Clear Sky Science · en
Functional protein design and enhancement with ontology reinforcement iteration
Teaching Computers to Build Better Proteins
Proteins do nearly all the work inside living cells, from breaking down food to fighting infections. Scientists would love to design new proteins on demand, such as tougher industrial enzymes or better medical tools, but turning computer ideas into real, working molecules is still hard. This study introduces a new system called ORI that links artificial intelligence directly to lab experiments, with the goal of making protein design faster, more reliable, and easier to control.
Why Designing Proteins Is So Difficult
Each protein is built from a long chain of amino acids, and tiny changes in this chain can dramatically alter how the molecule folds and behaves. Modern AI can predict protein shapes and even generate new sequences, but there is a stubborn gap between what looks good in a computer and what actually works in a test tube. Many designs fail because they are unstable, do not fold properly, or lack the desired activity, forcing researchers to rely on slow trial and error.
A Closed Loop Between Computer and Lab
The ORI framework tackles this problem by creating a feedback loop between AI models and real experiments 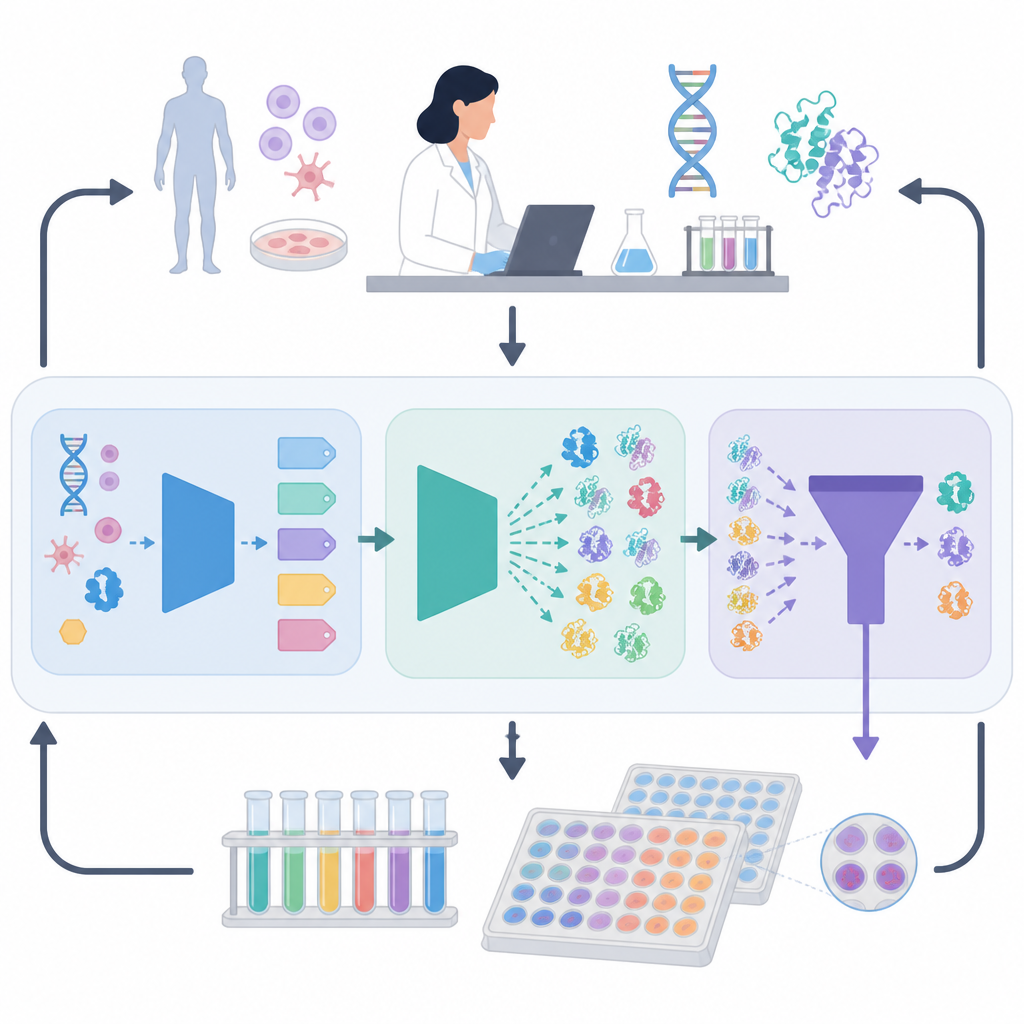
Learning from Experimental Feedback
What makes ORI different is that it does not stop at prediction; it learns from the lab’s successes and failures. Selected protein designs are synthesized and tested for how well they express, how active they are, and how they behave under stress. These results are then fed back into the AI using a strategy adapted from reinforcement learning, allowing the system to gradually favor sequence patterns that perform well in real life rather than only on paper. Over multiple rounds, the AI becomes better at proposing candidates that survive this real-world gauntlet 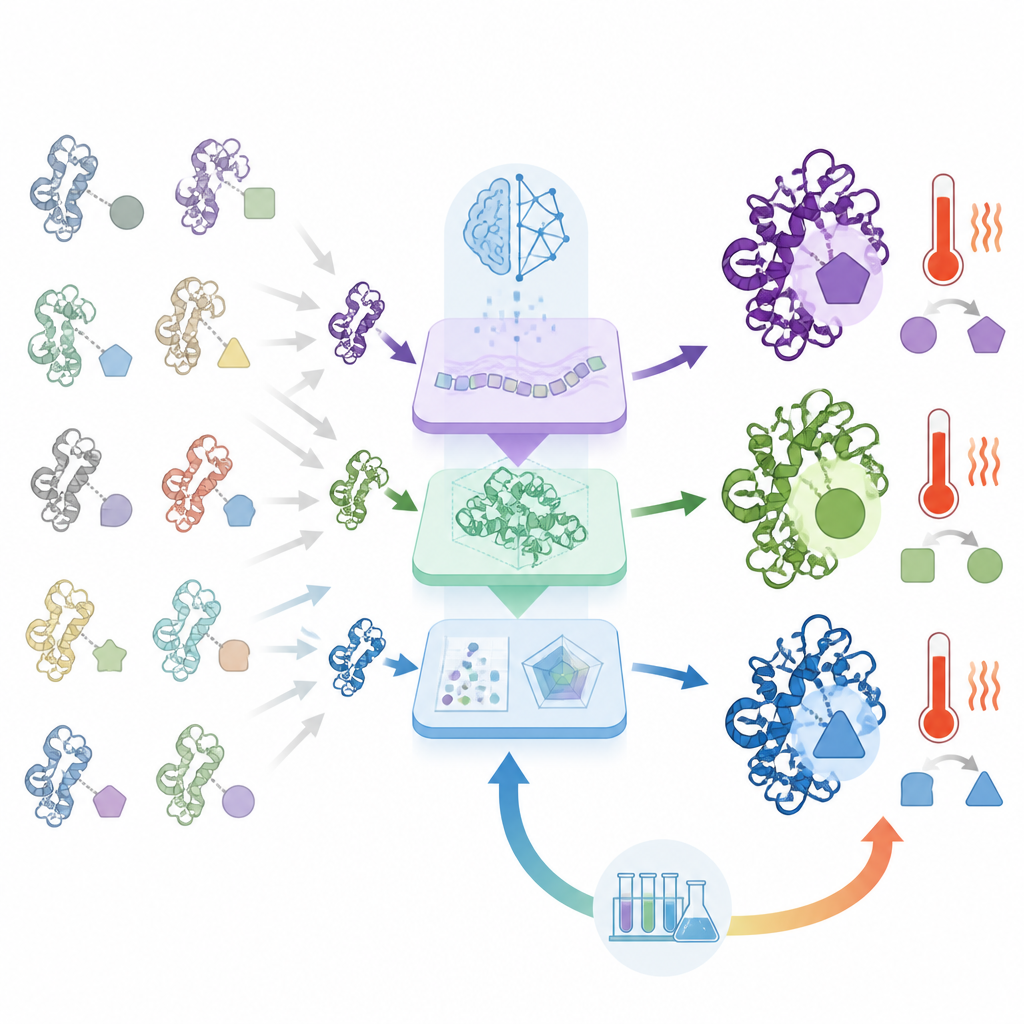
