Clear Sky Science · en
Heart-nosed bat alphacoronaviruses use human CEACAM6 to enter cells
Why bat viruses matter to all of us
After COVID-19, many people wonder where the next pandemic might come from. This study looks ahead by examining little-known coronaviruses that live in bats and asks a simple but crucial question: can any of them already get into human cells? By finding viruses that are “pre-adapted” in this way, scientists hope to spot potential threats early enough for vaccines, drugs and surveillance to be put in place before an outbreak starts.
Searching a crowded viral universe
Alphacoronaviruses are a large group of viruses that mostly infect bats, rodents and domestic animals, but a few already cause common colds in people. Testing every known strain in the lab would be impossible, so the researchers used a computational shortcut. From more than 2,700 spike protein sequences—the part of the virus that latches onto cells—they picked 40 that best captured the overall genetic diversity of the group. They then used these spikes to make harmless “pseudoviruses” that glow when they successfully enter cells, allowing rapid testing of which receptors, in which species, each virus can use.
Most bat viruses can’t use our usual doorways
Human coronaviruses that we already know about, such as the common cold viruses 229E and NL63 or SARS-CoV-2, use a handful of well-studied molecules on our cells as entry points, including proteins called ACE2, APN and DPP4. When the team tested their 40-virus panel against libraries of these receptors from many mammals, they found that very few alphacoronaviruses could use them at all, and almost none could use the human versions. This suggests that the classic coronavirus “door handles” are the exception rather than the rule in this branch of the virus family tree. For most bat alphacoronaviruses, some other, unknown route into cells must exist.
A heart‑nosed bat virus finds a new human handle
One virus, however, stood out. A strain called CcCoV‑KY43, originally found in heart‑nosed bats in Kenya, was able to enter two human cell lines derived from lung and gut tissue. To work out how, the researchers used the virus’s spike tip—the receptor-binding domain—as bait against a large panel of 759 human cell-surface proteins. Three closely related candidates popped up, all from the CEACAM family, which normally help cells stick together and are abundant on mucous surfaces. Follow‑up experiments narrowed this down to a single key partner: a protein called CEACAM6. When human cells that are usually resistant were engineered to make CEACAM6, they suddenly became infectable by the bat virus spike. Blocking or reducing CEACAM6 cut infection sharply, confirming that this protein acts as the virus’s doorway.
