Clear Sky Science · en
Systematic benchmarking demonstrates large language models have not reached the diagnostic accuracy of traditional rare-disease decision support tools
Why this matters for patients and doctors
When someone has a rare disease, getting the right diagnosis can take years of uncertainty, repeated tests, and visits to many specialists. At the same time, powerful new artificial intelligence systems called large language models are being hailed as potential game‑changers for medicine. This study asks a simple but crucial question: are these new AI chatbots actually better at spotting rare genetic diseases than the specialized tools doctors already use, or do they still have a way to go?
Rare diseases and the long road to answers
Although each rare disease affects relatively few people, there are more than 10,000 such conditions, and together they touch up to one in twelve individuals. Many of these patients endure a “diagnostic odyssey” lasting five to seven years before anyone can name what is wrong. For genetic conditions, one key step is to match a person’s combination of symptoms, lab results, and scan findings to the known patterns of specific diseases. Established computer programs already help with this by using standardized vocabularies of medical features to search through thousands of possible conditions.
Putting chatbots and traditional tools to the test
The researchers assembled a large collection of more than five thousand real but anonymized cases of patients with confirmed rare genetic or chromosomal diseases. Each case had been carefully converted into a structured format that encodes the person’s age, sex, symptoms, and test findings using a shared medical dictionary. From these structured records, the team automatically generated short case stories that could be given to seven different language models, including general‑purpose systems and models trained especially on medical text. In parallel, they fed the same structured data into Exomiser, a widely used rare‑disease decision‑support program, but without giving it any genetic sequencing information so the comparison would be fair.
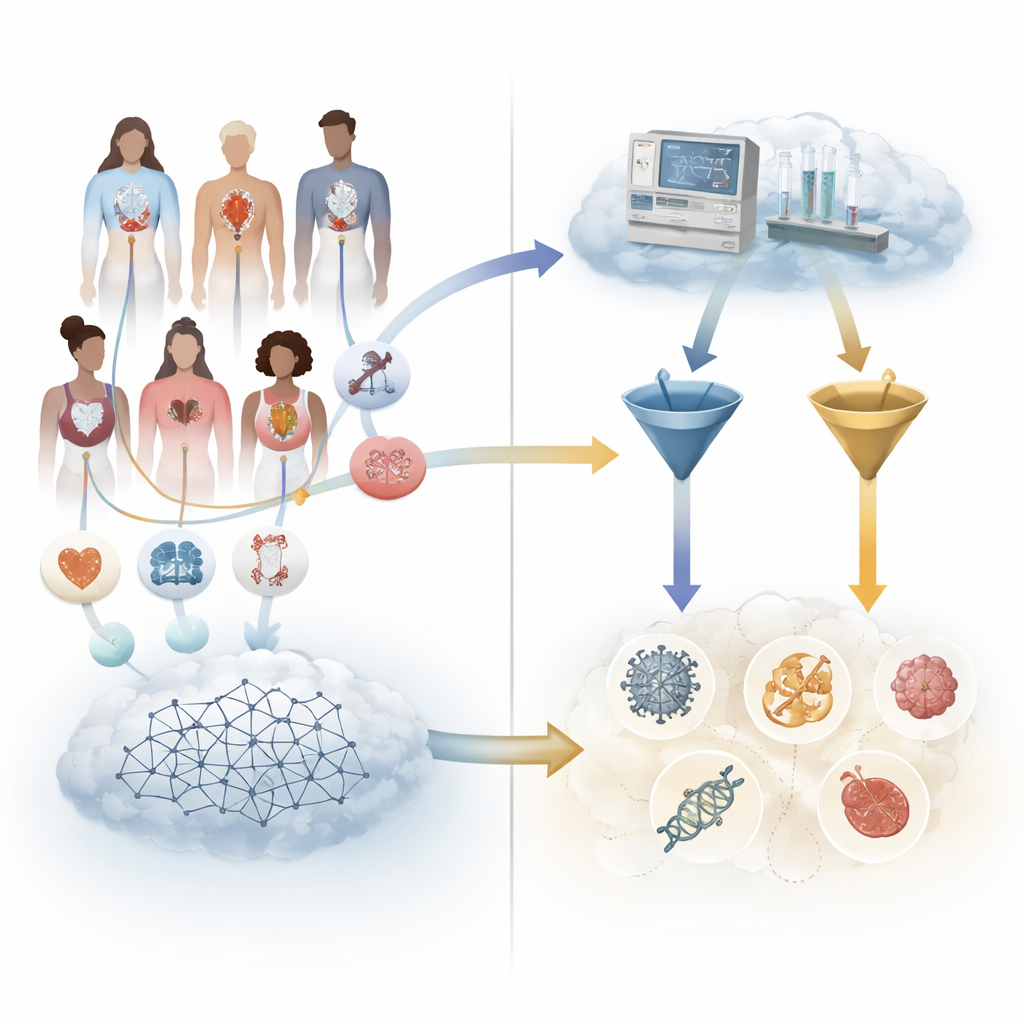
Scoring who found the right disease
Comparing the answers from chatbots and traditional software is harder than it sounds, because language models respond in free‑form text that may use different disease names or levels of detail. To avoid relying on human opinion about whether an answer was “close enough,” the team mapped every suggested diagnosis onto a unified disease catalogue. A model’s suggestion counted as correct if it matched the exact disease, an equivalent name, or a slightly more general version that clearly included the true condition. For each case, they then measured where the correct answer appeared in the model’s ranked list—first place, among the top three, or somewhere in the top ten.