Building Stronger, Hotter, and Dual-Function Enzymes
To show what ORI can do, the authors applied it to several enzyme design challenges. For lysozymes, which break down bacterial cell walls, the system generated many variants and, after learning from experimental feedback, produced versions with much higher activity than natural lysozymes, including one that was roughly one hundred times more active than a standard reference. ORI was also used to design chitinases, enzymes that digest chitin found in insect shells and fungi. By steering the model toward heat-resistant features, the team created chitinases that stayed active at temperatures around 85 degrees Celsius, far beyond what most natural versions can tolerate. Finally, the system engineered single proteins that combine both lysozyme and chitinase activities, and some of these dual-function enzymes outperformed their specialized natural counterparts in both tasks.
A New Way to Explore Protein Possibilities
For non-specialists, the key idea is that ORI turns protein design into an ongoing conversation between computer models and the wet lab, rather than a one-shot prediction. The system uses structured knowledge about protein types and properties to guide its imagination, and it listens carefully to what experiments say about each new design. By looping through this process, ORI can not only match natural proteins but sometimes surpass them in strength, heat resistance, or versatility. This suggests a future in which tailored proteins for medicine, industry, and environmental uses can be developed more efficiently and with greater confidence.
Citation: He, B., Qin, C., Zhao, Y. et al. Functional protein design and enhancement with ontology reinforcement iteration. Nat Commun 17, 4158 (2026). https://doi.org/10.1038/s41467-026-69855-6
Keywords: protein engineering, enzyme design, artificial intelligence, thermostable enzymes, multifunctional proteins