Clear Sky Science · en
Cut instance mixing: A domain-specific data augmentation method applied to gastrointestinal lesion detection
Why smarter training data matters for gut health
Doctors use tiny cameras to look inside our digestive tract and spot early signs of disease, such as small abnormal patches that can one day turn into cancer. But these warning signs can be extremely subtle, and computers that might help doctors find them need thousands of well-labeled images to learn what to look for. This paper presents a new way to “grow” realistic training images on a computer, making it easier to teach artificial intelligence systems to detect these hard-to-see lesions early.
The challenge of hidden warning spots
Lesions in the stomach and intestines—like intestinal metaplasia, early dysplasia, and small polyps—are important early warning signs for cancer, yet they often look only slightly different from healthy tissue. Human experts sometimes miss them, and computer models struggle too, mainly because there are not enough high-quality, carefully labeled images to learn from. Collecting and annotating endoscopy images is expensive, time-consuming, and ethically sensitive, especially for rare lesion types. Traditional tricks to boost training data—such as flipping, rotating, or blending whole images together—help in everyday photography, but they tend to blur or misplace the very fine details and boundaries that matter most in gastrointestinal exams.
From rough cut-and-paste to anatomy-aware mixing
Researchers have tried more advanced “cut-and-paste” methods that copy visible lesions from one image and drop them into another, or use generative models to create entirely new images. While these strategies increase variety, they often ignore the surrounding anatomy. A pasted lesion may land in an unrealistic place or clash in color and texture with nearby tissue, making the training images less believable. Generative models like GANs and diffusion networks can produce impressive pictures, but they demand very large datasets, are difficult to control, and risk inventing fake structures that do not exist in real patients. For tasks that depend on subtle color shifts and fine surface patterns—as in early stomach and colon disease—these shortcomings limit how much current augmentation methods can help.
A new way to blend lesions into realistic scenes
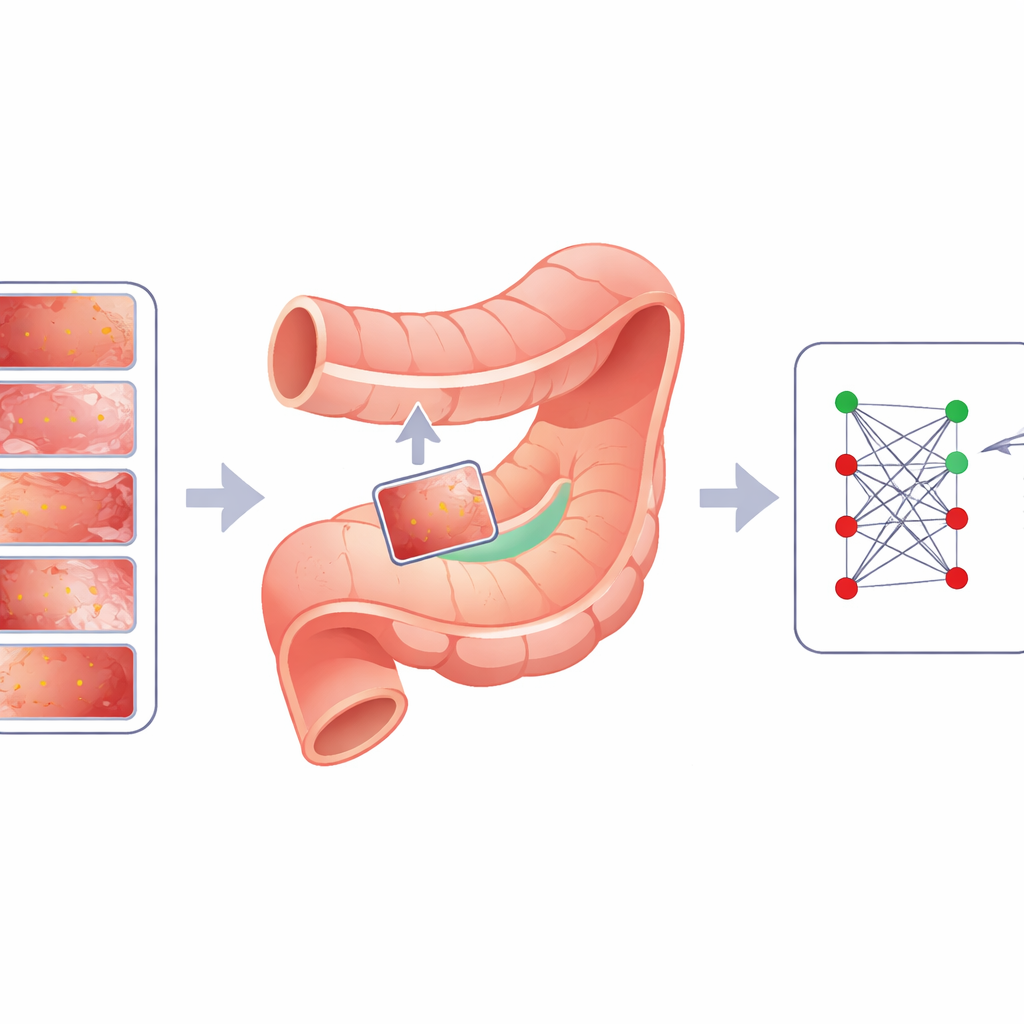
The authors introduce Cut Instance Mixing (CIM), a domain-specific approach designed specifically for gastrointestinal images. Instead of placing lesions randomly, CIM first analyzes a healthy image to find meaningful regions that resemble real mucosal patterns. It does this by clustering pixels into smooth, biologically plausible patches and then breaking them into smaller, tidy subregions. Next, CIM selects a real lesion from another image and searches for the subregion whose color and texture most closely match the lesion’s surroundings. The lesion is then positioned there and blended smoothly using techniques that preserve edges and lighting, with a tuning knob (called α) that controls how strong or subtle the lesion appears. The result is a synthetic image where the lesion looks as if it naturally formed in that spot, and because the original lesion label is known, the new image is automatically annotated for training.
