Clear Sky Science · en
Structural basis of membrane potential coupled vectorial CO₂ hydration by the DAB2 complex in chemolithoautotrophs
How microbes help lock away carbon
Hidden in the oceans and in sulfur-rich sediments are microbes that live entirely on inorganic chemicals and carbon dioxide. This study uncovers how one such group of bacteria uses a tiny molecular machine, called the DAB2 complex, to pull CO₂ out of its surroundings and convert it into a form the cell can use. Understanding this natural carbon pump matters for basic climate science and could eventually inspire new ways to capture CO₂ from the environment.
A microscopic carbon pump
Many microorganisms build their bodies from dissolved inorganic carbon, a mix of CO₂ and closely related molecules in water. They rely on a sluggish enzyme, RuBisCO, to bolt CO₂ onto organic compounds, a reaction at the heart of the global carbon cycle. Because RuBisCO is slow and easily disrupted, many microbes have evolved CO₂-concentrating mechanisms that stockpile usable carbon right where RuBisCO works. While these systems are well known in photosynthetic cyanobacteria, chemolithoautotrophs—bacteria that gain energy by oxidizing inorganic substances like sulfur—have remained mysterious. Earlier work pointed to a family of membrane complexes, collectively called DACs, that help these bacteria grow when carbon is scarce. One such system, DAB2 from the bacterium Halothiobacillus neapolitanus, can rescue carbon-starved E. coli, hinting that it actively accumulates inorganic carbon inside cells.
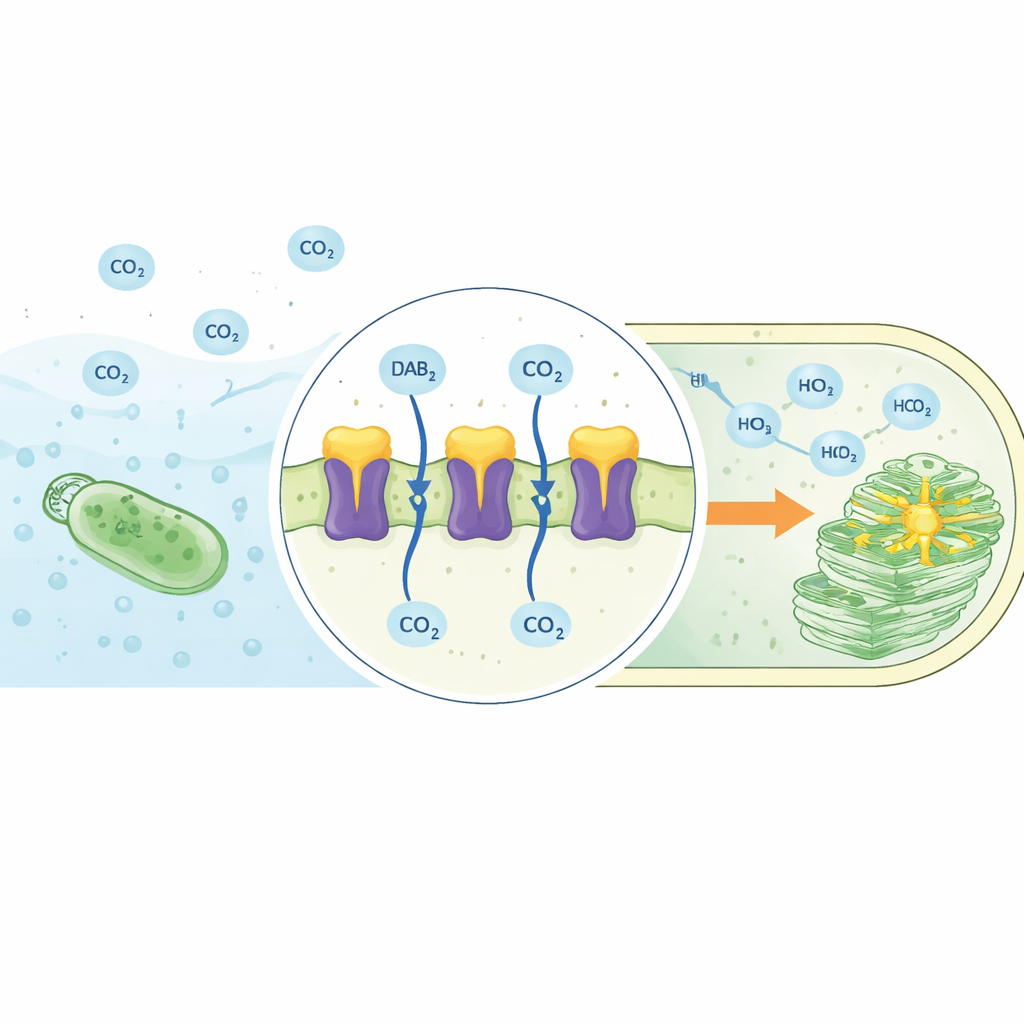
The DAB2 complex revealed in 3D
The authors used high-resolution cryogenic electron microscopy to determine the structure of the DAB2 complex embedded in a membrane-like environment. DAB2 turned out to be a two-part machine: an upper, soluble protein (DabA2) that sits on the inner side of the membrane, and a lower, membrane-spanning protein (DabB2) that forms a channel through the lipid layer. DabA2 resembles known carbonic anhydrases, enzymes that normally speed up the interconversion of CO₂ and