Clear Sky Science · en
X-linked cancer-associated polypeptide (XCP) from lncRNA1456 modulates PHF8 histone demethylase activity to regulate the epigenome, gene expression, and cellular pathways in breast cancer
Hidden Messages in the “Junk” Genome
For years, much of our DNA was dismissed as “junk” because it did not obviously encode proteins. This study shows that some of that overlooked genetic material can in fact produce powerful, previously unknown proteins that shape how cancers grow. By uncovering one such protein in breast tumors, the researchers reveal a new layer of gene control and point to fresh possibilities for diagnosis and treatment.
A Small Protein Hiding in Plain Sight
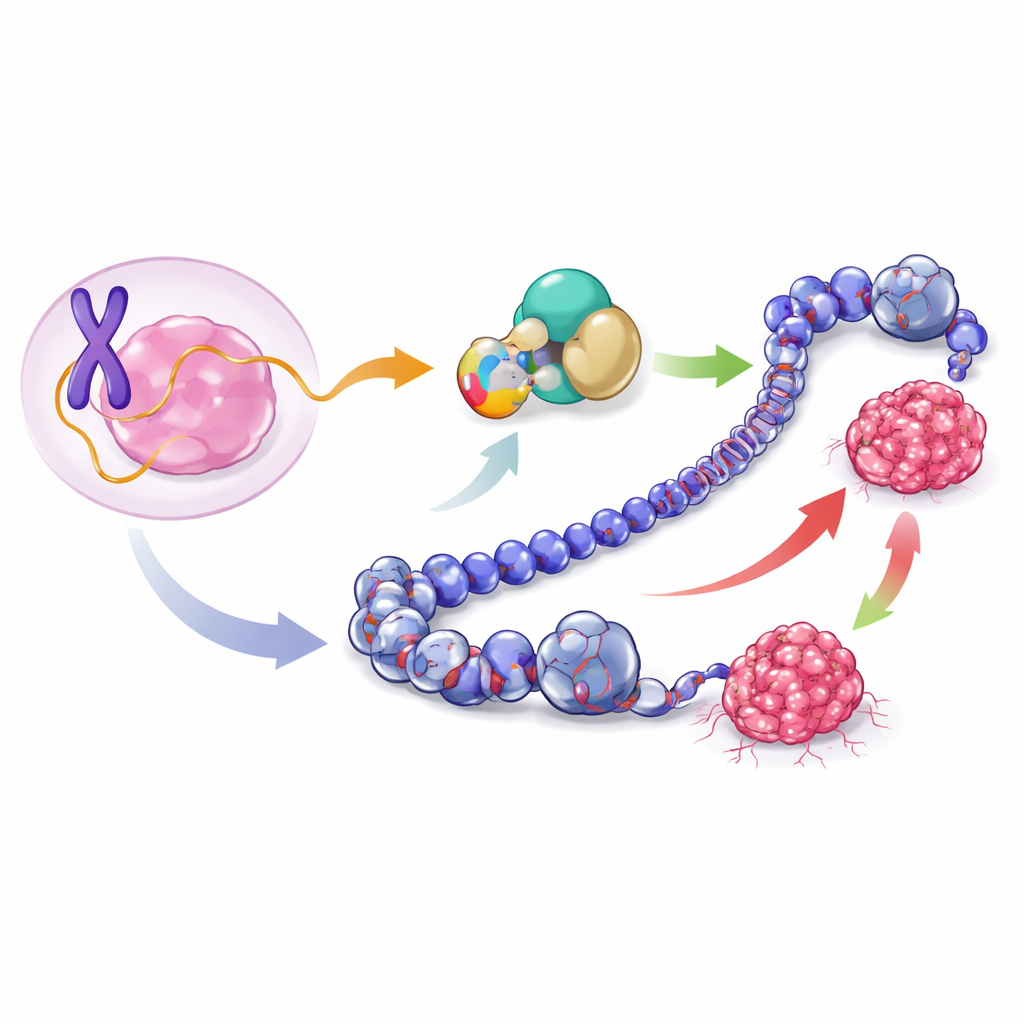
The team focused on a stretch of DNA on the X chromosome long assumed to make only a noncoding RNA molecule, called lncRNA1456. Using a combination of RNA sequencing and protein-detection methods, they discovered that this RNA actually carries instructions for a short protein of 132 building blocks, which they named XCP (X-linked Cancer-associated Polypeptide). XCP is produced from a region spanning the first two segments, or exons, of the RNA. In breast cancer cells of the luminal type—a common, typically hormone-sensitive subtype—XCP is made in abundance and is found mainly in the cell nucleus, where the DNA is housed and gene activity is controlled.
