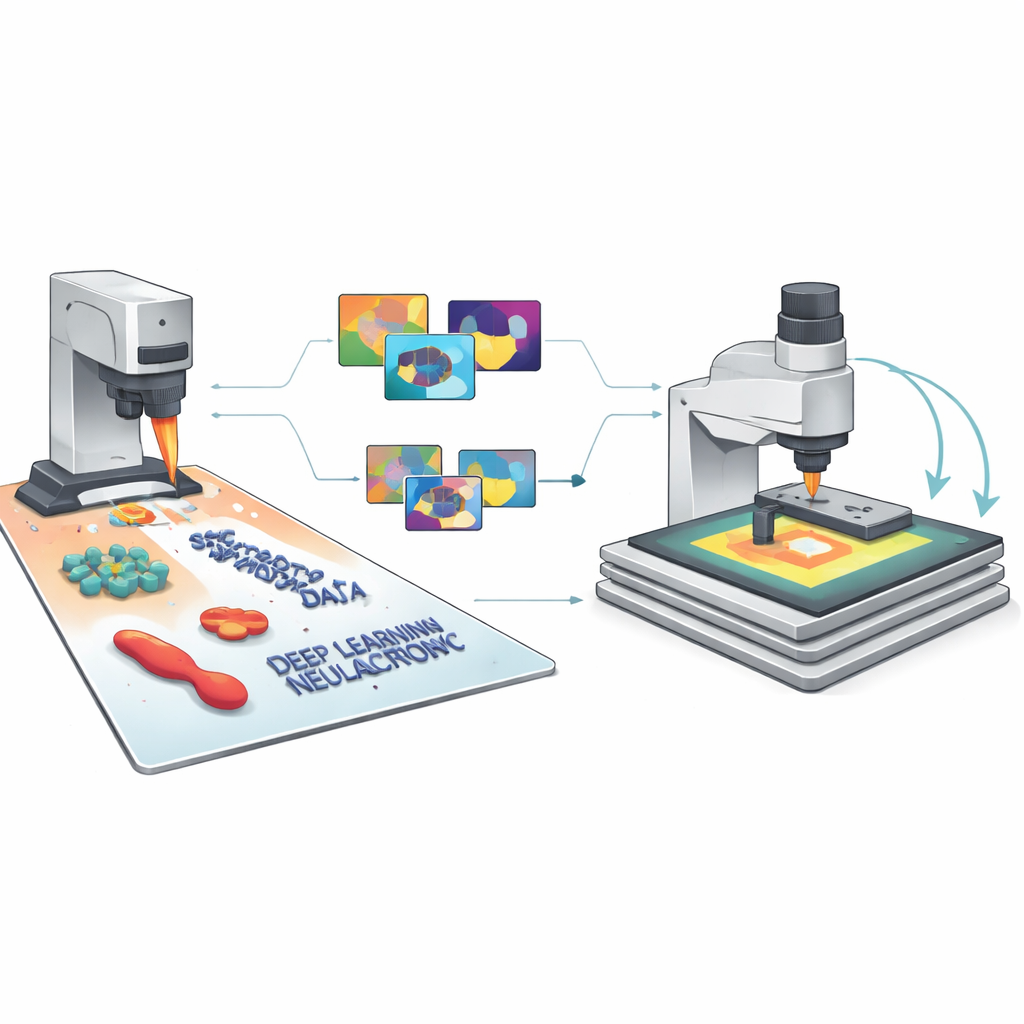
From Simulated Pixels to Reliable AI
Because each synthetic image comes with perfect labels for every feature and pixel, SimuScan can feed modern deep learning models with the kind of rich training material they normally lack in nanoscale imaging. The team tested several popular architectures—YOLOv8 for fast object detection, U-Net for detailed outlines, and Mask R-CNN for instance-level masks—using more than 5,000 synthetic images per task. Remarkably, models trained only on these artificial datasets performed strongly when evaluated on real AFM images of fabricated nanostructures and bacteria that had been painstakingly annotated by experts. Detection scores and segmentation accuracy were high across different shapes and cell types, showing that the simulated images had captured the essential appearance of these tiny structures and their common imaging artifacts.
