Clear Sky Science · en
Significance and challenges in dissecting cancer-bacteriome interactions
Why Tiny Tenants Matter for Cancer
Every person carries trillions of bacteria that quietly share our bodies, forming rich communities on our skin, in our mouths, lungs, and especially in our guts. This article explores a provocative idea: that these microbial neighbors can help drive cancers or, just as importantly, help hold them back. By examining how whole bacterial communities interact with tumors—and why it is so technically hard to study them—the authors argue that understanding this hidden ecosystem could open new paths for early detection and treatment of cancer.
The Hidden World Inside Us
The human body is home to an enormous “bacteriome,” thousands of bacterial species whose combined cell count rivals our own. Different organs host distinct mixes of bacteria, and those mixes shift over time and from person to person. Researchers now know that some bacterial species can push healthy cells toward cancer by damaging DNA, stoking long-term inflammation, or blocking the body’s built-in brakes on runaway growth. Others appear to do the opposite, rallying immune cells to attack tumors or changing the tumor environment in ways that slow cancer. Yet most of our resident bacteria have never been tested for their effects on cancer, leaving a vast unknown landscape of potential helpers and villains.
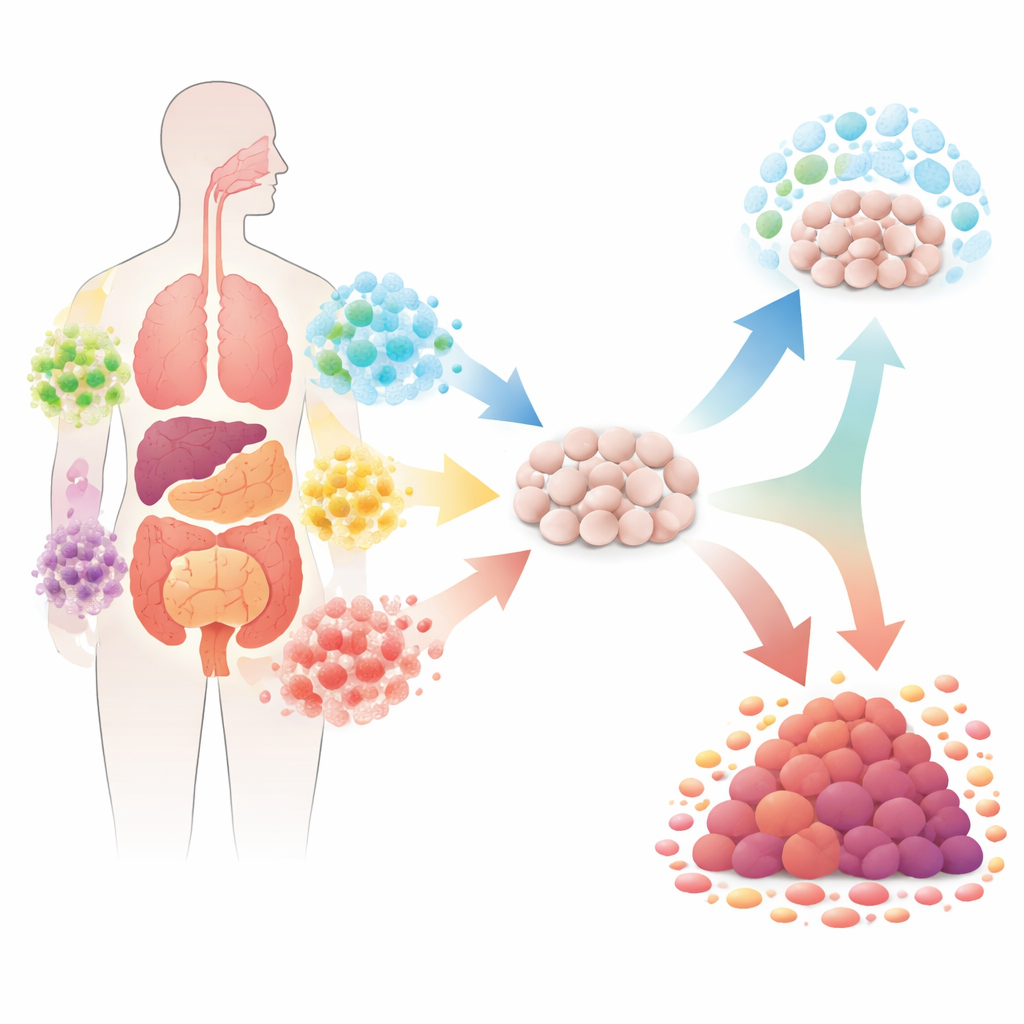
How Bacteria Can Nudge Cells Toward Tumors
The article reviews several well-studied examples that show how bacteria can help spark or fuel tumors. In the stomach, Helicobacter pylori triggers waves of immune cells that release reactive molecules, damaging DNA and promoting long-lasting inflammation that can lead to cancer. Certain strains of Escherichia coli manufacture a small, unstable chemical called colibactin that directly harms DNA, increasing mutation rates in nearby cells. In the colon, Fusobacterium nucleatum and Bacteroides fragilis can both disturb key signaling pathways that control how fast cells divide and when they die. Together, these cases reveal a common pattern: bacteria can shape cancer risk by repeatedly stressing a tissue’s DNA, immune balance, and growth controls over many years.
From Correlation to Cause
Many studies so far have compared the microbiomes of people with and without cancer, using DNA sequencing to catalog which bacteria are present. This work has linked specific species and broader bacterial communities to cancers of the gut, mouth, lungs, liver, skin, and other organs. But such snapshots have limits. They can show association, not proof that a microbe causes disease. They miss bacteria that are dangerous only in certain combinations, and they struggle to identify microbes that quietly protect against cancer. The authors outline several conceptual strategies—from studying one species at a time, to comparing large patient groups, to tracking the joint evolution of cancer cells and complex bacterial mixes—that each capture part of the picture but fall short of fully mapping how the bacteriome shapes tumors.