Where XCP Appears in the Body and in Tumors
To check whether XCP was just a lab artifact, the researchers surveyed many normal human tissues. They found that lncRNA1456 and its protein XCP are naturally active almost only in the testis and pancreas. In contrast, in breast cancer samples from large patient datasets, lncRNA1456 and XCP are strongly switched on in tumors but barely detectable in healthy breast tissue. Within breast cancers, high XCP levels show up most often in less aggressive, hormone receptor–positive tumors (luminal A, luminal B, and HER2-enriched types) and are rare in more aggressive basal-type tumors. This pattern suggests that cancer cells can improperly activate an otherwise tightly restricted X-linked gene, turning an unusual testis-associated protein into a cancer-associated one.
How XCP Steers Tumor Growth
The authors next asked what XCP actually does to cancer cells. They engineered breast cancer cell lines to make extra XCP on command and implanted these cells into mice to form tumors. In luminal MCF-7 cells, extra XCP sped up tumor growth, implying that XCP can act like an oncogene in this setting. Surprisingly, when the same protein was forced into basal-type MDA-MB-231 cells, tumors grew more slowly, hinting at a tumor-suppressive role in that context. By analyzing global gene activity in the resulting tumors, the team saw that XCP turns distinct sets of genes and pathways on and off in each cell type: in luminal cells it boosts cell-cycle and growth programs, whereas in basal cells it dampens pathways linked to invasion and tissue remodeling. Thus a single small protein can push cancer in opposite directions depending on the cellular environment.
A Molecular Partner That Rewrites the Epigenetic Code
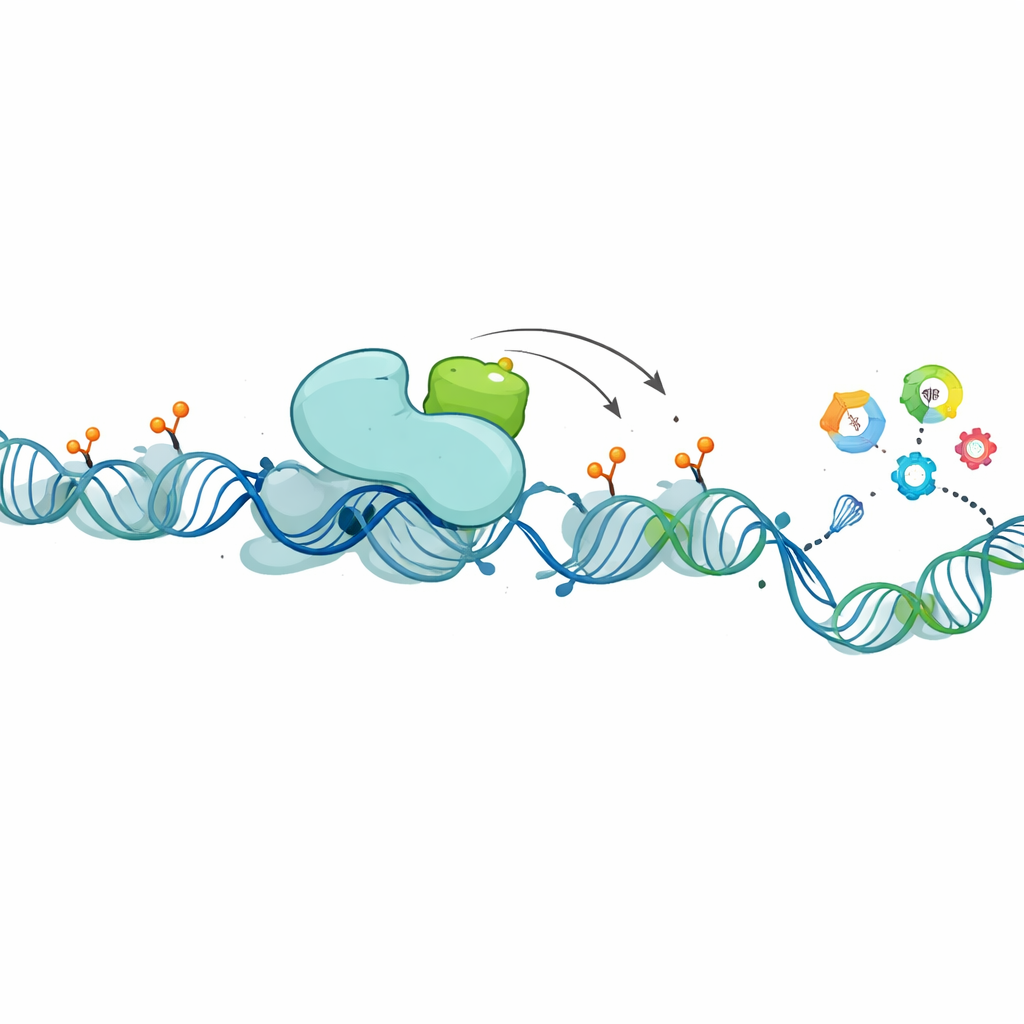
Digging deeper, the researchers searched for proteins that physically associate with XCP in the nucleus. They zeroed in on PHF8, a known “eraser” of certain chemical tags on histone proteins—the spools around which DNA is wrapped. These tags help determine whether nearby genes are active or silent. PHF8, like XCP, is encoded