Clear Sky Science · en
A deep learning approach to assess transendothelial cell trafficking performance
Watching Immune Cells Cross the Border
Our bodies depend on roaming immune cells that can leave the bloodstream, squeeze between vessel-lining cells, and enter tissues to fight infections and cancers. This border‑crossing act is hard to capture in the lab, especially under realistic blood‑flow conditions. The article presents a new way to watch this journey in detail and then let artificial intelligence (AI) rapidly score how well immune cells make it across, with an eye toward understanding diseases like pancreatic cancer and age‑related immune decline.
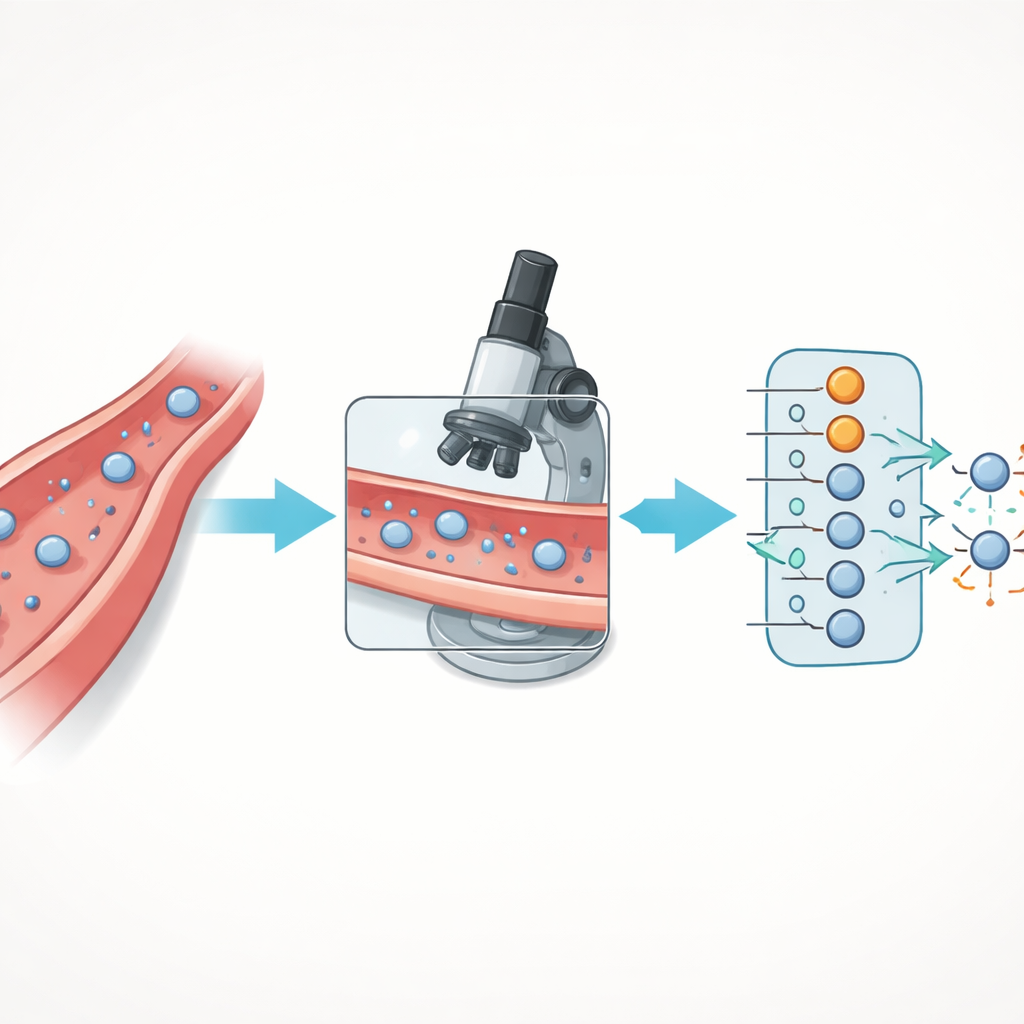
How Cells Sneak Out of Blood Vessels
Immune cells do not simply slip out of blood vessels at random. They follow a carefully choreographed series of steps: first they are caught by the vessel wall, then they roll along it, stop, crawl, and finally push between or through the lining cells to reach the tissue beyond. This multi‑step voyage, called transendothelial migration, relies on a web of surface hooks and signaling molecules on both the immune cells and the vessel‑lining cells. When this sequence is disturbed, immune defenses can falter, tumors can evade attack, or excessive cell entry can drive chronic inflammation and autoimmune disease.