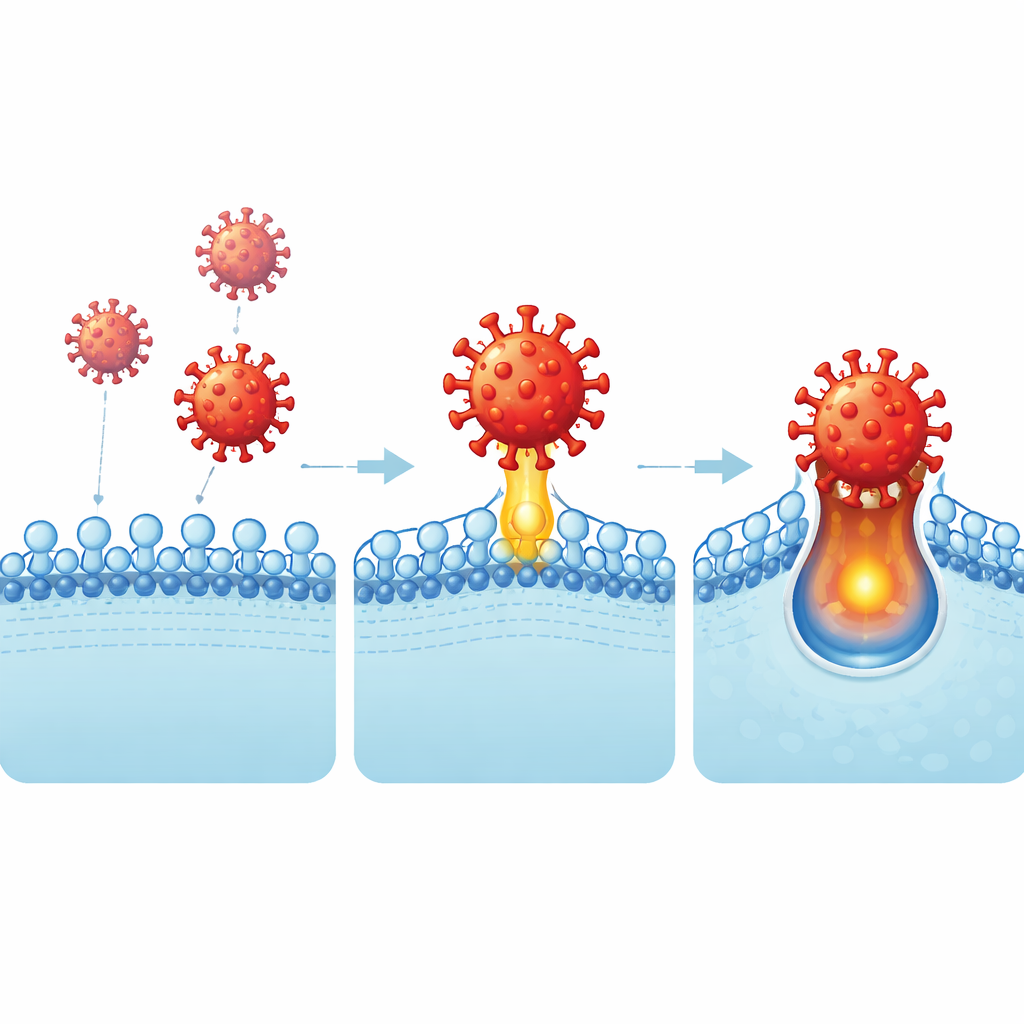
What the molecular embrace looks like
To see exactly how this interaction works, the team crystallized the bat virus binding domain together with CEACAM6 and solved the structure. They found that three tiny loops on the viral spike clasp the very tip of CEACAM6’s outermost domain, forming a snug interface dominated by hydrophobic (water‑repelling) contacts. This is the same region of CEACAM6 that usually contacts its partner proteins, meaning the virus has evolved to mimic or disrupt normal cell‑cell interactions. Subtle changes in just a few amino acids at this interface were enough to strengthen or weaken viral entry, explaining why close relatives such as CEACAM5 bind more weakly and do not function as effective receptors.
How widespread is this risky trait?
The story does not end with a single virus. Two additional coronaviruses from the same bat species in another part of Kenya, and several related viruses from horseshoe bats in China and Russia, also turned out to use CEACAM6-like proteins as entry doors. Some could use human CEACAM6, others only the bat versions, depending on fine‑tuned sequence details in both virus and host. By examining evolutionary trees, the authors conclude that the ability to use CEACAM6 probably arose at least twice, independently, in different alphacoronavirus lineages. Importantly, when they screened blood samples from hundreds of people living near the Kenyan bat colonies, they found only scattered signs of antibodies recognizing these bat viruses, arguing against major, ongoing spillover into humans, though rare or short-lived infections cannot be excluded.
What this means for future outbreaks
In everyday terms, this work shows that some bat coronaviruses already know how to open a particular “door” on human lung cells, even though they have not yet caused recognized disease. CEACAM6 is widespread in the human airways and appears to be used by a geographically dispersed set of alphacoronaviruses, especially in East Africa and parts of Eurasia. That combination—ready-made access to our cells and broad distribution in wildlife—marks these viruses as ones to watch. By identifying the exact receptor and mapping which viral strains can use it, the study provides concrete targets for surveillance, risk scoring, and eventually vaccines or antivirals, moving pandemic preparedness from guesswork toward a more systematic, evidence-based footing.
Citation: Gallo, G., Di Nardo, A., Lugano, D. et al. Heart-nosed bat alphacoronaviruses use human CEACAM6 to enter cells. Nature 653, 180–189 (2026). https://doi.org/10.1038/s41586-026-10394-x
Keywords: bat coronaviruses, CEACAM6, zoonotic spillover, virus receptors, pandemic preparedness