bicarbonate, but its architecture is heavily modified. It contains two related catalytic domains and a unique transmembrane “finger” that reaches down into DabB2. DabB2, in turn, looks strikingly similar to proton-conducting subunits from respiratory Complex I, a central player in cellular energy conversion. This combination suggests that DAB2 links a carbon-processing center to a proton-driven power source.
A hidden reaction chamber with gated tunnels
Closer inspection of DabA2 revealed a single zinc ion at the center of a deeply buried pocket, where CO₂ and bicarbonate bind. Unlike textbook carbonic anhydrases, this active site lacks a key amino acid that normally stabilizes the fleeting transition state of the reaction. The site is reachable only through narrow, winding tunnels lined mostly with water-repelling residues. Using both structural analysis and infrared spectroscopy, the researchers showed that the pocket can bind multiple CO₂ molecules strongly but does not, by itself, rapidly convert them to bicarbonate. The tight geometry of the tunnels creates bottlenecks that likely have to open and close in a controlled way, acting as gates that regulate when CO₂ can enter and when bicarbonate can leave.
Proton power and one-way traffic
DabB2’s structure and a series of targeted mutations indicate that it forms a pathway for protons—positively charged hydrogen ions—across the membrane. Key charged and polar residues align to form a continuous route for proton transfer, reminiscent of the “water wires” that carry protons in respiratory complexes. The unusual helical extension from DabA2 slots into DabB2 right where a second half-channel would be expected, apparently helping to form the outlet for protons. When the authors disrupted specific residues along this route, the complex could no longer support growth of carbon-starved E. coli, even though the proteins were still produced. Additional tests showed that DAB2 does not rely on sodium gradients, setting it apart from related complexes in some pathogens; instead, it appears to be driven purely by the membrane’s protonmotive force.