Why Traditional Lab Tests Fall Short
Standard lab assays for cell migration rarely reproduce the forces and structures that cells encounter inside real blood vessels. Simple filter‑based systems let researchers count how many cells move through a membrane, but they largely depend on gravity and cannot mimic the constant shear of flowing blood or reveal each step of the crossing sequence. Animal models capture more complexity but differ from human biology and are expensive and slow. A more realistic method, known as a flow‑based adhesion assay, allows researchers to grow human vessel‑lining cells in narrow channels and pump immune cells past them under controlled flow. Until now, this method has been held back by labor‑intensive, operator‑dependent manual scoring of thousands of cells frame by frame.
Turning Movies of Cells into Data with AI
The authors upgraded the flow assay and paired it with a deep learning model to automate analysis. They grew human endothelial cells inside microfluidic slides, stimulated them to display the right surface hooks, and then flowed fluorescently labeled T cells over this living barrier while recording high‑resolution images with a confocal microscope. Image processing steps removed background noise and separated individual T cells, allowing measurement of simple shape features such as cell area, how elongated they were, and how round their outlines appeared. Instead of asking the AI to distinguish every subtle phase of the journey, the team grouped cells into two key states: those still on or in the vessel wall and those that had fully crossed to the far side.
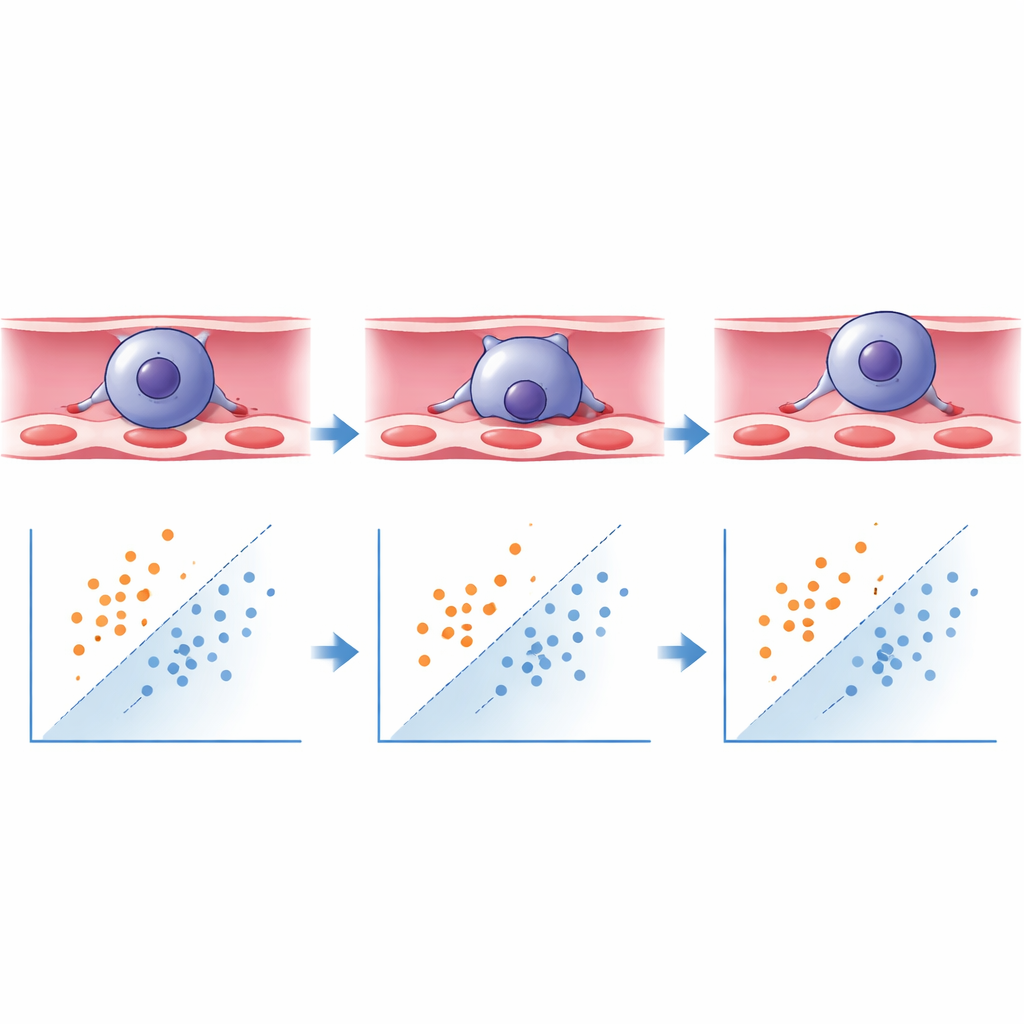
What the Neural Network Learned
Using the Keras/TensorFlow framework, the researchers trained a deep learning model to classify each segmented cell as “transmigrated” or “not yet transmigrated” based on these shape features. The model learned that elongated cells with a high aspect ratio were more likely to be in the midst of, or finished with, crossing, whereas more rounded cells tended to be still attached on the blood side. When tested on new, previously unseen images, the system correctly identified transmigrated cells with over 93% accuracy and non‑transmigrated cells with about 88% accuracy, comfortably exceeding common benchmarks for reliable automated classification. What used to require around ten hours of careful manual scoring by two blinded experts can now be done within minutes, with improved consistency and virtually no operator bias.
