Clear Sky Science · en
Molecular epidemiology of OXA-1054, a novel carbapenem-hydrolysing Class D β-lactamase, in Enterobacteriaceae isolated from wastewaters
Why germs in dirty water matter
Most of us think of antibiotic resistance as something that happens inside hospitals. But the drugs we use, and the germs they affect, do not stay there. They flow out with wastewater into treatment plants and rivers, where bacteria from many sources mix and swap genetic tricks for surviving our best medicines. This study tracks one such trick—a newly discovered resistance gene—in the wastewater system of Seville, Spain, to understand where it lives, how it moves, and why that matters for future infections.
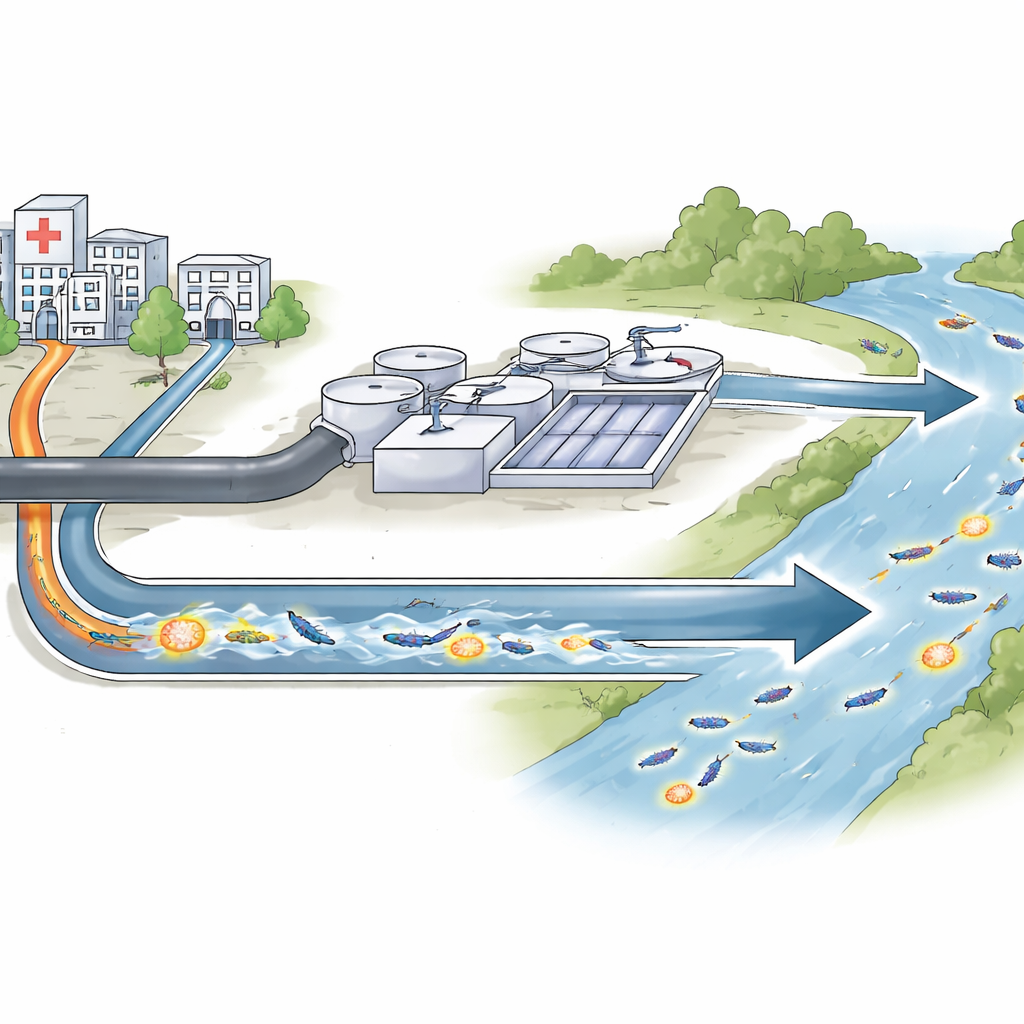
A new resistance trick in common gut bacteria
The researchers focused on a group of bacteria called Enterobacteriaceae, which includes many gut microbes that can cause serious infections. Some members of this group have learned to break down carbapenems, a last-resort family of antibiotics. The team found a previously unknown version of a resistance enzyme, named OXA-1054, in bacteria collected from hospital discharge pipes, city wastewater treatment plants, and the nearby Guadalquivir River. This enzyme belongs to a lineage called OXA-372, related to but distinct from the better-known OXA-48 family that has spread widely in hospitals. By cloning the new gene into harmless lab strains of Escherichia coli, the scientists showed that OXA-1054 can indeed raise resistance to several important drugs, including carbapenems.
Wastewater as a mixing bowl for resistance
Most of the OXA-1054-carrying bacteria were found in raw wastewater flowing into treatment plants, especially at one major facility, with a few isolates in hospital discharge water and in river water downstream. The gene appeared in several different species, such as Citrobacter, Enterobacter, and Raoultella, and in many unrelated clones within those species. This pattern suggests that the resistance gene is not confined to a single successful strain but is spreading across diverse bacteria that share the same aquatic environment. Although treatment reduced detectable bacteria in the treated effluent, the gene still showed up in the river at different times, hinting that environmental waters can act as a long-term reservoir.
A mobile package built to persist
Looking closely at the DNA surrounding the blaOXA-1054 gene, the team found it embedded in a recurring mobile module—essentially a small, portable genetic package. This package almost always contained two transposition genes, istA and istB, from the IS21 family, plus several copies of another mobile element type. In many cases, a second resistance gene (ampC) and clusters for resisting heavy metals like arsenic and mercury were located nearby. These add-ons mean that pollution with metals or other drugs could indirectly favor bacteria that also carry OXA-1054. The plasmids (small DNA circles) that housed this module often carried multiple “toxin–antitoxin” systems, genetic booby traps that help ensure the plasmid is not easily lost when bacteria divide, keeping the resistance trait in the population even when antibiotics are not present.
How it spreads without classic transfer
Surprisingly, most plasmids carrying OXA-1054 lacked the usual machinery for moving between bacteria by conjugation, and laboratory mating experiments failed to transfer the gene into a recipient strain. This points to a different mode of spread: instead of one highly mobile plasmid sweeping through bacterial communities, a conserved mobile platform containing blaOXA-1054 appears to hop into various plasmids and sometimes directly into bacterial chromosomes. Extra accessory genes—such as DNA helicases, DNA methyltransferases, and structural maintenance proteins—may help stabilize these platforms and support their movement and survival in stressful, polluted waters.
What this means for people and rivers
In simple terms, the study shows that a powerful new resistance gene is already circulating in the city’s wastewater and nearby river, even though it has not yet become common in hospital patients. It sits on a sturdy, mobile DNA package that helps it persist and hitchhike with other resistance traits, including defenses against heavy metals. This combination makes it more likely that the gene will endure in the environment and remain ready to enter human-associated bacteria. The work underscores that protecting our most valuable antibiotics requires watching not only clinics, but also the hidden microbial worlds in pipes, treatment plants, and rivers that connect human activity to the broader environment.
Citation: Monge-Olivares, L., González-Pinto, L., Pulido, M.R. et al. Molecular epidemiology of OXA-1054, a novel carbapenem-hydrolysing Class D β-lactamase, in Enterobacteriaceae isolated from wastewaters. npj Antimicrob Resist 4, 36 (2026). https://doi.org/10.1038/s44259-026-00209-4
Keywords: antibiotic resistance, wastewater, carbapenemase, mobile genetic elements, environmental microbiology