Putting the method to the test
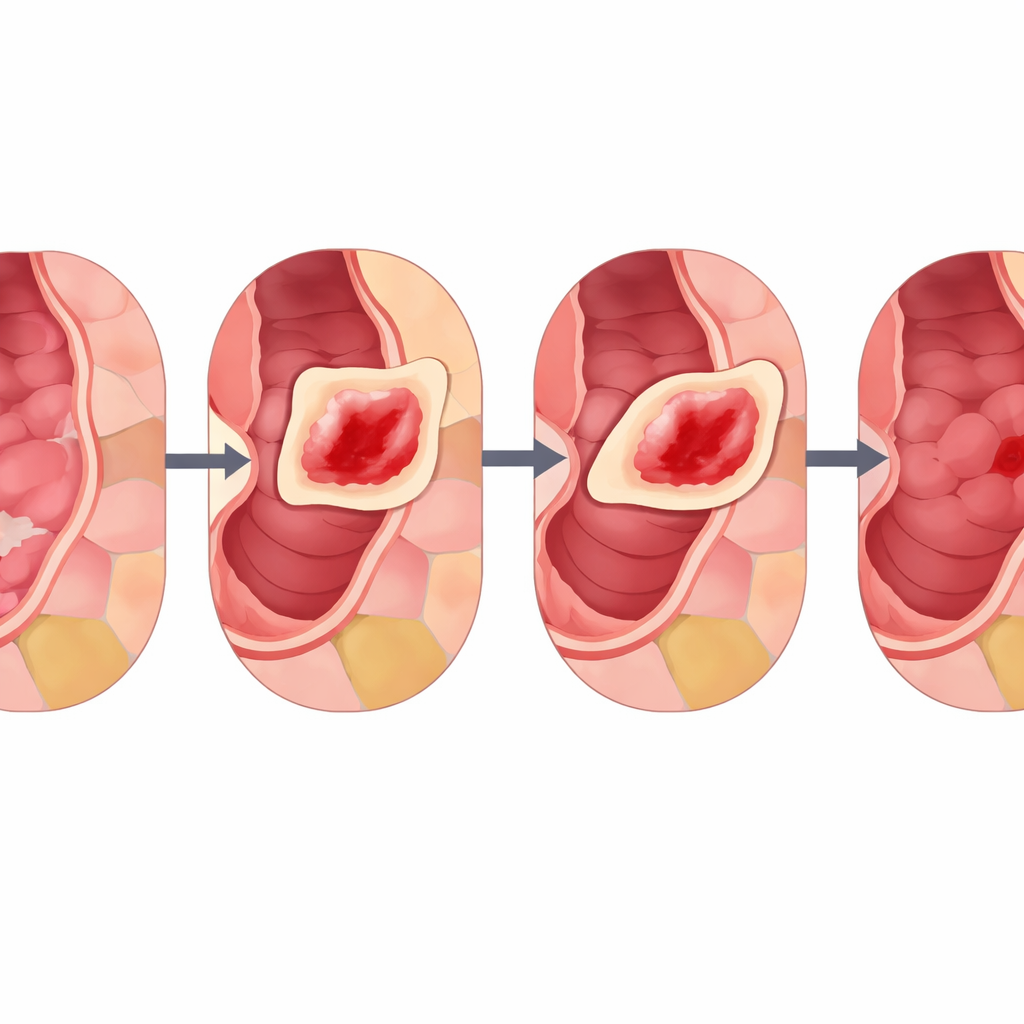
To see whether CIM truly helps, the team trained the same deep learning model on three different endoscopy collections: intestinal metaplasia, dysplasia, and colon polyps. For each dataset, they compared CIM against standard techniques like MixUp, CutMix, and simple random copy–paste, using strict cross-validation and an external test set from other hospitals for the polyp case. Across nearly all measures of performance—such as overall accuracy, ability to separate diseased from healthy tissue, and stability across repeated tests—CIM, especially with stronger blending (α around 0.8), came out on top. The researchers also inspected heatmaps that show where the model “looks” when it makes decisions. Models trained with CIM focused more precisely on true lesion regions, matching medical annotations better than rivals and suggesting that CIM helps the network learn more clinically meaningful features rather than shortcuts or noise.
What this means for future gut cancer screening
In simple terms, CIM gives doctors’ computers more and better practice images by inserting real lesions into believable new locations in a way that respects anatomy. This approach reduces the imbalance between common and rare conditions, strengthens detection of early and subtle disease, and works without heavy, hard-to-control generative models or extra helper networks. While the current work focuses on binary decisions—lesion or no lesion—the same strategy could be expanded to multiple lesion types and tested on other organ systems. If widely adopted, CIM and similar anatomy-aware data “mixing” tools could make computer-assisted endoscopy more reliable, helping specialists find dangerous changes in the digestive tract earlier and more consistently.
Citation: Neto, A., Almeida, E., Libânio, D. et al. Cut instance mixing: A domain-specific data augmentation method applied to gastrointestinal lesion detection. Sci Rep 16, 11941 (2026). https://doi.org/10.1038/s41598-026-42138-2
Keywords: gastrointestinal lesions, endoscopy imaging, data augmentation, deep learning, cancer screening