What the head‑to‑head comparison showed
Across all 5,213 cases, Exomiser clearly outperformed every language model tested. Using only symptom information, Exomiser placed the correct diagnosis first in about one out of three cases and within the top ten in well over half. The best language model, a reasoning‑focused system, reached first place in just under one quarter of cases and the top ten in a bit more than one third. Other medical‑focused models did noticeably worse, and one very large medical model almost never suggested the right disease. These patterns held up even when the researchers looked separately at heart, brain, or immune‑related disorders, and when they split cases by how detailed the symptom descriptions were.
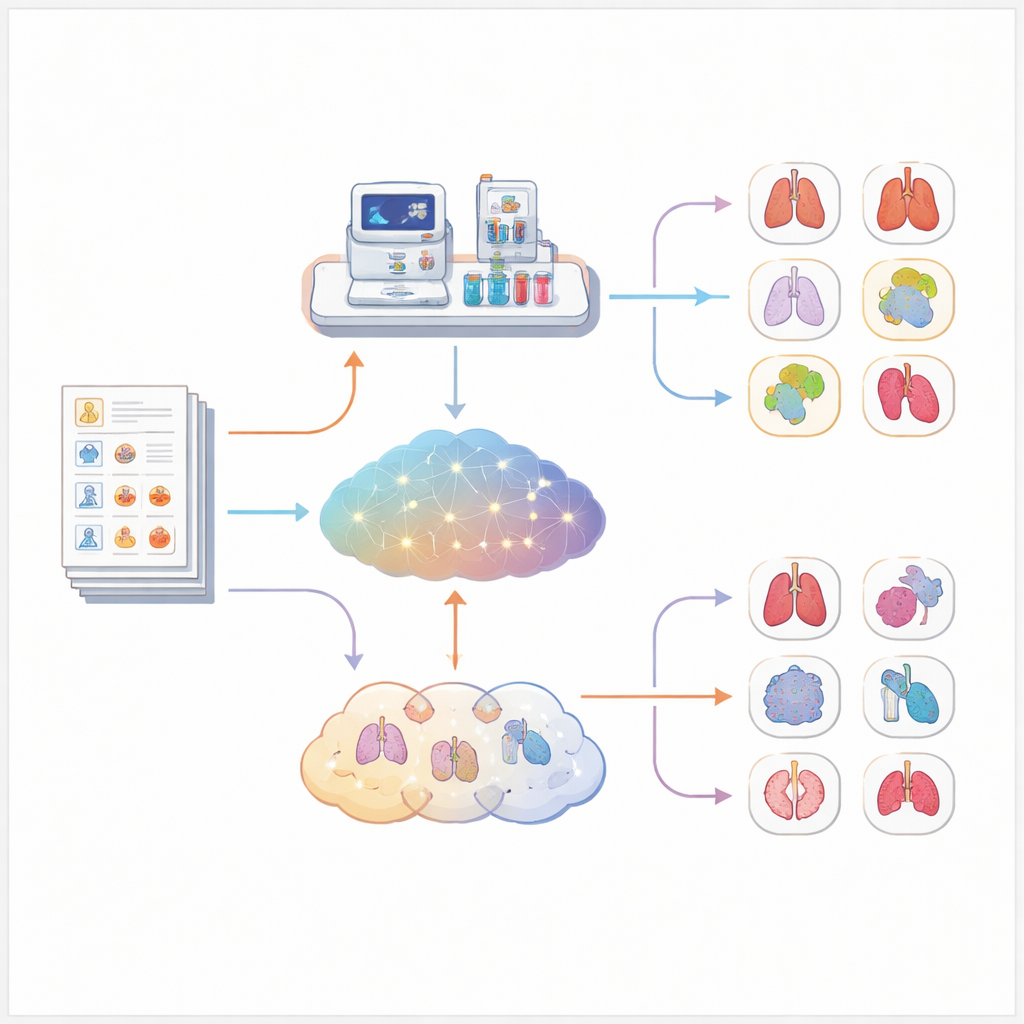
What this means for the future of AI in diagnosis
This study shows that, despite their impressive abilities with language, today’s large language models are not yet as reliable as specialized tools for diagnosing rare genetic diseases from symptom descriptions alone. They can still be useful as assistants—for example, helping doctors think of possibilities or explain conditions in plain language—but they should not replace established software when lives and long‑delayed answers are at stake. The authors argue that the most promising path forward is to weave language models into carefully designed diagnostic pipelines, where they work alongside, rather than instead of, trusted bioinformatics tools.
Citation: Reese, J.T., Chimirri, L., Bridges, Y. et al. Systematic benchmarking demonstrates large language models have not reached the diagnostic accuracy of traditional rare-disease decision support tools. Eur J Hum Genet 34, 498–504 (2026). https://doi.org/10.1038/s41431-026-02054-5
Keywords: rare diseases, medical diagnosis, large language models, clinical decision support, genetic disorders