Testing Immune Cells from Healthy People and Cancer Patients
To show that the method works across real‑world variation, the team applied their AI‑assisted assay to T cells from healthy donors of different ages and sexes, as well as from patients with pancreatic ductal adenocarcinoma, a cancer known for its highly hostile tumor environment and poor immune infiltration. Healthy donors showed no major differences in crossing performance by age or sex. In contrast, T cells from pancreatic cancer patients were markedly worse at completing the journey across the vessel‑lining cells than T cells from healthy people, regardless of whether the tumor carried a common KRAS mutation or whether the patient had received chemotherapy. These exploratory results hint that the tumor environment may leave a lasting mark on immune cells that weakens their ability to traffic effectively.
A New Tool for Studying and Tailoring Immune Traffic
By marrying a realistic flow‑based assay with deep learning, the authors have created a practical platform that turns complex image sequences into fast, standardized measurements of how well cells cross vessel walls. The approach is flexible enough to be retrained for other immune cells, tumor cells, or organ‑specific endothelial layers, and could be extended to more complex three‑dimensional vessel models in the future. For non‑specialists, the key message is that watching and scoring immune cell traffic no longer needs to be a slow, manual craft: with AI, it can become a robust, high‑throughput tool that helps scientists probe why some immune responses fail and how new treatments might restore the cells’ ability to reach the tissues where they are needed most.
Citation: Schumacher, T.M., Gottloeber, E.M., Koziel, E. et al. A deep learning approach to assess transendothelial cell trafficking performance. Sci Rep 16, 11602 (2026). https://doi.org/10.1038/s41598-026-46045-4
Keywords: immune cell trafficking, transendothelial migration, deep learning microscopy, flow-based adhesion assay, pancreatic cancer immunity