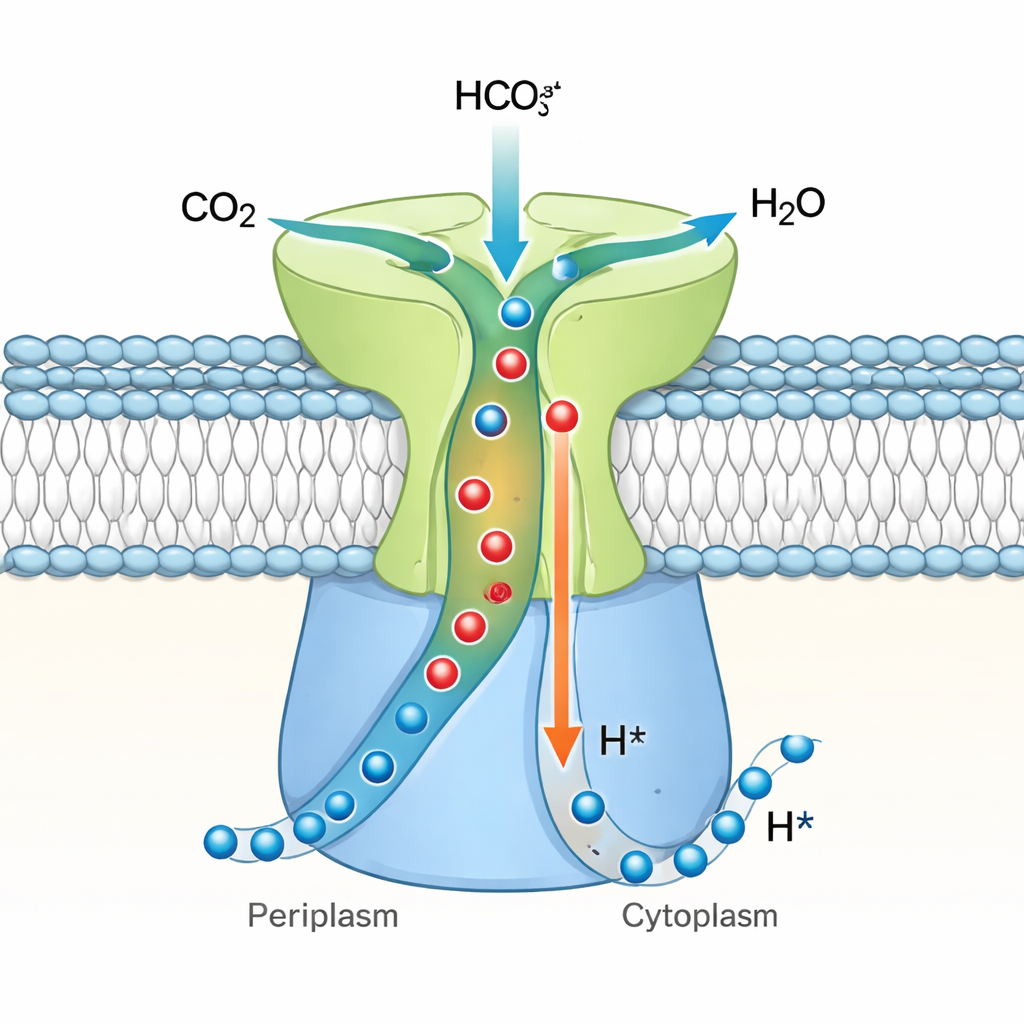
A one-way valve for cellular carbon
Bringing these findings together, the authors propose that DAB2 is a vectorial carbonic anhydrase—a one-way CO₂ hydrator tightly coupled to proton flow. In their model, CO₂ and water can slip into the buried active site when the system is in a “closed” resting state, but the geometry of the tunnels and steric hindrance prevent bicarbonate from coming back in, blocking the reverse reaction. When a proton gradient exists across the membrane, protons move through DabB2 and trigger subtle shifts in the DAB2 structure, likely transmitted by the DabA2 “finger.” These shifts open the tunnels and reorganize the active site so that CO₂ is efficiently converted to bicarbonate and released into the cell interior, while protons are carried across the membrane. This design lets chemolithoautotrophic bacteria tie carbon uptake directly to their energy status, ensuring that CO₂ capture proceeds only when there is enough proton power to drive it. The work establishes DAB2 as a prototype of a widespread, proton-driven family of carbon-converting machines and broadens our picture of how microbes engineer their own microscopic carbon pumps.
Citation: Lo, Y.K., Seletskiy, M., Bohn, S. et al. Structural basis of membrane potential coupled vectorial CO₂ hydration by the DAB2 complex in chemolithoautotrophs. Nat Commun 17, 4071 (2026). https://doi.org/10.1038/s41467-026-72558-7
Keywords: carbon dioxide uptake, chemolithoautotrophs, carbonic anhydrase, protonmotive force, membrane protein structure