Clear Sky Science · en
Patch type nucleotide sequence identities between genomes from many different species facilitate illegitimate recombination
Hidden Patterns in Life’s Genetic Code
Every living thing, from viruses and bacteria to wheat and whales, stores its genetic instructions in long strings of four chemical “letters.” This study asks a deceptively simple question: what happens when you line up the genetic code of two very different organisms and look for matching stretches? The answer turns out to be surprisingly universal—and may help explain how genomes constantly reshape themselves, fueling evolution and the emergence of new pathogens.
Short Matching Stretches Everywhere
The researchers began by comparing the full genetic sequence of the SARS-CoV-2 virus with a variety of other genomes, including human chromosomes, other viruses, bacteria, plants and animals. Instead of looking for long, obviously related segments, they paid attention to “patches” – short runs of identical letters interrupted by mismatches and gaps. Across more than 90 such cross-species comparisons, they found a striking regularity: about 40–50% of the positions lined up as exact matches, almost always arranged as these scattered, patchy stretches. This held true even for organisms that share no recent common ancestry and perform completely different biological roles.
Randomness That Looks the Same
To see whether these patchy identities reflected deep biological relationships or something more basic, the team created artificial control sequences. They shuffled real genomes to keep the same overall letter composition but scrambled their order, and they also generated fully random DNA strings with similar or fixed base frequencies. When they aligned these synthetic sequences with each other or with real genomes, they saw essentially the same pattern: many short exact matches spread irregularly, with overall identity again clustering around the mid‑40 percent range. They repeated the tests with different alignment programs and scoring settings, and the result hardly budged. The conclusion is that the four-letter alphabet itself, combined with typical genome sizes and letter frequencies, almost guarantees this patchy pattern.
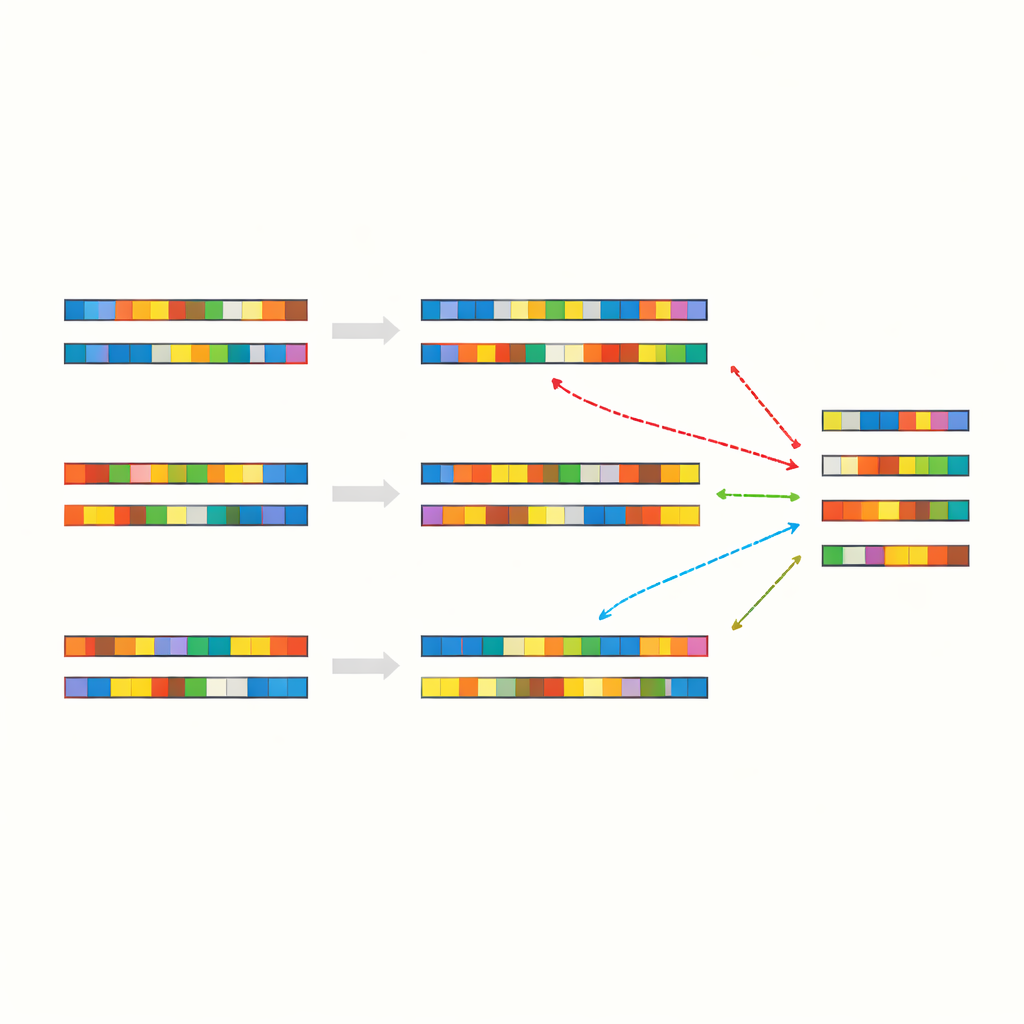
When Chance Becomes a Useful Signal
Patchy matches in DNA are not just a curiosity. Earlier studies, including work by the same group, have shown that similar patterns often appear right where foreign genetic material becomes permanently inserted into a host genome—for example, when certain viruses or mobile DNA elements integrate into animal cells. These events rely on “illegitimate recombination,” a catch-all term for cut-and-paste or copy-and-paste events that do not require long, perfectly matching stretches. The current study strengthens the idea that the ever-present patchy identities produced by basic statistics can act as convenient footholds for the cellular machinery that joins pieces of genetic material together. The authors even identify rare local regions where identity spikes far above random expectations, flagging them as potential hot spots where such recombination is especially likely.
Shaping Genomes Across Evolution
Because these patch patterns turn up in both coding and non-coding regions, in repetitive elements, and across wildly different species, the authors argue that they are a built-in feature of DNA rather than a side effect of particular genes. Over evolutionary time, this constant background of short matching stretches could have made it easier for early genomes to swap, rearrange, or insert new pieces, long before highly specialized enzymes and strict copying mechanisms evolved. In modern organisms, including fast-changing RNA viruses like SARS-CoV-2, the same statistical scaffolding may still help enable rare but consequential exchanges of genetic material with other viruses or even host cells, potentially giving rise to new variants with altered behavior.
What This Means for the Big Picture
To a non-specialist, the key message is that DNA’s four-letter code carries two kinds of information at once. One layer spells out genes and regulatory instructions. The other, more subtle layer is statistical: simply by using four letters with biased frequencies over long stretches, genomes inevitably share many scattered short matches. This study suggests that evolution has made use of that second layer, turning random-looking patterns into practical docking points for genetic reshuffling. In other words, the same simple rules that make sequences look patchily similar across the tree of life may also help living systems continually rewrite and adapt their own blueprints.
Citation: Weber, S., Ramirez, C.M. & Doerfler, W. Patch type nucleotide sequence identities between genomes from many different species facilitate illegitimate recombination. Sci Rep 16, 10524 (2026). https://doi.org/10.1038/s41598-026-44124-0
Keywords: genome recombination, DNA sequence patterns, genetic evolution, SARS-CoV-2 genetics, genome plasticity