Clear Sky Science · en
Rage against the mean: a perspective on measuring fitness of individual phage particles
Why tiny viruses matter to everyday life
Bacteriophages—viruses that infect bacteria—are the most abundant biological entities on Earth and are being explored as tools to fight antibiotic‑resistant infections, clean industrial pipes, and shape healthy microbiomes. Yet almost everything we know about them comes from methods that average the behavior of trillions of particles at once. This article argues that to truly harness phages for science and medicine, we must learn to measure what each individual virus particle does, not just what the crowd does on average.
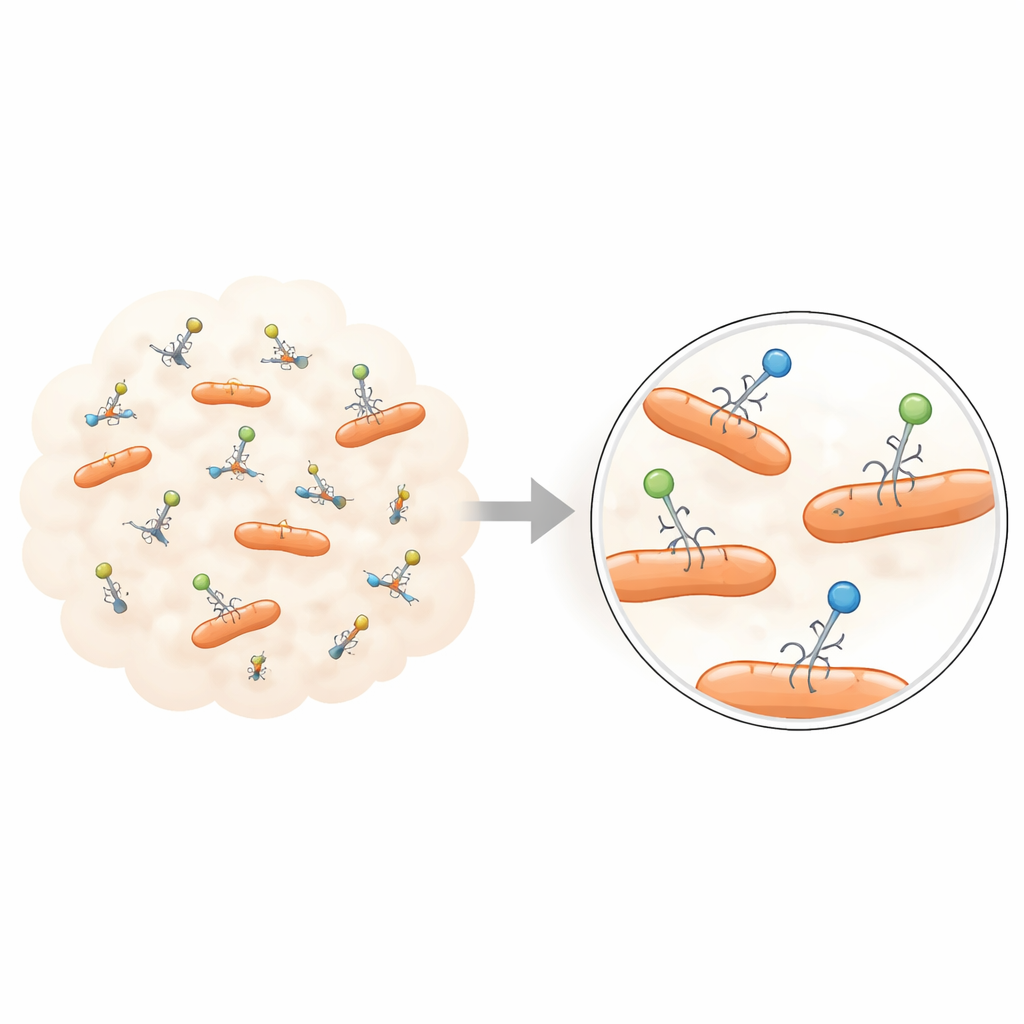
From crowd averages to individual stories
Classic phage experiments were invented in the early 1900s, when viruses were first discovered and visualized. These tools—such as plaque assays and one‑step growth curves—have been enormously successful for mapping the general life cycle of phages: how they attach to a bacterium, inject their genetic material, make copies of themselves, and burst the cell open. But they work by blending together huge populations of viruses and bacteria and then reading out a single number, like an average infection rate or average number of offspring. Any differences between individual virus particles are washed out, leaving a smooth picture that may hide the extremes that matter most for evolution, ecology, and therapy.
Following single viruses through an infection
New imaging and flow‑based technologies are beginning to reveal this hidden diversity. Using advanced optical microscopes, researchers can now label phages with fluorescent dyes or engineered glowing proteins and track their motion as bright spots. These experiments show that when phages first encounter a bacterial cell surface, they do not all behave alike: some bounce off quickly, others wander along the surface, and a fraction latch on tightly. The time each particle spends attached can vary over many seconds, and the motion of bound versus unbound particles reveals that there are multiple interaction states rather than one uniform adsorption step.