on the X chromosome, is highly expressed in testis, and is elevated in luminal breast cancers. Experiments showed that XCP and PHF8 bind to each other and are often present in the same tissues and tumor subtypes. When the team reduced lncRNA1456 (and therefore XCP) levels, PHF8’s grip on DNA weakened at hundreds of gene control regions, and many of those genes changed their activity. In biochemical tests, PHF8 alone only modestly removed a particular repressive tag from histones, but in the presence of XCP it became far more efficient, and this boosting effect increased with more XCP. In cells, expressing a normal version of lncRNA1456 lowered these repressive marks across the genome, whereas a mutant version unable to produce XCP did not, underscoring that it is the protein—not just the RNA—that drives the effect.
Why This Discovery Matters
This work reveals that a gene once labeled noncoding actually produces a small but influential protein that teams up with an epigenetic enzyme to reshape the activity of many genes in breast cancer. XCP helps PHF8 bind to chromatin and enhances its ability to strip off inhibitory chemical marks, altering growth-related pathways in a context-dependent way. For a lay audience, the key message is that our genome still harbors hidden proteins with substantial impact on disease. XCP is one such example: a testis-linked X chromosome protein that, when misexpressed in breast cancer, can either fuel or restrain tumor growth through fine-tuning how the DNA instruction book is read. As scientists continue to explore these “hidden” proteins, they may uncover new biomarkers and drug targets that were invisible when large parts of the genome were dismissed as noncoding.
Citation: Gadad, S.S., Camacho, C.V., Gong, X. et al. X-linked cancer-associated polypeptide (XCP) from lncRNA1456 modulates PHF8 histone demethylase activity to regulate the epigenome, gene expression, and cellular pathways in breast cancer. Oncogene 45, 1557–1571 (2026). https://doi.org/10.1038/s41388-026-03740-w
Keywords: breast cancer, noncoding RNA, epigenetics, microproteins, histone demethylase