Shrinking Cancer Experiments to Tiny Droplets
To overcome these gaps, the article highlights microfluidic technologies that shrink experiments into millions of microscopic droplets. Each droplet can act as a tiny test tube, holding a unique mixture of bacteria and human cells. Using clever chip designs, researchers can rapidly generate, grow, and sort these droplets based on simple signals, such as fluorescence that reports cancer cell growth. DNA “barcodes” inside droplets make it possible to later sequence all the genetic material in bulk yet still trace which bacteria and cells came from which droplet. Combined with powerful sequencing methods, this approach could finally let scientists test thousands of different mini-bacterial communities against cancer cells in parallel, revealing both cancer-promoting and cancer-suppressing combinations.
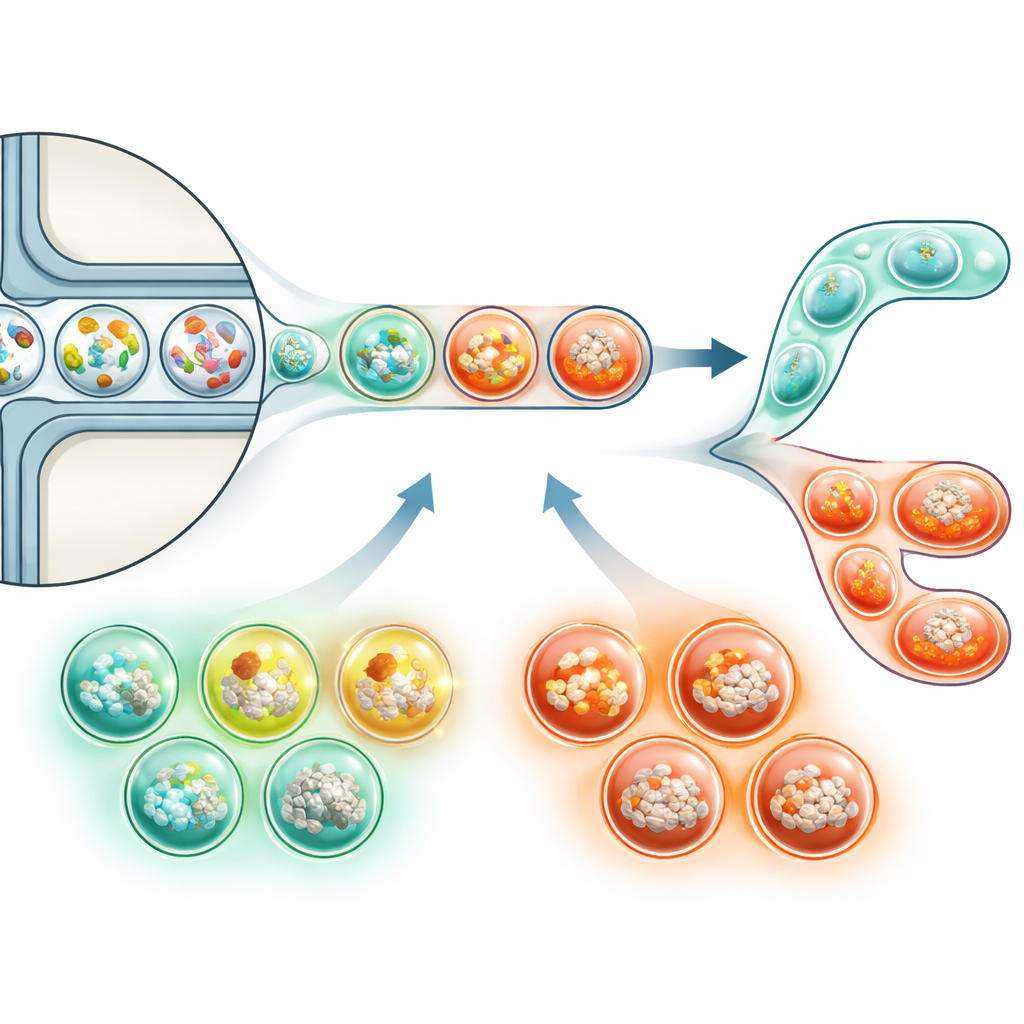
Making Sense of the Data Flood
Such high-throughput experiments generate enormous, complex data sets. The authors describe how statistical tools, dimensionality reduction techniques, network analysis, and modern artificial intelligence can help sift through this flood. These methods can highlight patterns, such as groups of bacteria that tend to appear together in aggressive cancers or in apparently protected tissues, and they can suggest testable hypotheses about how different microbes interact with each other and with tumors. Importantly, computational models grounded in biology can be used to translate these patterns into mechanistic ideas about cause and effect, guiding the next round of experiments.
Turning Microbial Ecology into Medicine
In the end, the article argues that cancer cannot be fully understood by focusing only on human genes and cells. Our resident bacteria form an ecological backdrop that can tilt the balance toward disease or health. By developing new ways to study entire bacterial communities alongside cancer cells—and by pairing these experiments with advanced data-analysis tools—researchers hope to move from merely spotting dangerous microbes to deliberately reshaping the bacteriome. The long-term vision is bold: to prevent or treat cancer not only by targeting tumors directly, but also by engineering the microbial ecosystems that live within us.
Citation: Alshareedah, I., Brunner, J.D., Chain, P.S.G. et al. Significance and challenges in dissecting cancer-bacteriome interactions. BJC Rep 4, 22 (2026). https://doi.org/10.1038/s44276-026-00229-7
Keywords: cancer microbiome, bacteriome, tumor microenvironment, microfluidics, microbial therapy