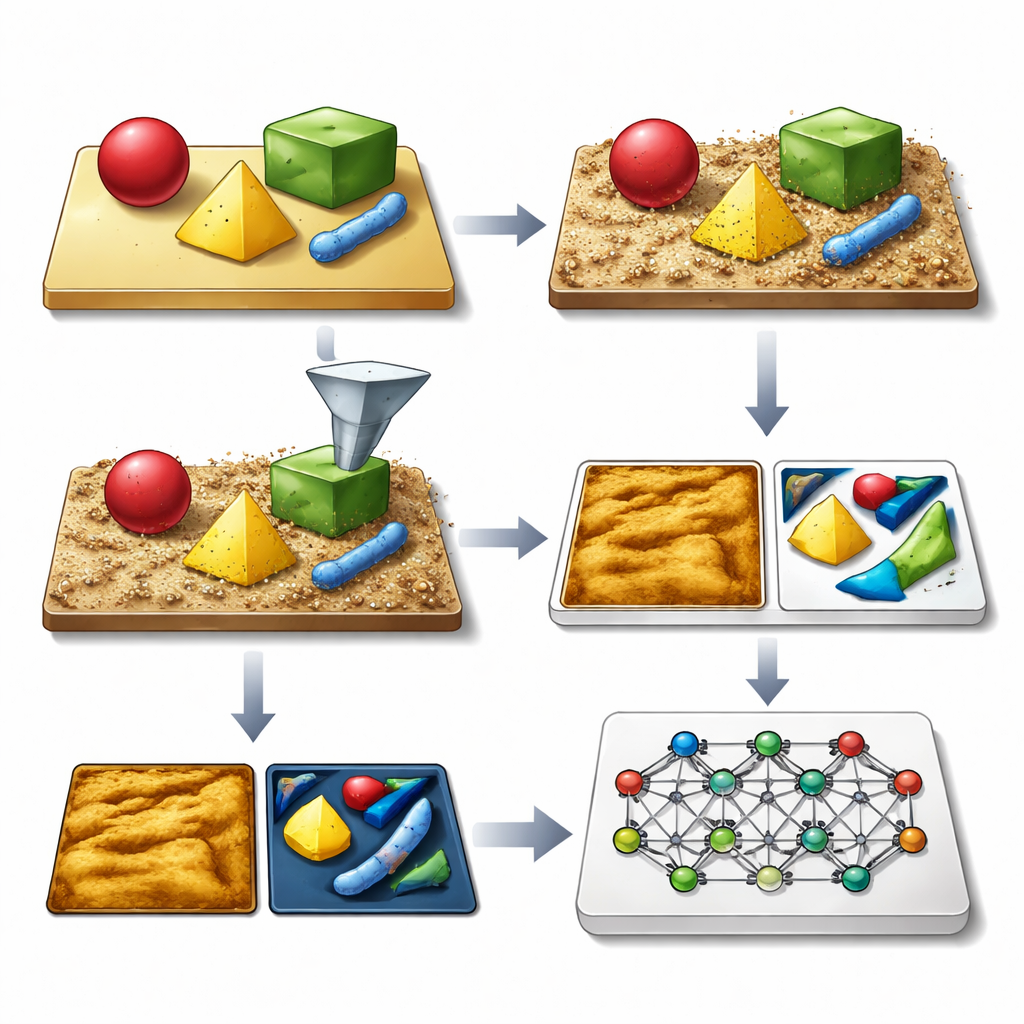
Letting the Microscope Decide Where to Look
The researchers then closed the loop between simulation, AI, and the physical instrument. In their semi-autonomous workflow, the AFM first takes a low-resolution overview scan over a relatively large area. A model trained on SimuScan data analyzes this image in real time, finds structures of interest—such as certain bacterial shapes or specific nanofabricated patterns—and selects promising regions for zoomed-in, high-resolution scans. The microscope automatically moves, rescans, and repeats this cycle across the sample, guided by simple user-defined rules like the desired number of objects or total area to cover. Using this approach, the system could autonomously find and image hundreds of individual bacteria, then measure variations in their size and shape across a population, something that would be prohibitively time-consuming to do by hand.
A New Path to Smarter Microscopes
To a non-specialist, the main takeaway is that SimuScan shows how “imaginary but realistic” data can help microscopes become more like self-driving instruments. By simulating both the tiny objects on a surface and the quirks of how the AFM sees them, the authors remove the need for large, manually labeled training sets and allow general-purpose AI models to work well on real experiments. This unlocks AFM studies that can explore larger areas, analyze many more objects, and adapt their behavior on the fly, making nanoscale characterization faster, more reproducible, and accessible to non-experts. In the long run, similar synthetic-data strategies could help bring autonomous, discovery-driven operation to many other types of scientific instruments.
Citation: Millan-Solsona, R., Checa, M., Brown, S.R. et al. Synthetic data-driven deep learning for label-free autonomous atomic force microscopy. Nat Commun 17, 3886 (2026). https://doi.org/10.1038/s41467-026-70421-3
Keywords: atomic force microscopy, synthetic data, deep learning, autonomous microscopy, nanostructure imaging