Watching viral genomes and bursts one cell at a time
Microscopy has also been used to catch the moment when a phage’s genetic material enters a cell. By staining viral DNA, scientists have watched the glow move from the capsid into the bacterium, and they find that some genomes stream in rapidly while others pause or stall for minutes before completing entry. Similarly, time‑lapse imaging of single infected cells can pinpoint when each cell bursts, showing a spread of lysis times even for genetically identical viruses in the same conditions. Microfluidic devices that hold bacteria in tiny channels, combined with fluorescent reporters for phage components, now allow researchers to follow individual infections from first contact through genome entry, viral production, and lysis within the very same cell.
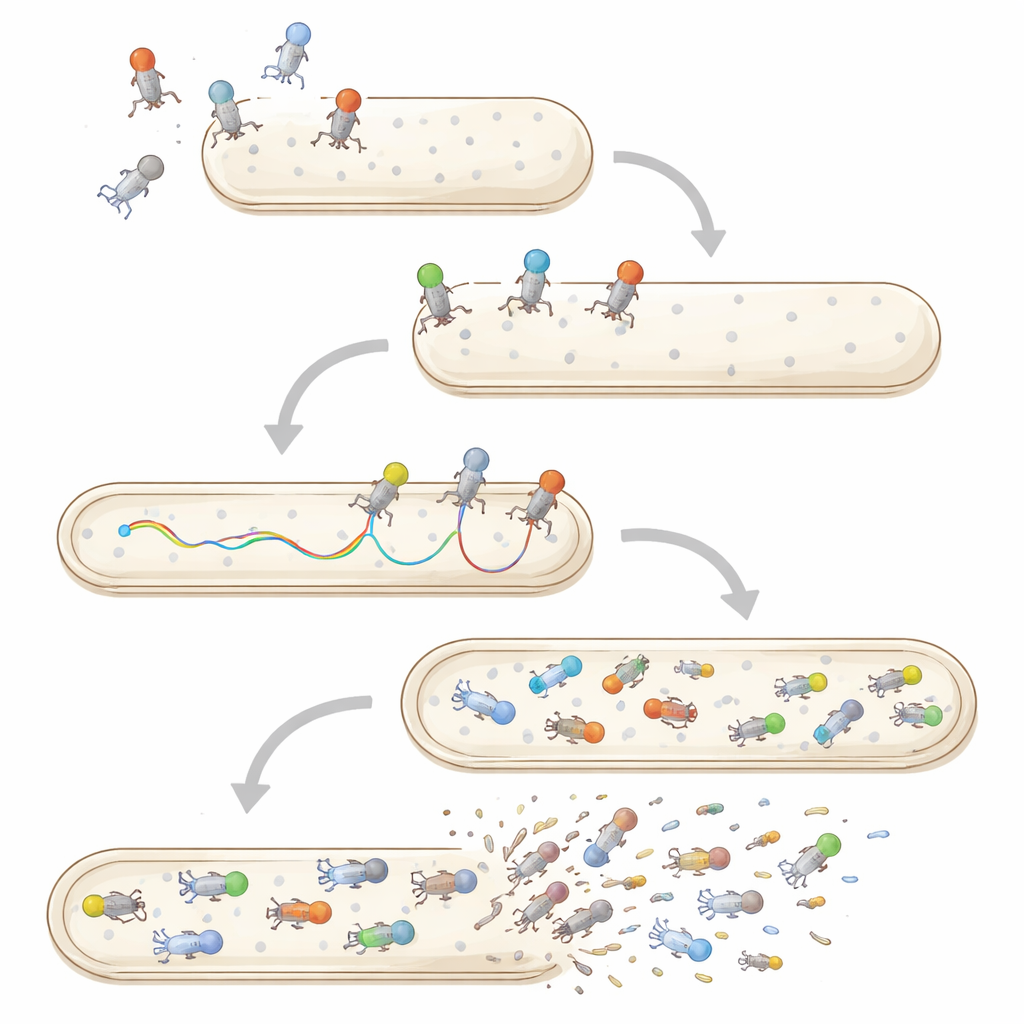
Multiple infections, stability, and life beyond bacteria
Single‑particle and single‑cell methods are also reshaping how we think about co‑infection and virus survival. When several phages attack the same bacterium, imaging and flow cytometry reveal that their genomes do not all enter or replicate equally; some are blocked or delayed, and one lineage may dominate the offspring. Outside bacteria, individual phage particles face stresses such as heat, acidity, or attack by human and animal cells. Traditional tests only report how many viruses remain infectious on average after such exposure, but newer approaches watch single particles diffusing through mucus or being engulfed by mammalian cells, hinting at large differences in which viruses persist or are quickly removed.
What this means for future medicine and technology
The authors argue that embracing this individual‑level variation will be crucial for both basic biology and phage‑based technologies. Evolution acts on differences between particles, so understanding which individual phages bind fastest, inject most reliably, replicate most efficiently, or remain stable longest can explain how viral strains adapt in oceans, soil, and microbiomes. The same information can guide the design of safer, more effective phage therapies and industrial applications—for example, choosing short‑lived phages for one‑time decontamination or long‑lived, highly stable phages for delivering useful genes to bacteria. By moving from averages to single‑particle measurements, researchers can better control which viral traits they select and manufacture, turning phages into more predictable and powerful tools.
Citation: Antani, J.D., Turner, P.E. Rage against the mean: a perspective on measuring fitness of individual phage particles. npj Viruses 4, 21 (2026). https://doi.org/10.1038/s44298-026-00187-4
Keywords: bacteriophages, single-particle virology, phage therapy, optical microscopy, viral evolution